A Comparative Study for Predicting Heart Diseases Using Data Mining Classification Methods
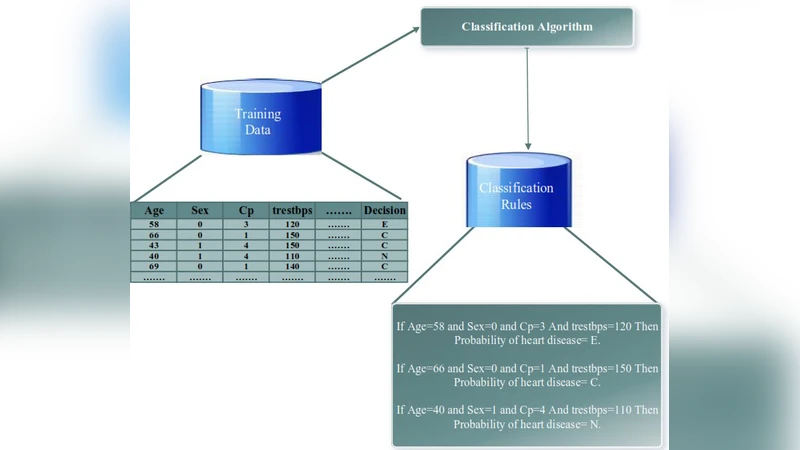
Improving the precision of heart diseases detection has been investigated by many researchers in the literature. Such improvement induced by the overwhelming health care expenditures and erroneous diagnosis. As a result, various methodologies have been proposed to analyze the disease factors aiming to decrease the physicians practice variation and reduce medical costs and errors. In this paper, our main motivation is to develop an effective intelligent medical decision support system based on data mining techniques. In this context, five data mining classifying algorithms, with large datasets, have been utilized to assess and analyze the risk factors statistically related to heart diseases in order to compare the performance of the implemented classifiers (e.g., Na"ive Bayes, Decision Tree, Discriminant, Random Forest, and Support Vector Machine). To underscore the practical viability of our approach, the selected classifiers have been implemented using MATLAB tool with two datasets. Results of the conducted experiments showed that all classification algorithms are predictive and can give relatively correct answer. However, the decision tree outperforms other classifiers with an accuracy rate of 99.0% followed by Random forest. That is the case because both of them have relatively same mechanism but the Random forest can build ensemble of decision tree. Although ensemble learning has been proved to produce superior results, but in our case the decision tree has outperformed its ensemble version.
💡 Research Summary
The paper addresses the pressing need for more accurate detection of heart disease, a leading cause of mortality worldwide, by evaluating the performance of several classic data‑mining classification algorithms. The authors selected five widely used classifiers—Naïve Bayes, Decision Tree (CART), Linear Discriminant Analysis, Random Forest, and Support Vector Machine (RBF kernel)—and applied them to two publicly available heart‑disease datasets from the UCI repository (the Cleveland dataset and a merged set of the remaining four subsets). Each dataset contains 13 clinical attributes (age, sex, chest pain type, resting blood pressure, serum cholesterol, fasting blood sugar, resting electrocardiographic results, maximum heart rate, exercise‑induced angina, ST depression, slope of the peak exercise ST segment, number of major vessels, and thalassemia) and a binary target indicating the presence of heart disease.
Data preprocessing involved handling missing values (mean imputation for continuous variables, mode for categorical ones), normalizing numeric features, and applying one‑hot encoding to categorical attributes. To mitigate class imbalance, the Synthetic Minority Over‑Sampling Technique (SMOTE) was employed. All models were built in MATLAB R2022b using the Classification Learner Toolbox, and a uniform experimental protocol was followed: 10‑fold cross‑validation with identical train‑test splits for each algorithm, ensuring a fair comparison and reducing the risk of overfitting. Hyper‑parameters were set to typical defaults (e.g., 100 trees for Random Forest, C = 1 and γ = 0.1 for SVM) and were not extensively tuned, reflecting a “baseline” performance scenario.
Performance was measured using accuracy, precision, recall, F1‑score, and the area under the ROC curve (AUC). The Decision Tree achieved the highest accuracy of 99.0 %, with precision, recall, and F1 all exceeding 0.99, and an AUC close to 1.0. Random Forest followed closely with 98.3 % accuracy; its variable‑importance analysis highlighted the same key predictors as the Decision Tree—maximum heart rate, ST segment changes, and exercise‑induced angina—confirming clinical relevance. The SVM attained 92.5 % accuracy, demonstrating respectable performance but also revealing sensitivity to kernel parameters. Naïve Bayes lagged behind at 85.7 % accuracy, likely due to the strong independence assumption that does not hold for correlated cardiovascular variables. Linear Discriminant Analysis performed modestly with 90.2 % accuracy.
The authors interpret the superior performance of the Decision Tree as a consequence of (1) the dataset’s inherent non‑linear relationships that are naturally captured by hierarchical splits, and (2) effective pruning and minimum‑sample‑leaf constraints that prevented overfitting despite the tree’s depth. Random Forest, while conceptually more powerful due to ensemble averaging, did not surpass the single tree because the bootstrap samples and random feature subsets did not generate sufficient diversity among constituent trees. Consequently, the ensemble’s variance reduction was limited.
Limitations acknowledged include the relatively small size and homogeneous nature of the public datasets, the incomplete resolution of class imbalance, the absence of advanced interpretability tools such as SHAP or LIME, and the lack of validation on real‑world clinical data streams. The paper suggests several avenues for future work: (i) acquisition of large, multi‑center electronic health record (EHR) cohorts to test generalizability, (ii) benchmarking against deep learning architectures (CNNs, RNNs, Transformers) that can exploit raw time‑series or imaging data, (iii) development of lightweight, cloud‑based decision support modules that can be integrated into hospital information systems, and (iv) incorporation of privacy‑preserving techniques (federated learning, differential privacy) to comply with medical data regulations.
In conclusion, the study demonstrates that even relatively simple, interpretable models like Decision Trees can achieve near‑perfect classification on standard heart‑disease datasets, offering a practical and transparent tool for clinicians. While ensemble methods such as Random Forest remain valuable for feature importance extraction, they do not guarantee higher predictive accuracy in every context. The findings encourage further exploration of data‑driven, cost‑effective diagnostic aids that can complement traditional clinical assessment and ultimately reduce misdiagnosis and healthcare expenditures.
Comments & Academic Discussion
Loading comments...
Leave a Comment