Are My EHRs Private Enough? -Event-level Privacy Protection
Privacy is a major concern in sharing human subject data to researchers for secondary analyses. A simple binary consent (opt-in or not) may significantly reduce the amount of sharable data, since many patients might only be concerned about a few sensitive medical conditions rather than the entire medical records. We propose event-level privacy protection, and develop a feature ablation method to protect event-level privacy in electronic medical records. Using a list of 13 sensitive diagnoses, we evaluate the feasibility and the efficacy of the proposed method. As feature ablation progresses, the identifiability of a sensitive medical condition decreases with varying speeds on different diseases. We find that these sensitive diagnoses can be divided into 3 categories: (1) 5 diseases have fast declining identifiability (AUC below 0.6 with less than 400 features excluded); (2) 7 diseases with progressively declining identifiability (AUC below 0.7 with between 200 and 700 features excluded); and (3) 1 disease with slowly declining identifiability (AUC above 0.7 with 1000 features excluded). The fact that the majority (12 out of 13) of the sensitive diseases fall into the first two categories suggests the potential of the proposed feature ablation method as a solution for event-level record privacy protection.
💡 Research Summary
The paper addresses a nuanced privacy problem in the sharing of electronic health records (EHRs) for secondary research: patients often care about the confidentiality of specific medical events rather than their entire record. To meet this need, the authors propose “event‑level privacy protection,” a method that selectively removes information related to a particular sensitive diagnosis while preserving the rest of the record for research use.
A list of 13 clinically relevant sensitive diagnoses (including HIV, anxiety disorders, depressive disorder, various reproductive conditions, and newborn complications) serves as the testbed. For each diagnosis, a case cohort of up to 5,000 patients is assembled from the Northwestern Medicine Enterprise Data Warehouse (NMEDW), which aggregates data from two hospitals (Northwestern Memorial Hospital and Lake Forest Hospital) covering the period October 2010 to September 2015. A control cohort of 30,000 patients without any of the listed sensitive diagnoses is also constructed, with stratified sampling to match the distribution of total diagnosis counts and thus mitigate bias from differing comorbidity burdens.
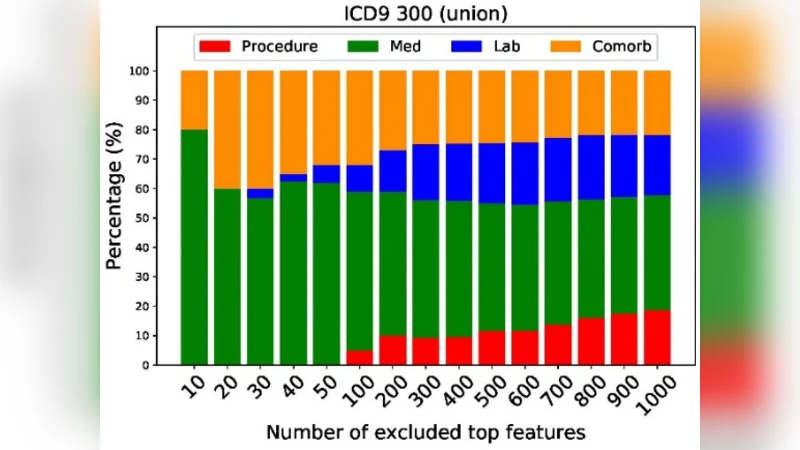
From each patient’s longitudinal record the authors extract three major data domains: laboratory test results, medication prescriptions, and procedural codes. All variables are binarized. Laboratory values are transformed into two binary flags (low vs. high) based on reference ranges; medications, procedures, and non‑sensitive diagnoses are represented by presence/absence flags. This yields several thousand binary features per cohort (e.g., 1,768 medication/procedure/comorbidity features for HIV).
The privacy risk is modeled as a supervised binary classification problem: can an attacker, given a mixed dataset of case and control patients, correctly identify which patients belong to a specific sensitive disease cohort? Logistic regression with L2 regularization is used as the classifier because of its interpretability and widespread adoption. Model training follows a 1:9 case‑to‑control split (≈500 cases, ≈4,500 controls) and stratified cross‑validation, reflecting a realistic scenario where an attacker has limited labeled data. Performance is measured by the area under the ROC curve (AUC).
To protect privacy, the authors introduce a feature‑ablation procedure. For each binary feature they compute a univariate relevance score using either the chi‑square statistic or the ANOVA F‑test, which quantifies the association between the feature and the disease label. Features are ranked by descending score, and the top k features are iteratively removed (k = 10, 20, …, 1000). After each removal step the logistic model is retrained and the AUC recomputed. This simulates the effect of progressively “scrubbing” the most predictive information from the released dataset.
The empirical results reveal three distinct patterns across the 13 diseases:
-
Fast‑declining identifiability (5 diseases). Removing fewer than 400 top features drives the AUC below 0.6, indicating that a relatively small set of highly predictive variables (often specific medications or procedures) is sufficient for identification.
-
Progressively declining identifiability (7 diseases). AUC falls below 0.7 only after eliminating between 200 and 700 top features, suggesting that identification relies on a broader, more distributed set of signals.
-
Slow‑declining identifiability (1 disease, HIV). Even after discarding 1,000 top features, the AUC remains above 0.7, reflecting the strong indirect signals (e.g., antiretroviral prescriptions, related lab abnormalities) that persist despite direct diagnosis removal.
Overall, 12 of the 13 conditions fall into the first two categories, demonstrating that targeted feature ablation can substantially reduce the risk of re‑identifying a patient’s sensitive condition while still leaving a useful dataset for secondary analysis.
The study contributes a practical, data‑driven framework for event‑level privacy that complements existing de‑identification standards (e.g., HIPAA). It highlights the importance of disease‑specific risk assessment and suggests that a modest amount of feature suppression can achieve a favorable privacy‑utility trade‑off for most conditions. Limitations include reliance on binary encoding (potential loss of granularity), confinement to two hospitals (raising questions about generalizability), and the possibility that residual indirect cues may still enable sophisticated attacks. Future work could explore multi‑institutional validation, hybrid approaches combining differential privacy noise with feature ablation, and adaptive algorithms that tailor the ablation depth to a user‑specified privacy budget.
Comments & Academic Discussion
Loading comments...
Leave a Comment