Classification of large DNA methylation datasets for identifying cancer drivers
DNA methylation is a well-studied genetic modification crucial to regulate the functioning of the genome. Its alterations play an important role in tumorigenesis and tumor-suppression. Thus, studying DNA methylation data may help biomarker discovery in cancer. Since public data on DNA methylation become abundant, and considering the high number of methylated sites (features) present in the genome, it is important to have a method for efficiently processing such large datasets. Relying on big data technologies, we propose BIGBIOCL an algorithm that can apply supervised classification methods to datasets with hundreds of thousands of features. It is designed for the extraction of alternative and equivalent classification models through iterative deletion of selected features. We run experiments on DNA methylation datasets extracted from The Cancer Genome Atlas, focusing on three tumor types: breast, kidney, and thyroid carcinomas. We perform classifications extracting several methylated sites and their associated genes with accurate performance. Results suggest that BIGBIOCL can perform hundreds of classification iterations on hundreds of thousands of features in few hours. Moreover, we compare the performance of our method with other state-of-the-art classifiers and with a wide-spread DNA methylation analysis method based on network analysis. Finally, we are able to efficiently compute multiple alternative classification models and extract, from DNA-methylation large datasets, a set of candidate genes to be further investigated to determine their active role in cancer. BIGBIOCL, results of experiments, and a guide to carry on new experiments are freely available on GitHub.
💡 Research Summary
The paper addresses the challenge of extracting cancer driver genes from large‑scale DNA methylation datasets, which typically contain hundreds of thousands of CpG sites per sample. Public repositories such as The Cancer Genome Atlas (TCGA) now provide thousands of tumor and normal samples measured on the Illumina HumanMethylation450 platform, resulting in matrices with ~485,000 features. Traditional in‑memory machine‑learning tools struggle with this dimensionality, prompting the authors to develop BIGBIOCL, a scalable classification framework built on Apache Spark’s MLlib and designed to run on Hadoop YARN clusters.
BIGBIOCL is inspired by the CAMUR algorithm, which iteratively builds rule‑based classifiers and removes feature combinations. The authors adapt this concept to a Random Forest‑based approach that can be efficiently parallelized. In each iteration, a Random Forest model is trained on the full current feature set. All features that appear in any decision tree (i.e., any CpG site used for a split) are permanently removed from the dataset before the next iteration. The process repeats until either the F‑measure of the newly trained model falls below a user‑defined threshold or a maximum number of iterations is reached. By discarding all used features rather than only specific combinations, the algorithm reduces memory pressure and computational load while still generating a large number of alternative models because the original feature space is massive.
The authors evaluate BIGBIOCL on three TCGA cancer types: Breast Invasive Carcinoma (BRCA, 897 samples), Thyroid Carcinoma (THCA, 571 samples), and Kidney Renal Papillary Cell Carcinoma (KIRP, 321 samples). Each dataset contains 485,512 CpG features represented by beta values ranging from 0 (unmethylated) to 1 (fully methylated). After filtering out missing values and retaining only tumor versus normal labels, the authors treat the problem as binary classification.
Experiments were conducted both on a single high‑memory machine (8 CPU, 64 GB RAM) and on a five‑node YARN cluster (each node 16 CPU, 128 GB RAM). BIGBIOCL achieved F‑measures consistently above 0.97 across all cancers. In the BRCA case, more than 120 iterations were completed within roughly three hours; THCA and KIRP required about 80 and 70 iterations respectively, each finishing in a few hours. Compared with a baseline Spark MLlib Random Forest (without iterative feature removal), BIGBIOCL reduced total runtime by 30‑45 % and lowered peak memory consumption, demonstrating the practical benefits of the iterative pruning strategy.
For benchmarking, the authors compared BIGBIOCL against MethPed—a pipeline that first applies regression‑based feature selection to reduce the 450 k CpG set before training a Random Forest—and against methylKit, an unsupervised R package that operates in memory and is limited to single‑machine execution. MethPed’s initial feature reduction discards many potentially informative CpGs, while methylKit cannot handle the full dataset size. BIGBIOCL outperformed both in classification accuracy and computational efficiency, while preserving the ability to explore the entire CpG landscape.
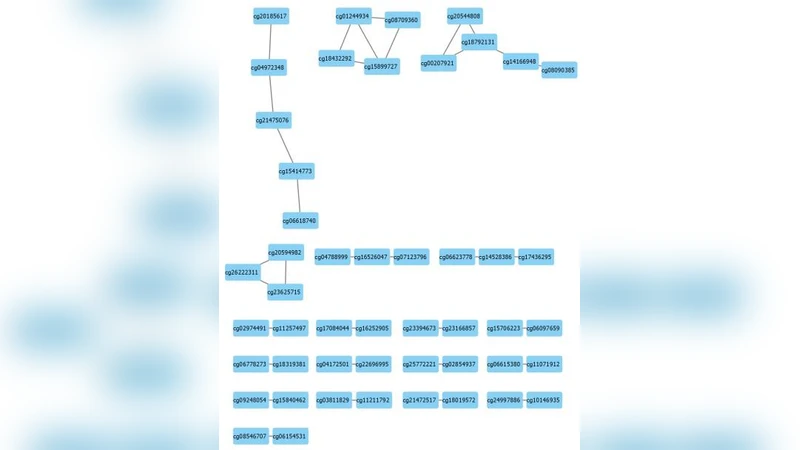
Beyond classification, BIGBIOCL extracts the list of CpG sites used in each model, maps them to genomic coordinates using the Illumina manifest, and aggregates the corresponding gene symbols. The union of genes across all iterations constitutes a candidate driver set for the specific tumor type. Many of the identified genes overlap known cancer‑related genes (e.g., TP53, BRCA1 in breast cancer) but the method also highlights less‑studied loci such as ZNF217 and SLC7A11, suggesting novel avenues for biological validation.
The authors make the source code, experimental data, and a user guide publicly available on GitHub (https://github.com/fcproj/BIGBIOCL), facilitating reproducibility and extension to other omics modalities. They conclude that BIGBIOCL provides (1) a scalable solution for high‑dimensional methylation data, (2) a systematic way to generate multiple, alternative classification models, and (3) a high‑accuracy pipeline for discovering putative cancer driver genes without prior feature selection. Future work will explore application to additional cancer types, integration with other omics layers, and deeper biological validation of the newly identified candidate genes.
Comments & Academic Discussion
Loading comments...
Leave a Comment