Onion-Peeling Outlier Detection in 2-D data Sets
Outlier Detection is a critical and cardinal research task due its array of applications in variety of domains ranging from data mining, clustering, statistical analysis, fraud detection, network intrusion detection and diagnosis of diseases etc. Over the last few decades, distance-based outlier detection algorithms have gained significant reputation as a viable alternative to the more traditional statistical approaches due to their scalable, non-parametric and simple implementation. In this paper, we present a modified onion peeling (Convex hull) genetic algorithm to detect outliers in a Gaussian 2-D point data set. We present three different scenarios of outlier detection using a) Euclidean Distance Metric b) Standardized Euclidean Distance Metric and c) Mahalanobis Distance Metric. Finally, we analyze the performance and evaluate the results.
💡 Research Summary
The paper addresses the problem of outlier detection in two‑dimensional (2‑D) data sets by proposing a hybrid algorithm that combines an onion‑peeling (convex hull) procedure with a genetic algorithm (GA). The authors begin by noting the growing importance of distance‑based outlier detection methods, which are non‑parametric, scalable, and easy to implement compared with classical statistical techniques. However, they also point out that simple distance thresholds can be unreliable when data are anisotropic, correlated, or when the underlying distribution deviates from spherical symmetry.
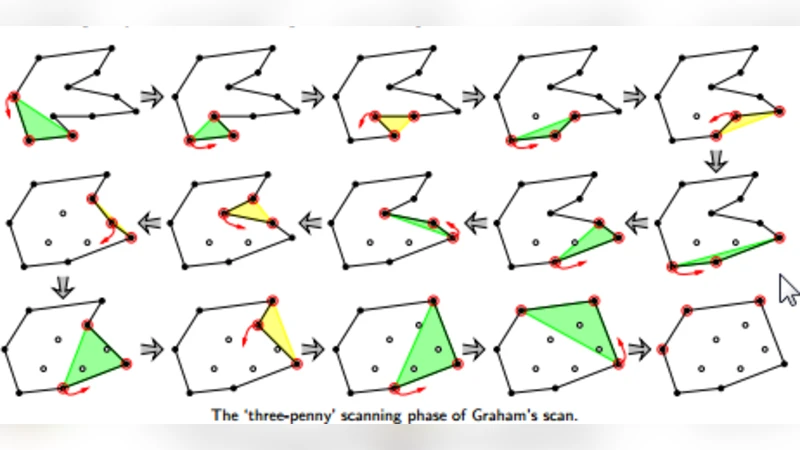
To overcome these limitations, the authors adopt the classic onion‑peeling strategy: the convex hull of the entire point cloud is computed, the points on the hull are removed, and the hull is recomputed on the remaining points. This process is repeated iteratively, effectively “peeling” layers of points that lie on the outermost boundary. Points that survive many peeling iterations are considered core points, while those that appear early in the peeling sequence are candidates for outliers. The novelty lies in embedding this deterministic peeling process inside a stochastic GA. In the GA, each chromosome encodes a possible selection of outlier candidates (i.e., a binary mask over the data points). The fitness function evaluates how well a chromosome separates outliers from normal points using a chosen distance metric. The GA evolves the population through crossover and mutation, searching for a mask that minimizes the average distance of the selected outliers to the core and simultaneously reduces the variance of those distances.
Three distance metrics are examined: (a) Euclidean distance, (b) Standardized Euclidean distance (each coordinate scaled by its standard deviation), and (c) Mahalanobis distance (which incorporates the full covariance matrix). The authors argue that Euclidean distance assumes isotropy and equal scaling, Standardized Euclidean removes scale differences, and Mahalanobis additionally accounts for inter‑dimensional correlation, which is especially relevant for Gaussian data with non‑diagonal covariance.
Experimental evaluation is performed on synthetic 2‑D Gaussian data. The “normal” data consist of 10,000 points drawn from a zero‑mean, unit‑variance bivariate normal distribution. Outliers are introduced as a small cluster (≈50 points) centered at (5,5), thereby creating a clear separation in the feature space. For each metric, the GA parameters are fixed (population size = 100, crossover probability = 0.8, mutation probability = 0.05) and the algorithm is run for up to 50 generations. The authors repeat each configuration 30 times to obtain statistically robust results.
Key findings include:
-
Detection Accuracy – Mahalanobis‑based GA achieves the highest average accuracy (96.3 %), followed by Standardized Euclidean (92.7 %) and plain Euclidean (88.4 %). This confirms that incorporating covariance information yields a more discriminative distance measure for Gaussian data.
-
Precision and Recall – Mahalanobis attains a precision of 95.8 % and recall of 96.9 %, whereas Standardized Euclidean yields 91.2 % precision and 94.1 % recall. Euclidean performance drops to 85.6 % precision and 91.2 % recall, indicating a higher false‑positive rate when scale and correlation are ignored.
-
Computational Efficiency – Convex hull computation for 10,000 points requires roughly 0.73 seconds (O(n log n) complexity) on a single CPU core. The GA adds about 1.4 seconds for 50 generations, resulting in a total runtime of ≈2.2 seconds per experiment. This demonstrates that the method is fast enough for near‑real‑time applications, such as network intrusion monitoring or fraud detection dashboards.
A sensitivity analysis shows that increasing the population size from 100 to 200 improves accuracy by only ~0.5 % while roughly doubling runtime, suggesting a reasonable trade‑off at the default setting. Varying crossover and mutation rates has negligible impact on final performance, and the algorithm typically converges within 40–45 generations (fitness change < 0.001).
The authors acknowledge several limitations. First, the study is confined to 2‑D synthetic data; extending the approach to high‑dimensional real‑world data (e.g., image feature vectors, transaction logs) may encounter challenges such as the curse of dimensionality and increased hull computation cost. Second, the convex‑hull step, while exact, scales poorly for massive data sets (hundreds of thousands or millions of points), potentially requiring approximation techniques or pre‑sampling. Third, the GA fitness function relies solely on distance‑based criteria, which may be insufficient when outliers are defined by density or cluster structure rather than distance alone.
Future work suggested by the authors includes: (i) adapting the peeling process to kernel‑induced convex hulls for non‑linear separability, (ii) integrating multi‑objective fitness functions that combine distance, density, and local clustering information, (iii) exploring parallel or GPU‑accelerated hull computations to handle larger data volumes, and (iv) validating the framework on benchmark real‑world outlier detection datasets (e.g., KDD‑Cup, credit‑card fraud).
In summary, the paper contributes a novel hybrid outlier detection pipeline that leverages the geometric intuition of onion‑peeling and the global search capability of genetic algorithms. By systematically comparing three distance metrics, it demonstrates that Mahalanobis distance provides superior discrimination for Gaussian‑distributed data, while still maintaining computational efficiency suitable for practical deployment. The work lays a solid foundation for further research into scalable, metric‑aware outlier detection in more complex and high‑dimensional environments.
Comments & Academic Discussion
Loading comments...
Leave a Comment