A Determination Scheme for Quasi-Identifiers Using Uniqueness and Influence for De-Identification of Clinical Data
Objectives; The accumulation and usefulness of clinical data have increased with IT development. While using clinical data that needs to be identifiable to obtain meaningful information, it is essential to ensure that data is de-identified and unnecessary clinical information is minimized to protect personal information. This process requires criteria and an appropriate method as there are clear identifiers as well as quasi-identifiers that are not readily identifiable. Methods; To formulate such a method, first, primary quasi-identifiers were selected by classifying information in 20 clinical personal information database tables into Direct-Identifier (DID), Quasi-Identifier (QI), Sensitive Attribute (SA), and Non-Sensitive Attribute (NSA) according to its type. Secondary QIs were then selected by assessing the risk for outliers by measuring uniqueness values of the selected data and scoring re-identification by calculating equivalence class of the influence on other data on QI removal. Third, the risk of re-identification of data users was numeralized and classified. Lastly, the final QI according to user class was determined by comparing the calculated re-identification scores to the threshold values of user classes. Results; Eventually, final QIs ranging from a minimum of 18 to a maximum of 28 were selected by making an assumption about user classes and using it as criteria. Conclusions; The QI selection method presented by the current investigators can be used by researchers at the final checkup stage before they de-identify the selected QIs. Therefore, clinical data users can securely and efficiently use clinical data containing personal information by objectively selecting QIs using the method proposed in the present study.
💡 Research Summary
The paper addresses a critical gap in clinical data de‑identification: the systematic selection of quasi‑identifiers (QIs) that balance re‑identification risk against data utility. While direct identifiers (DIDs) such as names or social security numbers are unambiguously removed under regulations like HIPAA, QIs are more ambiguous; they do not identify an individual on their own but can do so when combined with other attributes. Existing practice often relies on expert intuition, leading to inconsistent QI sets and either excessive data loss or insufficient privacy protection.
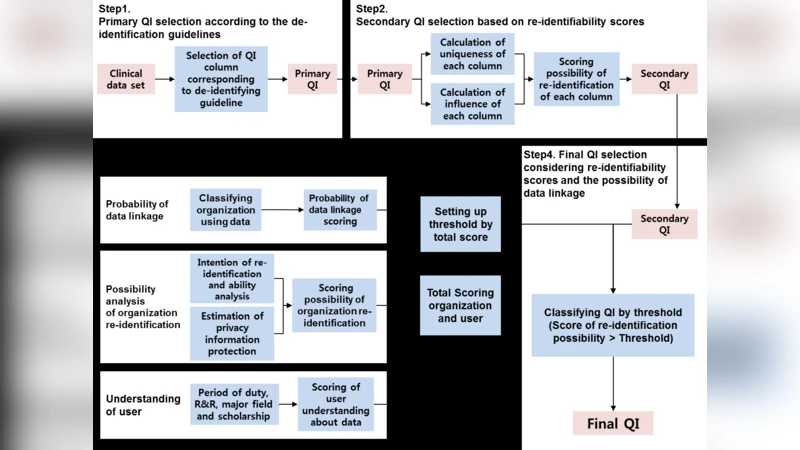
To resolve this, the authors propose a multi‑step, quantitative framework and apply it to the electronic medical record (EMR) system of Korea’s National Cancer Center. The dataset comprises 20 clinical tables, of which 17 contain personal information and three additional tables hold death‑related data. First, each column is classified into one of four categories: Direct Identifier (DID), Quasi‑Identifier (QI), Sensitive Attribute (SA), or Non‑Sensitive Attribute (NSA), following HIPAA guidance and expert review. This yields an initial pool of primary QIs.
The second step introduces two novel metrics for each primary QI:
-
Uniqueness – the proportion of records that have a unique value in that column. A higher uniqueness ratio indicates that the column alone can single out individuals, raising re‑identification risk.
-
Influence – the change in the number of equivalence classes (i.e., distinct QI value combinations) when the column is removed. If the total number of equivalence classes drops sharply, the column exerts strong influence on other attributes, meaning it can amplify re‑identification when combined with them.
The authors compute both metrics for every candidate column, sum them to obtain a “re‑identifiability score,” and select those with the highest scores as secondary QIs.
The third step incorporates contextual risk by grading both the data‑receiving institution and the individual researcher. Institutions are stratified into high, medium, and low categories based on their typical access to clinical data (e.g., national health insurance agencies receive a high grade, while general government agencies receive a low grade). Corresponding weights of 10, 5, and 1 are assigned. Researchers are evaluated on four binary questions concerning intent and capability to re‑identify (e.g., potential harm to subjects, monetary gain, contractual prohibitions, and ability to link external data). Each “yes” answer adds one point, yielding a maximum researcher score of four.
Privacy protection measures are also scored via six binary items (e.g., existence of security service contracts, staff training, management plans). Here, each “no” answer adds one point, with a maximum of six, indicating weaker protection.
All these scores are aggregated and compared against pre‑defined thresholds that reflect acceptable risk levels for each user‑institution combination. The final QI set is then determined: depending on the assumed user class, the authors report a range of 18 to 28 columns being retained as QIs that must be de‑identified.
The results demonstrate that the framework can produce a transparent, reproducible QI selection process that reduces unnecessary data suppression while keeping re‑identification risk within acceptable bounds. The authors argue that this method can serve as a “final checklist” for researchers before de‑identifying data, ensuring consistent application across projects.
In the discussion, the authors acknowledge several limitations. The uniqueness and influence metrics are column‑centric and may not capture complex multivariate interactions that could also lead to re‑identification. The weighting scheme for institutions and researchers is based on expert judgment and may need adaptation for other health systems or regulatory environments. Moreover, the study is confined to a single national cancer center’s EMR, so external validation on heterogeneous datasets is required.
Nevertheless, the contribution is significant: by quantifying QI risk and integrating policy‑driven user grading, the paper offers a pragmatic tool for data custodians, privacy officers, and investigators seeking to balance data utility with privacy compliance in clinical research.
Comments & Academic Discussion
Loading comments...
Leave a Comment