Building a Controlled Vocabulary for Standardizing Precision Medicine Terms
Rapid advances of technology and development of research in precision medicine domain have led to the production of different types of biomedical data. Standard medical vocabularies were shown to be limited in dealing with such heterogeneous data and consequently, new controlled vocabulary for data integration and normalization has been proposed. In this study, the precision medicine vocabulary (PMV), which is a controlled vocabulary for terms used in precision medicine, is built based on the method of data integration in Unified Medical Language System (UMLS). It now covers ten top semantic types of disease, drug, gene, gene variation and so on. In total of 1,372,967 concepts and 4,567,208 terms have been integrated from widely used databases related with precision medicine.
💡 Research Summary
The paper presents the design, construction, and evaluation of the Precision Medicine Vocabulary (PMV), a controlled vocabulary aimed at standardizing the heterogeneous terminology used in precision medicine research. Recognizing that the Unified Medical Language System (UMLS) – while extensive – lacks the granularity required for the molecular‑level concepts central to precision medicine, the authors set out to create a domain‑specific lexical resource that can support data integration, knowledge discovery, and the development of a Precision Medicine Knowledge Base (PMKB).
The scope of PMV is defined around ten top‑level semantic types that reflect the core entities of precision medicine: Anatomical Structure, Gene, Gene Product, Mutation, Cell, Disease, Phenotypic Abnormality, Biological Pathway, Biological Function, and Chemical/Drug. These categories were chosen to capture both clinical and molecular dimensions, from tissue anatomy to genomic variants and therapeutic agents.
Data collection began with a systematic extraction of concepts from UMLS, filtered by the ten semantic types and limited to human species and English language entries. After pruning, 52 source vocabularies were retained, including broad resources such as MeSH, NCIt, and SNOMED CT, as well as domain‑specific databases like HGNC, OMIM, HPO, DrugBank, and RxNorm. The authors then enriched this foundational set by integrating additional heterogeneous resources (DrugBank, ClinVar, NCBI Gene) that were either incompletely represented in UMLS or entirely absent.
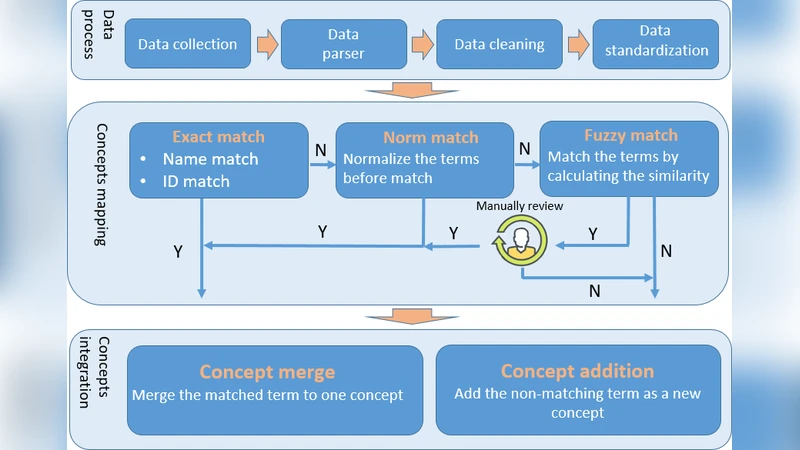
Integration followed a four‑step matching pipeline: (1) exact match using concept name, identifier, and source; (2) normalized match employing the UMLS “Norm” lexical tool (which strips possessives, replaces punctuation, removes stop‑words, lower‑cases, tokenizes, and alphabetically sorts tokens); (3) fuzzy match to capture near‑matches; and (4) expert manual review of the top five fuzzy candidates. Matched terms were merged into existing PMV concepts; unmatched terms gave rise to new concepts. This process yielded 4,571 new drug concepts with 150,629 terms from DrugBank, 21,172 gene concepts with 220,328 terms from NCBI Gene, and 294,712 mutation concepts with 316,630 terms from ClinVar.
PMV adopts a simplified identifier scheme derived from UMLS: MCID for concepts, MAID for terms, and MTID for semantic types. Each concept links to at least one term, and a preferred term is selected based on source priority. Semantic types are organized hierarchically; the top‑level taxonomy mirrors UMLS inheritance, while lower‑level classifications are built by mapping terms from MeSH, NCIt, HPO, ClinVar, and NCBI Gene into a “Subclass_of” structure.
Quantitative comparison with UMLS across the ten semantic types shows that PMV surpasses UMLS in areas most relevant to precision medicine: gene (66,022 vs 46,948 concepts), mutation (320,672 vs 25,715), and pathway (2,157 vs 0). Conversely, UMLS retains higher counts in broader medical categories such as anatomical structure and basic biological function, reflecting its more general scope. The authors interpret these results as evidence that PMV provides superior coverage for molecular and therapeutic entities while still maintaining reasonable breadth in clinical concepts.
The discussion highlights PMV’s strengths—high coverage of precision‑medicine‑specific entities, scalability through modular integration, and open‑source availability—and acknowledges gaps, notably limited representation of basic medical concepts and the need for finer‑grained semantic typing. Future work includes expanding the taxonomy (e.g., incorporating Pathway Commons), enriching term attributes and inter‑term relationships, developing an automated integration pipeline for mapping, merging, exporting, and version control, and publishing the database in multiple formats (RDF, JSON) beyond the current MySQL implementation.
In conclusion, the authors have delivered a robust, extensible controlled vocabulary that addresses a critical bottleneck in precision medicine data interoperability. By aggregating over 1.3 million concepts and 4.5 million terms from a wide array of authoritative sources, PMV lays the groundwork for more effective data integration, knowledge base construction, and ultimately, the translation of genomic insights into clinical practice.
Comments & Academic Discussion
Loading comments...
Leave a Comment