A reaction network scheme which implements the EM algorithm
A detailed algorithmic explanation is required for how a network of chemical reactions can generate the sophisticated behavior displayed by living cells. Though several previous works have shown that reaction networks are computationally universal and can in principle implement any algorithm, there is scope for constructions that map well onto biological reality, make efficient use of the computational potential of the native dynamics of reaction networks, and make contact with statistical mechanics. We describe a new reaction network scheme for solving a large class of statistical problems including the problem of how a cell would infer its environment from receptor-ligand bindings. Specifically we show how reaction networks can implement information projection, and consequently a generalized Expectation-Maximization algorithm, to solve maximum likelihood estimation problems in partially-observed exponential families on categorical data. Our scheme can be thought of as an algorithmic interpretation of E. T. Jaynes’s vision of statistical mechanics as statistical inference.
💡 Research Summary
The paper presents a novel construction that embeds the Expectation‑Maximization (EM) algorithm into the continuous‑time dynamics of a chemical reaction network (CRN). Starting from the information‑geometric view of EM, the authors reinterpret each EM iteration as a pair of projections: an E‑projection (information projection) that minimizes the extended Kullback‑Leibler (KL) divergence D(x‖y) over a constrained affine subspace X, and an M‑projection (mixture projection) that minimizes the same divergence over a log‑convex exponential family Y. Alternating these projections yields the classical EM update.
To realize this mathematically in a physical system, the authors design a CRN composed of two interacting subnetworks. The species set S is partitioned into latent variables X (representing the hidden data) and parameters θ (the model parameters to be inferred). A finite set of integer vectors B defines elementary reactions of the form b⁺ → b⁻, where b⁺ and b⁻ are the positive and negative parts of each vector. The mass‑action kinetics of these reactions generate ordinary differential equations (ODEs) for the concentrations of X and θ.
The E‑projection subnetwork changes only the X‑species while using θ‑species as catalysts. With θ held fixed, the ODEs drive the system toward the point that minimizes D(x‖y(θ)) subject to the linear observation constraints (e.g., s = Sx). The M‑projection subnetwork does the opposite: it alters only the θ‑species, catalyzed by X, thereby minimizing D(x‖y(θ)) with respect to θ while keeping x fixed. Both subnetworks are constructed to possess a detailed‑balance point q, which ensures that the KL‑divergence acts as a Lyapunov function: dD/dt ≤ 0, with equality only at detailed balance.
A crucial technical contribution is the avoidance of critical siphons—subsets of species that, once depleted, cannot be regenerated—by careful choice of reaction vectors. This guarantees that the Global Attractor Conjecture holds, i.e., every trajectory converges to the unique detailed‑balance point within its stoichiometric compatibility class. Consequently, the combined CRN implements a continuous‑time EM algorithm: the pair (x(t),θ(t)) evolves such that D(x(t)‖y(θ(t))) monotonically decreases, and the trajectory converges to a local minimum (x*,θ*) that satisfies the observation constraints.
The authors substantiate their construction with rigorous theorems. Theorem 7 establishes that for any detailed‑balanced CRN without critical siphons, the mass‑action dynamics decrease D(x‖y) and converge to the E‑projection. Theorem 10 and Theorem 11 prove analogous results for the M‑projection subnetwork, introducing a new class of nonlinear CRNs whose Lyapunov functions have a time‑dependent second argument—a novelty beyond existing results that only handle static target points. Theorem 12 ties the two subnetworks together, showing that the full system’s dynamics satisfy dD/dt ≤ 0 and therefore converge to a stationary point of the EM objective.
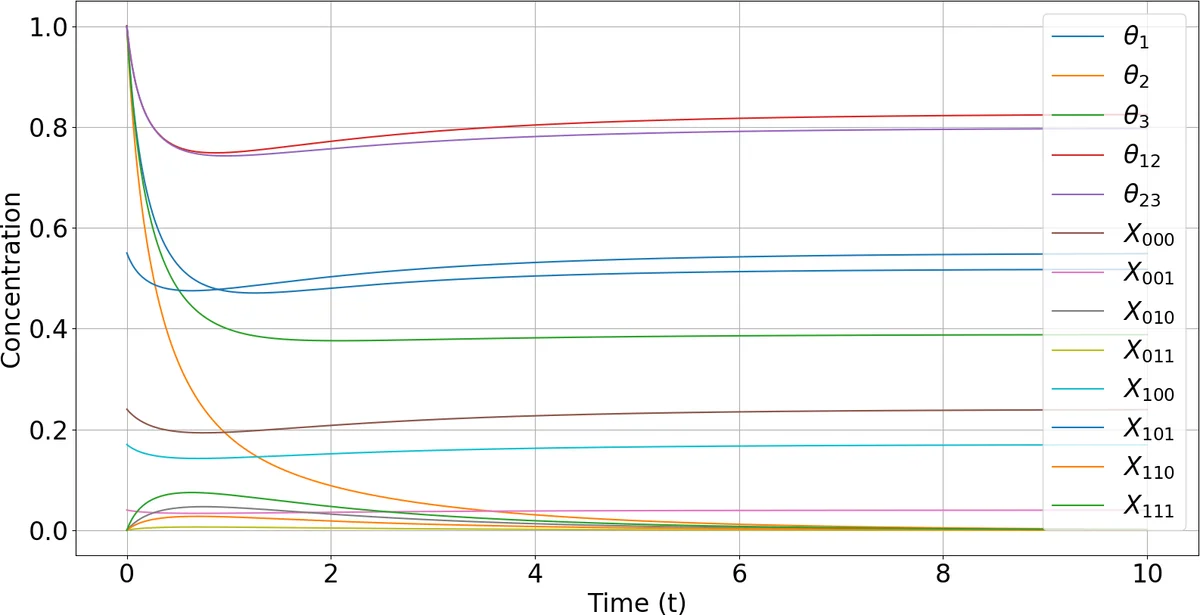
An illustrative example uses a three‑sided die whose outcome probabilities depend on two hidden parameters. The authors write out eight reactions that implement the EM steps for this model, demonstrating how the concentrations of X₁, X₂, X₃ (latent counts) and θ₁, θ₂ (parameters) evolve under mass‑action kinetics to recover the maximum‑likelihood estimates from only aggregated observations (total rolls and the number of rolls yielding either side X₁ or X₂).
Beyond the algorithmic novelty, the paper emphasizes biological plausibility: all reactions are of the simple creation‑annihilation or catalytic type, mirroring real biochemical processes. The mapping between free energy in physics and KL‑divergence in information theory provides a unified thermodynamic interpretation of statistical inference. In this view, a cell could perform Bayesian updating of its environmental model simply by the natural relaxation of its internal reaction network.
In summary, the work bridges three domains—chemical reaction network theory, information geometry, and statistical learning—by showing that a carefully engineered CRN can autonomously execute the EM algorithm. This opens avenues for synthetic biology implementations of sophisticated inference mechanisms, for molecular computing platforms that exploit native chemical dynamics, and for deeper theoretical insights into how physical systems embody statistical reasoning.
Comments & Academic Discussion
Loading comments...
Leave a Comment