Bayesian Unmixing using Sparse Dirichlet Prior with Polynomial Post-nonlinear Mixing Model
A sparse Dirichlet prior is proposed for estimating the abundance vector of hyperspectral images with a nonlinear mixing model. This sparse prior is led to an unmixing procedure in a semi-supervised scenario in which exact materials are unknown. The nonlinear model is a polynomial post-nonlinear mixing model that represents each hyperspectral pixel as a nonlinear function of pure spectral signatures corrupted by additive white noise. Simulation results show more than 50% improvement in the estimation error.
💡 Research Summary
The paper addresses hyperspectral unmixing in a realistic semi‑supervised scenario where the exact endmember signatures are unknown but a large spectral library is available. To cope with the inherent non‑linearity of many natural scenes, the authors adopt the Polynomial Post‑Nonlinear Mixing Model (PPNMM), which represents each pixel as a linear mixture of endmember spectra followed by a second‑order polynomial transformation and additive white Gaussian noise. This model captures both linear and bilinear interactions while keeping computational complexity modest.
A central contribution is the introduction of a Sparse Symmetric Dirichlet prior for the abundance vector. The Dirichlet distribution’s concentration parameter α controls sparsity: values α < 1 force most abundance components toward zero, reflecting the practical assumption that only a few endmembers from a huge library actually contribute to any given pixel. By setting α to a small value, the prior simultaneously performs endmember selection and abundance estimation, eliminating the need for a separate Endmember Extraction Algorithm (EEA).
The Bayesian hierarchy is completed with Jeffreys’ prior for the noise variance σ², a zero‑mean Gaussian prior for the nonlinear coefficient b (with variance τ²), and an inverse‑Gamma hyper‑prior for τ². The resulting joint posterior over (a, b, σ², τ²) is analytically intractable, so the authors employ a Metropolis‑Within‑Gibbs sampler. In each iteration they sample a from its Dirichlet conditional, b from a Gaussian conditional, σ² from its Jeffreys conditional, and τ² from an inverse‑Gamma conditional. The algorithm runs for 10 000 MCMC iterations with a 1 000‑iteration burn‑in, and the posterior means are taken as point estimates.
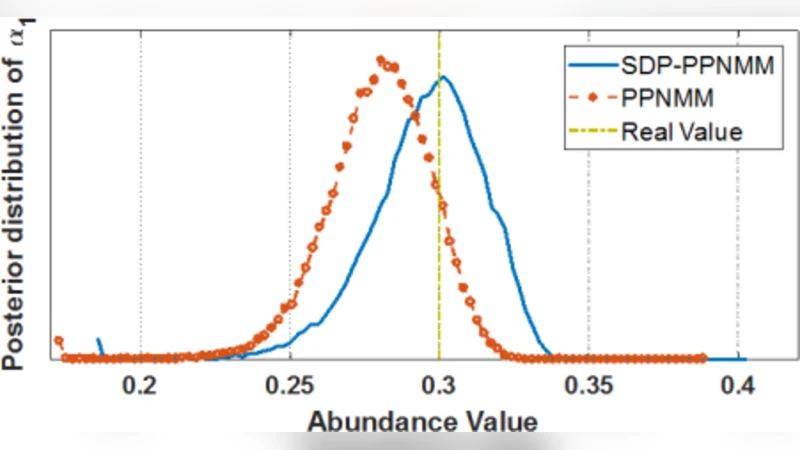
Experimental validation uses synthetic data generated from the USGS spectral library. Six endmembers are randomly drawn to form a library; only two are actually mixed to create a pixel, yielding an abundance vector with four zero entries—an explicit test of sparsity. The proposed SDP‑PPNMM method is compared against the conventional PPNMM that employs a uniform Dirichlet prior (α = 1). Over 20 Monte‑Carlo runs, the SDP‑PPNMM achieves a mean‑square error (MSE) of 0.0238 × 10⁻² versus 0.05 × 10⁻² for the baseline, a reduction of more than 50 %. Reconstruction error (RE) also improves slightly (5.17 % vs 5.30 %). Posterior distributions of the non‑zero abundances are sharper and more concentrated around the true values, while the zero‑abundance components exhibit a pronounced peak at zero, confirming effective sparsity enforcement.
In summary, the paper presents a hierarchical Bayesian unmixing framework that (i) removes the dependency on prior endmember extraction, (ii) handles nonlinear mixing via a tractable second‑order polynomial model, and (iii) leverages a sparsity‑inducing Dirichlet prior to jointly select relevant endmembers from a large library. The demonstrated >50 % reduction in estimation error underscores the practical advantage of the approach. Future work could explore adaptive learning of the concentration parameter α, incorporation of higher‑order nonlinearities, or application to real‑world hyperspectral datasets.
Comments & Academic Discussion
Loading comments...
Leave a Comment