Using routinely collected patient data to support clinical trials research in accountable care organizations
Background: More than half (57%) of pharma clinical research spend is in support of clinical trials. One reason is that Electronic Health Record (EHR) systems and HIPAA privacy rules often limit how broadly patient information can be shared, resulting in laborious human efforts to manually collect, de-identify, and summarize patient information for use in clinical studies. Purpose: Conduct feasibility study for a Rheumatoid Arthritis (RA) clinical trial in an Accountable Care Organization. Measure prevalence of RA and related conditions matching study criteria. Evaluate automation of patient de-identification and summarization to support patient cohort development for clinical studies. Methods: Collect original clinical documentation directly from the provider EHR system and extract clinical concepts necessary for matching study criteria. Automatically de-identify Protected Health Information (PHI) protect patient privacy and promote sharing. Leverage existing physician expert knowledge sources to enable analysis of patient populations. Results: Prevalence of RA was four percent (4%) in the study population (mean age 53 years, 52% female, 48% male). Clinical documentation for 3500 patient were extracted from three (3) EHR systems. Grouped diagnosis codes revealed high prevalence of diabetes and diseases of the circulatory system, as expected. De-identification accurately removed 99% of PHI identifiers with 99% sensitivity and 99% specificity. Conclusions: Results suggest the approach can improve automation and accelerate planning and construction of new clinical studies in the ACO setting. De-identification accuracy was better than previously approved requirements defined by four (4) hospital Institutional Review Boards.
💡 Research Summary
The paper presents a feasibility study that demonstrates how routinely collected electronic health record (EHR) data can be leveraged to accelerate patient cohort identification for clinical trials within an Accountable Care Organization (ACO). Using rheumatoid arthritis (RA) as a case study, the authors extracted raw clinical documentation from three disparate EHR platforms (Practice Fusion, GE Centricity, and drchrono) for 3,500 patients affiliated with the NACORS network. The extracted data were transformed into standardized vocabularies—including ICD‑9/10, SNOMED‑CT, RxNorm, LOINC, and HL7 demographic codes—and then grouped using expert‑curated hierarchies such as the Clinical Classification System (CCS) and UMLS Metathesaurus. This standardization enabled consistent identification of diagnoses, medications, and laboratory panels across heterogeneous systems.
A central component of the workflow was an automated de‑identification engine developed by Medal. The engine identified and redacted protected health information (PHI) with 98.7 % sensitivity and 99.1 % specificity on a human‑reviewed sample of 1,224 PHI tokens, surpassing the thresholds set by four institutional review boards. De‑identified records were stored as HIPAA Limited Data Sets, preserving a link to the original source for possible re‑identification under controlled circumstances (e.g., patient consent for trial enrollment).
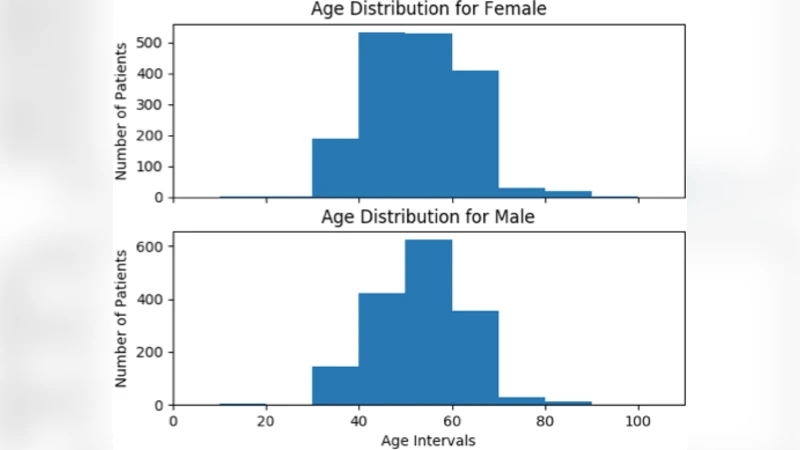
For cohort definition, the authors applied a standard RA case definition that incorporated SNOMED concepts (e.g., 156471009, 69896004) and 281 ICD‑10‑CM codes such as “Rheumatoid Arthritis with rheumatoid factor.” Patients were also grouped under the broader category “Diseases of the musculoskeletal system and connective tissue” to capture comorbid conditions relevant to trial eligibility. Demographic analysis revealed an average age of 53 years and a near‑equal gender distribution (52 % female, 48 % male). Disease prevalence profiling showed high rates of endocrine/metabolic disorders (61 %), circulatory system diseases (47 %), and musculoskeletal disorders (36 %). The prevalence of RA within this ACO population was approximately 4 %, notably higher than the <1 % prevalence reported in the general U.S. population, underscoring the value of aggregating data across multiple primary‑care sites.
The discussion emphasizes that, despite near‑universal EHR adoption in the United States, data sharing across independent practices remains hampered by system heterogeneity and privacy regulations. The presented pipeline—comprising non‑customized EHR extraction (via print, CCD, or API), robust standardization, and high‑accuracy de‑identification—offers a cost‑effective alternative to labor‑intensive manual chart review and fax‑based record exchange. By enabling rapid, reproducible cohort construction, the approach can shorten the site‑selection and patient‑recruitment phases of clinical trials, especially for conditions with low prevalence such as RA. Limitations include the modest sample size and the lack of a gold‑standard clinical validation of the RA diagnoses beyond code‑based definitions. Future work should scale the methodology to larger, multi‑institutional datasets and integrate prospective trial enrollment workflows.
In conclusion, the study demonstrates that routinely collected EHR data, when coupled with automated standardization and de‑identification, can substantially improve the efficiency and feasibility of clinical trial support in ACO settings. This strategy promises reduced trial costs, accelerated timelines, and broader inclusion of diverse patient populations, thereby enhancing the generalizability of trial outcomes.
Comments & Academic Discussion
Loading comments...
Leave a Comment