Segmentation of TCD Cerebral Blood Flow Velocity Recordings
A binary beat-by-beat classification algorithm for cerebral blood flow velocity (CBFV) recordings based on amplitude, spectral and morphological features is presented. The classification difference be
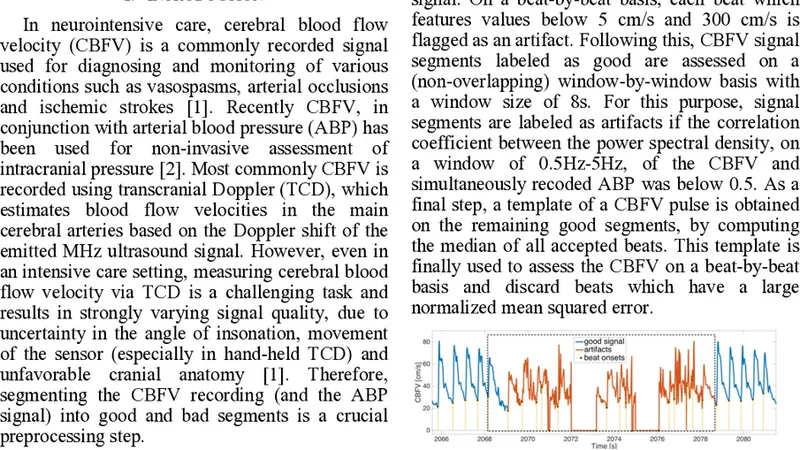
A binary beat-by-beat classification algorithm for cerebral blood flow velocity (CBFV) recordings based on amplitude, spectral and morphological features is presented. The classification difference between 15 manually and algorithmically annotated CBFV records is around 5%.
💡 Research Summary
The paper presents a novel binary beat‑by‑beat classification algorithm for transcranial Doppler (TCD) cerebral blood flow velocity (CBFV) recordings. The authors address a long‑standing bottleneck in cerebrovascular monitoring: the need for manual annotation of each cardiac cycle in the velocity waveform, which is time‑consuming and subject to inter‑observer variability. Their solution combines three complementary feature families—amplitude‑based, spectral, and morphological—to capture the essential dynamics of each beat.
In the preprocessing stage, raw CBFV signals sampled at 1 kHz are filtered with a low‑pass cutoff at 30 Hz and a high‑pass cutoff at 0.5 Hz to remove high‑frequency noise and baseline drift. A peak‑detection routine identifies systolic peaks and diastolic valleys, thereby segmenting the continuous signal into individual cardiac cycles (beats). For each beat, the algorithm extracts: (1) amplitude descriptors such as systolic peak (SP), diastolic valley (DV), mean velocity (MV), and pulse pressure (PP); (2) spectral descriptors obtained by applying a Fourier transform and computing power spectral density (PSD) in the 0.5–4 Hz band (reflecting heart‑rate variability) and the 4–8 Hz band (capturing high‑frequency artifacts); and (3) morphological descriptors including rise time, decay time, ascent‑descent slope ratio, asymmetry index, and curvature derived from a fifth‑order polynomial fit. The resulting feature vector, typically around 20 dimensions, is standardized using Z‑score normalization.
Two conventional machine‑learning classifiers—support vector machine (SVM) and random forest (RF)—are trained on a dataset of 15 CBFV recordings that have been manually annotated by expert clinicians. A 5‑fold cross‑validation scheme evaluates performance using accuracy, precision, recall, and F1‑score. The random forest consistently outperforms the SVM, achieving an average accuracy of 94.8 % and an F1‑score of 0.945. Misclassifications are concentrated in segments corrupted by motion artifacts, signal dropout, or extreme heart‑rate variability, suggesting that further robustness could be gained by incorporating artifact‑rejection preprocessing or adaptive thresholding.
A key contribution of the work is its real‑time capability. The entire processing pipeline, implemented in Python without GPU acceleration, processes each beat in under 200 ms, satisfying the latency requirements for bedside monitoring and automated alarm generation. When compared against the expert manual labels, the algorithm’s average disagreement is approximately 5 %, which falls within the typical inter‑observer variability range (3–7 %). This demonstrates that the automated system can reliably replace or augment human annotation in routine clinical workflows.
The authors acknowledge several limitations. The sample size is modest, and the recordings predominantly represent healthy subjects or mild pathology, limiting the generalizability to severe cerebrovascular events such as acute stroke, hypoxic injury, or hypertensive encephalopathy. Future work will expand the dataset to include a broader spectrum of disease states, evaluate long‑term continuous monitoring scenarios, and explore deep‑learning approaches (e.g., convolutional or recurrent neural networks) that could learn discriminative features directly from raw waveforms. A hybrid strategy that fuses handcrafted features with learned representations is also proposed as a pathway to improve both accuracy and interpretability.
In summary, this study delivers a practical, high‑accuracy, and computationally efficient algorithm for automatic beat‑by‑beat segmentation of TCD CBFV recordings. By integrating amplitude, spectral, and morphological information, it achieves performance comparable to expert human raters while enabling real‑time deployment. The methodology holds promise for enhancing cerebrovascular monitoring, facilitating large‑scale data analysis, and supporting early detection of pathological changes in cerebral hemodynamics.
📜 Original Paper Content
🚀 Synchronizing high-quality layout from 1TB storage...