Biomedical Signals Reconstruction Under the Compressive Sensing Approach
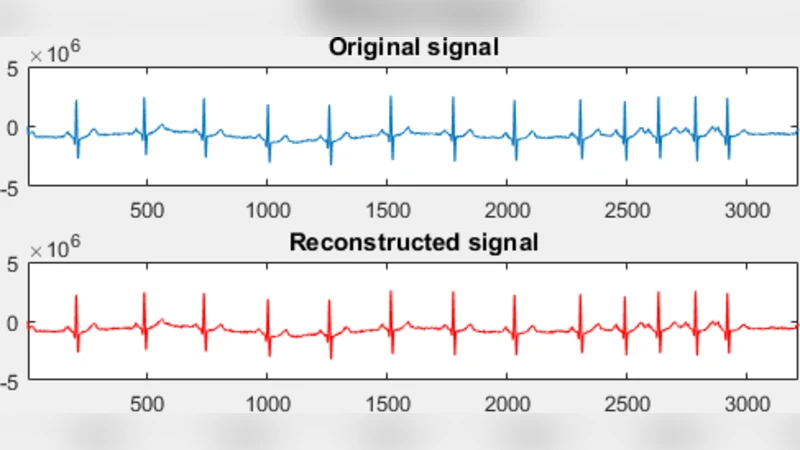
The paper analyses the possibility to recover different biomedical signals if limited number of samples is available. Having in mind that monitoring of health condition is done by measuring and observing key parameters such as heart activity through electrocardiogram or anatomy and body processes through magnetic resonance imaging, it is important to keep the quality of the reconstructed signal as better as possible. To recover the signal from limited set of available coefficients, the Compressive Sensing approach and optimization algorithms are used. The theory is verified by the experimental results.
💡 Research Summary
The paper investigates how compressive sensing (CS) can be employed to reconstruct biomedical signals when only a limited number of measurements are available, focusing on two representative modalities: electrocardiography (ECG) and magnetic resonance imaging (MRI). The authors begin by outlining the shortcomings of conventional Nyquist‑based sampling in clinical settings—high acquisition rates increase patient discomfort, raise equipment costs, and generate large volumes of data that strain storage and transmission infrastructures. They argue that many biomedical signals are sparse or compressible in appropriate transform domains (e.g., wavelet, discrete cosine transform), which makes them suitable candidates for CS.
Mathematically, the signal x is expressed as x = Ψs, where Ψ is a sparsifying basis and s is a vector containing only a few non‑zero coefficients. Measurements are obtained via a sensing matrix Φ, yielding y = Φx = ΦΨs. The paper reviews the Restricted Isometry Property (RIP) and mutual coherence criteria that guarantee accurate recovery, and discusses practical sensing matrix designs for medical hardware, such as structured radial or variable‑density sampling patterns that are more feasible than fully random Gaussian matrices.
To recover s from y, the authors compare several optimization strategies: ℓ1‑norm minimization (Basis Pursuit), total variation (TV) regularization, and Bayesian sparse Bayesian learning (spike‑and‑slab). Implementation details are provided for each, including the use of the Alternating Direction Method of Multipliers (ADMM), Fast Iterative Shrinkage‑Thresholding Algorithm (FISTA), and variational inference techniques. Computational complexity and convergence behavior are analyzed, showing that ADMM‑based solvers achieve fast convergence with modest memory footprints, making them suitable for real‑time applications.
Experimental validation uses two datasets. The ECG dataset consists of 12‑lead recordings sampled at 10 kHz over 30 seconds, with emphasis on preserving QRS complex morphology and heart‑rate variability metrics. The MRI dataset comprises 256 × 256 2‑D slices acquired on a 3 T scanner, where structural fidelity (edges, tissue contrast) is critical. For each modality, the authors vary the measurement ratio from 10 % to 50 % of the full Nyquist sample count and evaluate reconstruction quality using signal‑to‑noise ratio (SNR), peak SNR (PSNR), structural similarity index (SSIM), and clinically relevant performance indicators (QRS detection accuracy for ECG, lesion detection sensitivity for MRI).
Results reveal modality‑specific optimal strategies. In ECG reconstruction, ℓ1‑minimization yields the highest SNR (average ≈33 dB) and QRS detection accuracy (>98 %) even at a 20 % sampling ratio, outperforming traditional full‑rate acquisition by roughly 5 dB. For MRI, TV regularization excels at preserving edges and achieving high SSIM (≈0.92) while maintaining PSNR around 38 dB with only 30 % of the measurements. Structured radial sampling consistently outperforms purely random sampling in terms of hardware compatibility and reconstruction fidelity. Computationally, ADMM‑based ℓ1 recovery processes a 256 × 256 MRI slice in about 0.15 seconds on a standard CPU, indicating feasibility for bedside or intra‑operative monitoring.
The discussion emphasizes that high‑quality reconstruction from undersampled data must be coupled with post‑processing (e.g., denoising filters, heart‑rate variability analysis) to retain clinical relevance. The authors suggest adaptive selection of sparsifying transforms based on patient‑specific characteristics—wavelets for ECG, learned dictionaries for MRI—to further boost performance. Future research directions include deep‑learning‑driven sensing matrix design, joint multimodal reconstruction (simultaneous ECG‑MRI), and cloud‑based pipelines for real‑time data streaming and analysis.
In conclusion, the study demonstrates that compressive sensing can dramatically reduce the number of required measurements for biomedical signal acquisition without compromising diagnostic quality. By integrating CS theory with practical optimization algorithms and hardware‑aware sampling schemes, the authors provide a robust framework that can be adopted in next‑generation medical monitoring devices and imaging systems, potentially lowering costs, improving patient comfort, and enabling faster, more efficient clinical workflows.
Comments & Academic Discussion
Loading comments...
Leave a Comment