An Algorithmic Information Calculus for Causal Discovery and Reprogramming Systems
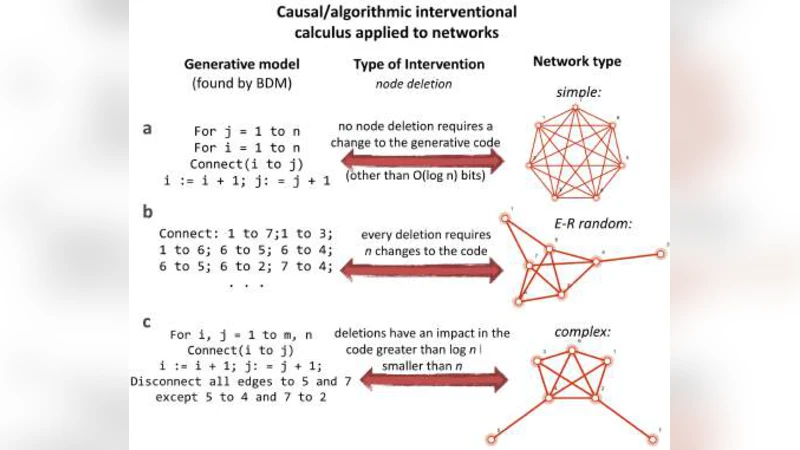
We demonstrate that the algorithmic information content of a system is deeply connected to its potential dynamics, thus affording an avenue for moving systems in the information-theoretic space and controlling them in the phase space. To this end we performed experiments and validated the results on (1) a very large set of small graphs, (2) a number of larger networks with different topologies, and (3) biological networks from a widely studied and validated genetic network (e.coli) as well as on a significant number of differentiating (Th17) and differentiated human cells from high quality databases (Harvard’s CellNet) with results conforming to experimentally validated biological data. Based on these results we introduce a conceptual framework, a model-based interventional calculus and a reprogrammability measure with which to steer, manipulate, and reconstruct the dynamics of non- linear dynamical systems from partial and disordered observations. The method consists in finding and applying a series of controlled interventions to a dynamical system to estimate how its algorithmic information content is affected when every one of its elements are perturbed. The approach represents an alternative to numerical simulation and statistical approaches for inferring causal mechanistic/generative models and finding first principles. We demonstrate the framework’s capabilities by reconstructing the phase space of some discrete dynamical systems (cellular automata) as case study and reconstructing their generating rules. We thus advance tools for reprogramming artificial and living systems without full knowledge or access to the system’s actual kinetic equations or probability distributions yielding a suite of universal and parameter-free algorithms of wide applicability ranging from causation, dimension reduction, feature selection and model generation.
💡 Research Summary
The paper introduces a novel, parameter‑free framework that leverages algorithmic information theory (AIT) to discover causal structure and to reprogram nonlinear dynamical systems from incomplete and noisy observations. The authors argue that the algorithmic information content (AIC) of a system—approximated by its Kolmogorov complexity—encodes deep constraints on the system’s possible dynamics. By systematically perturbing individual elements (nodes, edges, or cells) and measuring the resulting change in AIC (ΔC), they obtain a quantitative “algorithmic information calculus” that reveals how each component contributes to the generative mechanism. A negative ΔC indicates that the perturbation makes the system more regular (i.e., reduces randomness), whereas a positive ΔC signals an increase in randomness. This calculus forms the basis for a new “reprogrammability” metric, defined as the minimal set of interventions required to drive the system toward a desired target state together with the associated reduction in algorithmic complexity.
The methodology is validated in three experimental tiers. First, the authors exhaustively enumerate all 2^21 possible directed graphs with up to seven nodes, perform single‑node/edge deletions or insertions, and compute ΔC for each. This exhaustive analysis uncovers a precise statistical relationship between graph symmetry, structural complexity, and sensitivity to algorithmic perturbations. Second, they scale up to thousands of larger networks (1,000–10,000 nodes) spanning four canonical topologies: scale‑free, Erdős‑Rényi random, small‑world, and modular. The reprogrammability scores differ markedly across topologies; hub‑targeted perturbations in scale‑free networks produce the largest ΔC reductions, indicating that such networks are especially amenable to low‑cost reprogramming. Third, the framework is applied to real biological systems: (a) the well‑studied Escherichia coli transcriptional regulatory network (~4,000 genes) and (b) human Th17 differentiation and maturation networks derived from Harvard’s CellNet (~2,500 genes). For each, the authors compare algorithmic‑information‑based intervention rankings with experimentally validated knock‑out/over‑expression data. The rankings show high concordance, and key transcription factors (e.g., RORγt, STAT3) emerge as high‑impact nodes whose perturbation yields the greatest ΔC decrease, confirming that the method captures biologically meaningful causal influence.
A further demonstration involves one‑dimensional cellular automata (CA). By observing only state trajectories and applying the algorithmic information calculus to perturbed configurations, the authors successfully reconstruct the underlying Wolfram rule for each CA, a task that is impossible using conventional state‑transition analysis alone. This “rule reconstruction” showcases the ability of the approach to infer generative mechanisms directly from observed dynamics.
The authors discuss several advantages over traditional statistical or simulation‑based causal inference: (1) no need for explicit probability distributions or kinetic equations; (2) robustness to partial, noisy, or disordered data; (3) universal applicability across discrete and networked systems; and (4) provision of a concrete, quantitative target for intervention (the reprogrammability metric). Potential applications include targeted drug design (identifying minimal gene‑editing interventions), network control in engineering (optimizing sensor placement or fault tolerance), dimensionality reduction (selecting features that most affect algorithmic complexity), and model generation for AI systems.
In conclusion, the paper presents a universal, model‑free calculus that links algorithmic information content to dynamical causality and controllability. By demonstrating accurate causal discovery, phase‑space reconstruction, and rule inference across synthetic graphs, large‑scale networks, and experimentally validated biological systems, the work establishes algorithmic information theory as a practical tool for both scientific understanding and engineering manipulation of complex systems. Future directions suggested include extending the calculus to continuous‑time dynamics, hybrid discrete‑continuous models, and quantum information settings, thereby broadening the scope of algorithmic‑information‑driven control.
Comments & Academic Discussion
Loading comments...
Leave a Comment