Identification of Key Proteins Involved in Axon Guidance Related Disorders: A Systems Biology Approach
Axon guidance is a crucial process for growth of the central and peripheral nervous systems. In this study, 3 axon guidance related disorders, namely- Duane Retraction Syndrome (DRS) , Horizontal Gaze Palsy with Progressive Scoliosis (HGPPS) and Congenital fibrosis of the extraocular muscles type 3 (CFEOM3) were studied using various Systems Biology tools to identify the genes and proteins involved with them to get a better idea about the underlying molecular mechanisms including the regulatory mechanisms. Based on the analyses carried out, 7 significant modules have been identified from the PPI network. Five pathways/processes have been found to be significantly associated with DRS, HGPPS and CFEOM3 associated genes. From the PPI network, 3 have been identified as hub proteins- DRD2, UBC and CUL3.
💡 Research Summary
The present study employs a comprehensive systems‑biology workflow to uncover the molecular underpinnings of three human axon‑guidance disorders: Duane Retraction Syndrome (DRS), Horizontal Gaze Palsy with Progressive Scoliosis (HGPPS), and Congenital Fibrosis of the Extraocular Muscles type 3 (CFEOM3). First, disease‑associated genes were retrieved from three public repositories—GeneCards (v3.0), STITCH (v5.0), and the Comparative Toxicogenomics Database (CTD). After merging and de‑duplicating the results, a curated list of 76 candidate genes was assembled (Table 1).
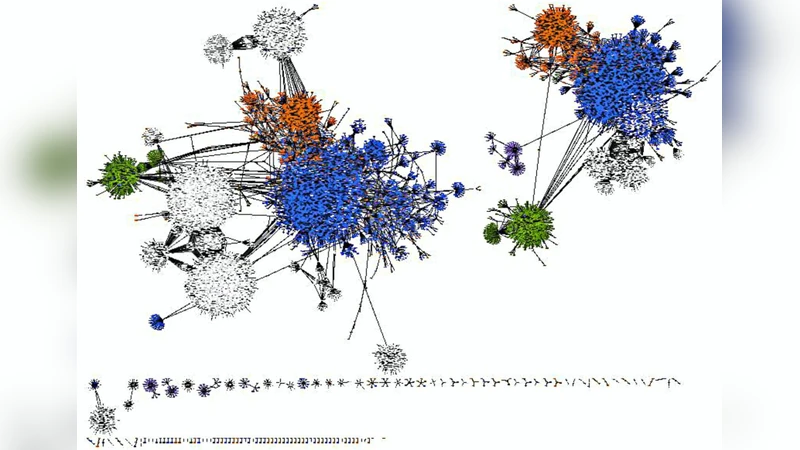
Next, protein‑protein interactions (PPIs) for these genes were extracted from two curated interaction databases, BioGRID (v3.4) and MINT (2012). Integration of the two sources yielded a large network comprising 24,062 nodes (proteins) and 29,226 edges (interactions), which was visualized in Cytoscape. Centrality analysis was performed with the CytoNCA plug‑in, calculating degree, betweenness, and closeness for each node. The three proteins that consistently ranked highest across all three metrics were the dopamine D2 receptor (DRD2), the poly‑ubiquitin gene (UBC), and Cullin‑3 (CUL3). These were designated as hub proteins (Table 2).
To explore functional sub‑structures, the MCODE algorithm was applied (degree cut‑off = 2, max depth = 100, node score cut‑off = 0.2, k‑core = 2). Seven highly interconnected modules were identified (Table 3). Each module represents a biologically coherent cluster: for example, Module 1 is enriched for chaperonin and cytoskeletal transport proteins (ASB‑3, CCT2, etc.), Module 2 contains ubiquitin‑proteasome components (ARRB2, GRB2, HGS, UBC), and Module 3 groups transcription‑regulatory factors (CUL3, DAPK1, etc.).
Pathway enrichment of the module proteins was carried out using the JEPETTO plug‑in for KEGG analysis. Five pathways emerged as statistically significant: dorso‑ventral axis formation, phototransduction, olfactory transduction, bladder cancer, and nucleotide excision repair (Table 4). The dorso‑ventral axis formation pathway is particularly relevant because it governs early neural tube patterning and the establishment of axonal trajectories, directly linking the identified network to the phenotypes of DRS, HGPPS, and CFEOM3.
Finally, transcriptional regulators were identified through the TRRUST database. Ten transcription factors—CIITA, TFAP4, RFX5, SOX2, POU5F1, REST, GATA3, STAT3, JUN, and EP300—were found to target the disease‑associated genes (Table 5). Several of these, such as SOX2 and POU5F1, are well‑known stem‑cell and neuro‑progenitor regulators, suggesting that dysregulation of early developmental transcription programs may contribute to the observed axon‑guidance defects.
The discussion contextualizes the three hub proteins within existing neurobiology literature. DRD2 is a classic dopamine receptor implicated in a wide spectrum of neuropsychiatric and movement disorders, indicating that dopaminergic signaling may intersect with axon‑guidance pathways. UBC maintains cellular ubiquitin pools, a critical factor for protein quality control under stress; its centrality hints at a broader role for ubiquitin homeostasis in neuronal survival and axonal maintenance. CUL3 functions as the scaffold of an E3 ubiquitin‑ligase complex, and prior work has shown its involvement in axonal arborization and dendritic elaboration, making it a plausible regulator of the structural remodeling required for proper eye‑movement control.
In conclusion, by integrating gene‑disease mining, large‑scale PPI mapping, centrality and module detection, pathway enrichment, and transcription‑factor analysis, the authors construct a detailed molecular network that links DRS, HGPPS, and CFEOM3. The identification of DRD2, UBC, and CUL3 as hub proteins provides concrete targets for future functional validation, drug‑screening, and possibly biomarker development. This systems‑level perspective not only clarifies shared pathogenic mechanisms across the three disorders but also establishes a methodological framework applicable to other neurodevelopmental conditions.
Comments & Academic Discussion
Loading comments...
Leave a Comment