Ultrafast imaging of cell elasticity with optical microelastography
Elasticity is a fundamental cellular property that is related to the anatomy, functionality and pathological state of cells and tissues. However, current techniques based on cell deformation, atomic force microscopy or Brillouin scattering are rather slow and do not always accurately represent cell elasticity. Here, we have developed an alternative technique by applying shear wave elastography to the micrometer scale. Elastic waves were mechanically induced in live mammalian oocytes using a vibrating micropipette. These audible frequency waves were observed optically at 200,000 frames per second and tracked with an optical flow algorithm. Whole cell elasticity was then mapped using an elastography method inspired by the seismology field. Using this approach, we show that the elasticity of mouse oocyte is decreased when the oocyte cytoskeleton is disrupted with cytochalasin B. The technique is fast (less than 1 ms for data acquisition), precise (spatial resolution of a few micrometers), able to map internal cell structures, robust, and thus represents a tractable novel option for interrogating biomechanical properties of diverse cell types.
💡 Research Summary
This paper introduces a novel method for measuring cellular elasticity called optical micro‑elastography, which adapts shear‑wave elastography concepts from seismology to the micrometer scale. The authors generate high‑frequency shear waves inside live mouse oocytes by contacting the cell with a vibrating micropipette that oscillates at 15 kHz with a 20 µm amplitude. The resulting wave propagation is captured with an ultra‑high‑speed camera operating at 200,000 frames per second through a 100× microscope objective. An optical‑flow algorithm extracts pixel‑wise displacement fields from the image sequence, and a “passive” elastography algorithm—originally developed for passive seismic imaging—calculates the shear‑wave speed (cₛ). Assuming a known cell density, the shear modulus G is obtained directly from G = ρ cₛ², providing absolute elasticity values in kilopascals without requiring any stress‑distribution model.
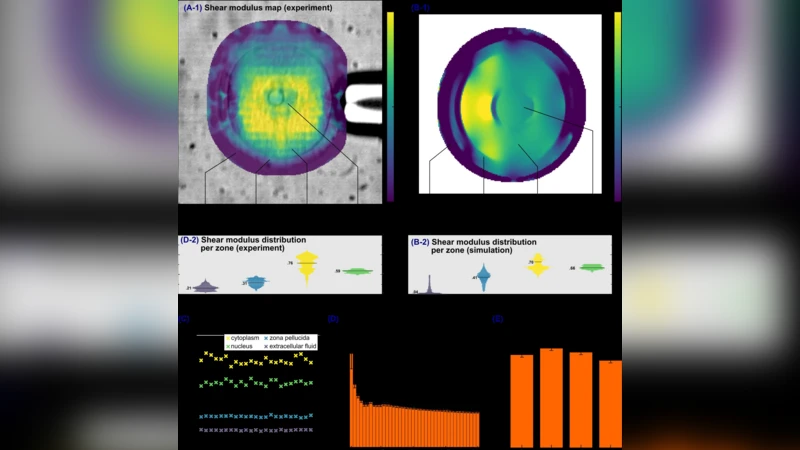
The technique achieves sub‑millisecond acquisition: reliable elasticity maps are produced from as few as 5 frames (0.025 ms) but stabilize after about 160 frames (≈0.8 ms). Spatial resolution is on the order of ten micrometers, sufficient to distinguish the zona pellucida, cytoplasm, nucleus, and surrounding fluid in an 80 µm‑diameter oocyte. Median shear moduli measured were 0.31 kPa (zona pellucida), 0.76 kPa (cytoplasm), 0.59 kPa (nucleus), and a small artifact (~0.21 kPa) in the extracellular fluid, attributed to spurious displacement detection. Reproducibility was demonstrated by 23 consecutive measurements taken 2.5 ms apart, showing no temporal drift in any region.
Robustness tests revealed that varying the vibration amplitude between 8 µm and 20 µm did not affect the estimated modulus, indicating that precise control of the excitation amplitude is not critical. The authors also examined the effect of actin disruption using cytochalasin B. Confocal imaging confirmed a marked reduction in cortical actin, and optical micro‑elastography detected a statistically significant softening of the whole cell (p < 0.002), with decreases observed in both cytoplasm and nucleus. Measurements across developmental stages (germinal vesicle, 2‑cell, 4‑cell embryos) showed comparable elasticity, suggesting that the method can be applied to cells at different maturation states.
Compared with existing approaches—static deformation assays (micropipette aspiration, magnetic‑bead twisting, optical tweezers), atomic‑force microscopy, particle‑tracking microrheology, and Brillouin scattering—the new method offers several distinct advantages. It is non‑contact, non‑invasive, and provides absolute, localized elasticity values in a single acquisition lasting less than a millisecond, thereby eliminating confounding biological changes that can occur during longer measurements. Unlike AFM, it can probe internal structures rather than just the surface, and unlike Brillouin microscopy, it measures shear rather than bulk modulus, which is more sensitive to cytoskeletal alterations. The simplicity of the setup—standard inverted microscope, two micropipettes, and a high‑speed camera—makes it accessible to many laboratories.
Limitations include the current spatial resolution (~10 µm), which may be insufficient for smaller cells or sub‑cellular organelles. Higher excitation frequencies (>30 kHz) and faster imaging would be needed to resolve finer features. Additionally, the method assumes a purely elastic, isotropic medium; viscoelastic effects and mixed wave modes (surface vs. bulk) are not fully disentangled. Nonetheless, the authors propose extensions such as multi‑frequency excitation, three‑dimensional mapping by varying focal depth, and the use of phase‑contrast imaging to improve particle tracking.
In summary, optical micro‑elastography provides a rapid, precise, and versatile platform for quantifying cellular elasticity at the micrometer scale. Its ability to generate whole‑cell elasticity maps in sub‑millisecond timescales opens the door to real‑time monitoring of dynamic cellular processes, drug responses, and disease‑related mechanical alterations. Future work aimed at improving resolution, incorporating viscosity measurements, and validating against gold‑standard techniques will further solidify its role in cellular biomechanics research.
Comments & Academic Discussion
Loading comments...
Leave a Comment