Towards Commodity, Web-Based Augmented Reality Applications for Research and Education in Chemistry and Structural Biology
This article reports prototype web apps that use commodity, open-source technologies for augmented and virtual reality to provide immersive, interactive human-computer interfaces for chemistry, structural biology and related disciplines. The examples, which run in any standard web browser and are accessible at https://lucianoabriata.altervista.org/jsinscience/arjs/armodeling/ together with demo videos, showcase applications that could go well beyond pedagogy, i.e. advancing actual utility in research settings: molecular visualization at atomistic and coarse-grained levels in interactive immersive 3D, coarse-grained modeling of molecular physics and chemistry, and on-the-fly calculation of experimental observables and overlay onto experimental data. From this playground, I depict perspectives on how these emerging technologies might couple in the future to neural network-based quantum mechanical calculations, advanced forms of human-computer interaction such as speech-based communication, and sockets for concurrent collaboration through the internet -all technologies that are today maturing in web browsers- to deliver the next generation of tools for truly interactive, immersive molecular modeling that can streamline human thought and intent with the numerical processing power of computers.
💡 Research Summary
The paper presents a suite of prototype web applications that bring augmented reality (AR) and virtual reality (VR) to chemistry, structural biology, and related fields using only standard web browsers. By leveraging open‑source, client‑side JavaScript libraries—A‑Frame for declarative 3D scene construction, AR.js for marker‑based camera tracking, Three.js for WebGL rendering, Cannon.js (via the A‑Frame‑physics plugin) for real‑time rigid‑body dynamics, annyang for speech recognition, and Google Charts for data visualization—the author demonstrates that sophisticated, interactive molecular tools can be deployed without any software installation or specialized hardware.
The prototypes are organized into two main categories. The first set focuses on small molecules. Starting from a pure HTML example that displays lysine and glutamate side chains attached to Hiro and Kanji markers, successive layers of functionality are added: real‑time distance measurement, Coulombic force calculation, clash detection, haptic feedback via phone vibration, stochastic proton‑transfer simulations, and a Diels‑Alder reaction visualizer that animates bond formation as the reactants approach. A particularly notable demo computes paramagnetic pseudocontact chemical shifts and line broadening for a probe hydrogen atom moving around a metmyoglobin heme group, generating a synthetic NMR spectrum on the fly with Google Charts.
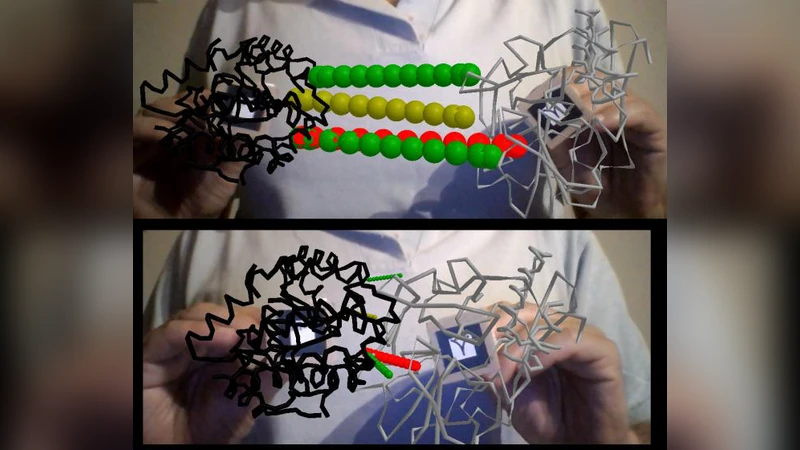
The second set addresses macromolecules, primarily proteins. Users can load PDB files and view them at full atomistic detail, or at coarse‑grained resolutions (one bead per residue centered on Cα atoms, or multi‑bead schemes reminiscent of MARTINI). The applications incorporate physics‑based interactions: Cannon.js supplies contact forces and constraints, enabling users to feel repulsion or attraction as they manipulate protein fragments with AR markers. Advanced examples overlay real‑time small‑angle X‑ray scattering (SAXS) profiles, allowing direct comparison between computed scattering curves and experimental data. A manual docking tool lets users guide protein–protein interfaces by moving markers, with contact data updating instantly.
The author acknowledges that true quantum‑mechanical energy evaluations are too computationally intensive for in‑browser execution. However, he points out that recent machine‑learning potentials (e.g., ANI, SchNet, DeepMD) can approximate quantum calculations orders of magnitude faster, making real‑time, physics‑accurate reactivity exploration feasible within a web AR/VR context. Future integration with emerging WebXR standards could eliminate the need for fiducial markers, while socket‑based networking would enable multi‑user collaborative sessions. Voice commands, already demonstrated via annyang, hint at more natural human‑computer interaction.
Overall, the work demonstrates that web‑based AR/VR has matured from a novelty visualization tool into a viable platform for interactive molecular modeling, data analysis, and collaborative research. By building entirely on open standards, the prototypes are highly accessible, portable across devices, and readily extensible with emerging computational and interaction technologies. This positions web AR/VR as a promising foundation for the next generation of immersive scientific tools that blend human intuition with the computational power of modern chemistry and structural biology workflows.
Comments & Academic Discussion
Loading comments...
Leave a Comment