Mittens: An Extension of GloVe for Learning Domain-Specialized Representations
We present a simple extension of the GloVe representation learning model that begins with general-purpose representations and updates them based on data from a specialized domain. We show that the resulting representations can lead to faster learning and better results on a variety of tasks.
💡 Research Summary
The paper introduces “Mittens,” a straightforward extension of the GloVe word‑embedding model that enables domain‑specific fine‑tuning while preserving the information contained in large, general‑purpose pretrained vectors. Traditional GloVe learns word vectors wᵢ and context vectors eⱼ by minimizing a weighted least‑squares loss that forces the inner product wᵢ·eⱼ (plus bias terms) to approximate the logarithm of the co‑occurrence count Xᵢⱼ. The authors first rewrite this objective in a fully vectorized matrix form, J = f(X)·(Wᵀ·F·W + b·1ᵀ + 1·\tilde bᵀ − g(X)), which allows efficient implementation in NumPy and TensorFlow and yields GPU‑accelerated training speeds comparable to the original C implementation.
Mittens augments the vectorized GloVe loss with a retrofitting regularizer: for every word i for which an external pretrained embedding rᵢ exists (the set R), a penalty µ‖(wᵢ + eᵢ) − rᵢ‖² is added. The hyper‑parameter µ controls the trade‑off between (a) adapting to the new domain’s co‑occurrence statistics (the original GloVe term) and (b) staying close to the prior embedding (the regularizer). When µ = 0 the model reduces to vanilla GloVe; as µ grows, the learned vectors are forced to remain near the external vectors.
The authors conduct three families of experiments.
-
Sentiment (IMDB) domain – Using the unlabeled half of the IMDB movie‑review corpus, they build a co‑occurrence matrix (window = 10, distance‑weighted counts) and train 50‑dimensional embeddings. Three baselines are compared: (i) “External GloVe” (pretrained on Wikipedia + Gigaword), (ii) “IMDB GloVe” (trained from scratch on the IMDB data), and (iii) Mittens (starting from External GloVe and fine‑tuning on IMDB with µ = 0.1). Classification is performed by representing each review as the sum of its word vectors and feeding this to a Random Forest. Mittens achieves 77.4 % accuracy, outperforming External GloVe (62.0 %) and IMDB GloVe (72.2 %). Correlation analysis shows that Mittens’ vectors retain a higher Pearson ρ (≈ 0.47) between inner products and log co‑occurrences than vanilla GloVe (≈ 0.37), suggesting that the domain data smooths noisy statistics while preserving the original distributional structure.
-
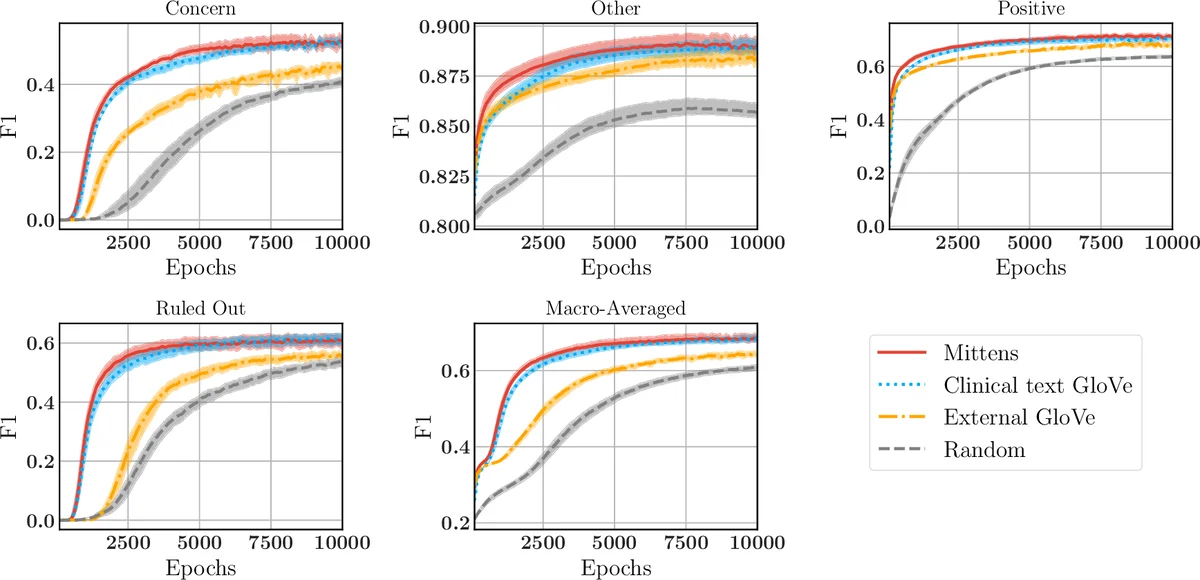
Clinical text – From 100 K de‑identified clinical notes (≈ 1.3 M “SOAP” segments), a 6,519‑word vocabulary is built (frequency threshold = 500). Mittens and GloVe are trained on the same co‑occurrence matrix (µ = 0.1 for Mittens). Both achieve similar ρ ≈ 0.51, indicating that the regularizer does not destroy the GloVe objective. To evaluate downstream utility, 3,206 sentences containing disease mentions are manually labeled into four categories (Positive diagnosis, Concern, Ruled Out, Other). A single‑layer LSTM (50‑dim hidden state) is trained, allowing the embeddings to be updated. Across ten random train/test splits, Mittens converges faster and reaches slightly higher macro‑F1 scores for all categories, confirming that a warm‑start accelerates learning in sequence‑labeling tasks.
-
Transfer to SNOMED CT edge prediction – The authors test whether Mittens’ embeddings generalize beyond the training domain. They select the five largest SNOMED CT semantic subgraphs (e.g., “disorder”, “procedure”), represent each node by summing the vectors of the words in its canonical name, and train a Random Forest to predict whether an edge exists between two nodes (balanced positive/negative set). Nodes in train and test splits are disjoint. Mittens consistently yields marginally higher accuracy (≈ 0.5–1 % absolute) than both Random (baseline) and External GloVe, demonstrating that the domain‑adapted vectors retain useful semantic information for downstream biomedical graph tasks.
Across all experiments, the key findings are: (a) the vectorized GloVe implementation enables efficient retrofitting; (b) a single scalar µ suffices to balance fidelity to the prior embeddings against adaptation to new co‑occurrence statistics; (c) Mittens accelerates convergence and often improves final performance in both classification and sequence‑labeling settings; (d) the approach is robust enough to transfer to related but distinct tasks (edge prediction in a large medical ontology). The authors conclude that Mittens offers a simple, effective way to combine massive general‑purpose corpora with modest domain‑specific data, and they suggest extending the retrofitting idea to newer contextual models such as BERT or ELMo.
Comments & Academic Discussion
Loading comments...
Leave a Comment