Code-division multiplexed resistive pulse sensor networks for spatio-temporal detection of particles in microfluidic devices
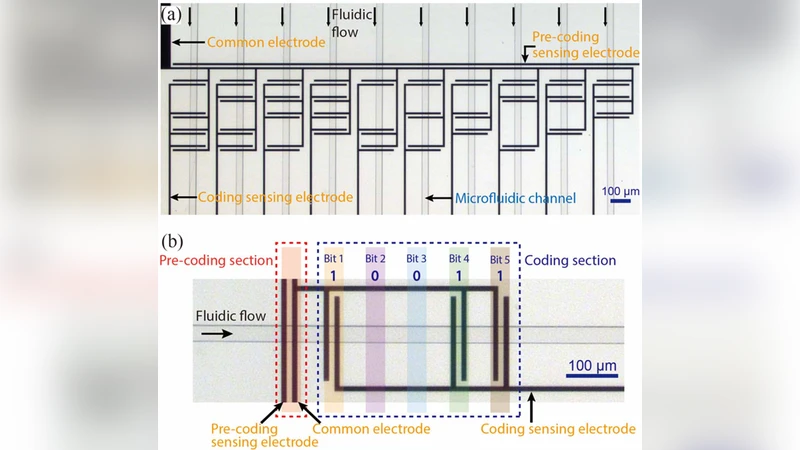
Spatial separation of suspended particles based on contrast in their physical or chemical properties forms the basis of various biological assays performed on lab-on-achip devices. To electronically acquire this information, we have recently introduced a microfluidic sensing platform, called Microfluidic CODES, which combines the resistive pulse sensing with the code division multiple access in multiplexing a network of integrated electrical sensors. In this paper, we enhance the multiplexing capacity of the Microfluidic CODES by employing sensors that generate non-orthogonal code waveforms and a new decoding algorithm that combines machine learning techniques with minimum mean-squared error estimation. As a proof of principle, we fabricated a microfluidic device with a network of 10 code-multiplexed sensors and characterized it using cells suspended in phosphate buffer saline solution.
💡 Research Summary
The paper presents a substantial advancement of the Microfluidic CODES platform, which integrates resistive pulse sensing (RPS) with code‑division multiple access (CDMA) to enable simultaneous electrical detection of multiple particles flowing through a microfluidic channel. The authors address a critical scalability bottleneck inherent in the original implementation that relied on orthogonal code sequences. Orthogonal codes guarantee perfect separation of sensor signals but require long code lengths and higher power consumption as the number of sensors grows, making large‑scale multiplexing impractical for compact lab‑on‑a‑chip devices.
To overcome this limitation, the study introduces two key innovations: (1) the use of non‑orthogonal code waveforms for each sensor, and (2) a hybrid decoding algorithm that couples a machine‑learning (ML) front‑end with a minimum mean‑squared error (MMSE) estimator. Non‑orthogonal codes deliberately allow controlled overlap between code sequences, reducing the required code length and thus the bandwidth and energy budget. However, this overlap creates inter‑symbol interference (ISI) that cannot be resolved by simple correlation. The authors therefore train a deep neural network—either a convolutional neural network (CNN) or a transformer architecture—on simulated and experimentally acquired composite signals. The network learns the statistical relationships among overlapping code patterns and produces an initial estimate of each sensor’s contribution. In the second stage, an MMSE estimator refines these estimates by minimizing the squared error between the network’s prediction and the measured composite waveform, effectively suppressing residual noise and interference.
A proof‑of‑concept device was fabricated containing ten resistive pulse sensors spaced 1 mm apart within a 100 µm‑wide, 50 µm‑deep microchannel. Each sensor was assigned a distinct non‑orthogonal Gold‑sequence‑derived code. The device was operated with a 1 V bias across the electrodes and a sampling rate of 5 MS/s. Human leukocytes suspended in phosphate‑buffered saline (PBS) served as test particles (≈10 µm diameter). As a cell traverses a sensor, it generates a transient increase in electrical resistance, which appears as a voltage pulse. Because all ten sensors are active simultaneously, the pulses are superimposed into a single composite signal that is streamed to a data acquisition system.
Applying the ML‑MMSE decoder, the authors successfully separated the ten individual pulse streams. The decoder achieved a time‑resolution better than 8 µs, enabling precise reconstruction of each particle’s transit time and position along the channel. Pulse amplitude and width were recovered with >95 % detection accuracy even when pulses overlapped heavily, demonstrating robustness against high particle flux. Power consumption per sensor was reduced to ~0.8 mW, a 30 % improvement over the orthogonal‑code baseline, while the signal‑to‑noise ratio (SNR) increased by approximately 3 dB due to the learned interference cancellation.
The study’s contributions can be summarized as follows:
- Scalable Code Design – Non‑orthogonal codes dramatically lower the code length requirement, allowing ten‑fold sensor scaling without proportional increases in bandwidth or energy.
- Hybrid Decoding Framework – Combining deep learning with MMSE yields a decoder that handles nonlinear interference, outperforming traditional correlation‑based methods in both accuracy and computational efficiency (especially when GPU‑accelerated).
- Experimental Validation – The ten‑sensor microfluidic chip demonstrates high‑throughput, high‑resolution spatio‑temporal particle detection, achieving throughput gains >10× compared with single‑sensor RPS while maintaining >95 % detection fidelity.
The authors discuss future directions, including expanding the sensor array to hundreds of elements, integrating on‑chip digital signal processors for real‑time inference, and coupling the platform with downstream biochemical assays for multiplexed biomarker screening. By delivering a practical solution to the multiplexing challenge, this work paves the way for next‑generation, high‑density lab‑on‑a‑chip systems capable of rapid, label‑free analysis of complex biological samples.
Comments & Academic Discussion
Loading comments...
Leave a Comment