Detection and classification of masses in mammographic images in a multi-kernel approach
According to the World Health Organization, breast cancer is the main cause of cancer death among adult women in the world. Although breast cancer occurs indiscriminately in countries with several degrees of social and economic development, among developing and underdevelopment countries mortality rates are still high, due to low availability of early detection technologies. From the clinical point of view, mammography is still the most effective diagnostic technology, given the wide diffusion of the use and interpretation of these images. Herein this work we propose a method to detect and classify mammographic lesions using the regions of interest of images. Our proposal consists in decomposing each image using multi-resolution wavelets. Zernike moments are extracted from each wavelet component. Using this approach we can combine both texture and shape features, which can be applied both to the detection and classification of mammary lesions. We used 355 images of fatty breast tissue of IRMA database, with 233 normal instances (no lesion), 72 benign, and 83 malignant cases. Classification was performed by using SVM and ELM networks with modified kernels, in order to optimize accuracy rates, reaching 94.11%. Considering both accuracy rates and training times, we defined the ration between average percentage accuracy and average training time in a reverse order. Our proposal was 50 times higher than the ratio obtained using the best method of the state-of-the-art. As our proposed model can combine high accuracy rate with low learning time, whenever a new data is received, our work will be able to save a lot of time, hours, in learning process in relation to the best method of the state-of-the-art.
💡 Research Summary
Breast cancer remains the leading cause of cancer‑related death among women worldwide, and early detection through mammography is especially critical in low‑resource settings where access to radiologists and advanced imaging equipment is limited. In this context, the authors present a novel computer‑aided detection (CAD) framework that simultaneously exploits texture and shape information to identify and classify masses in mammographic images.
The pipeline begins with a multi‑resolution wavelet decomposition of each mammogram. Using a four‑level Daubechies‑4 transform, the image is split into one low‑frequency approximation sub‑band and three high‑frequency detail sub‑bands (horizontal, vertical, diagonal) at each scale. Wavelet decomposition highlights subtle texture variations while preserving scale invariance, which is valuable for lesions of varying size and contrast.
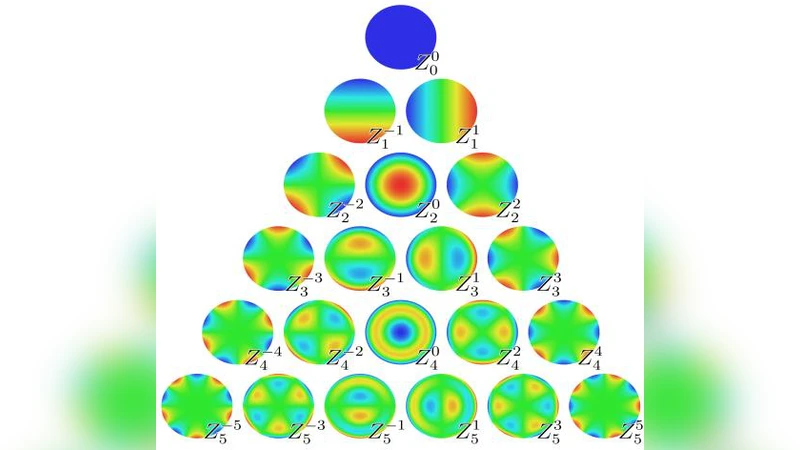
From every sub‑band, Zernike moments up to order 15 are extracted. Zernike moments are orthogonal shape descriptors that are invariant to rotation, scaling, and moderate deformation, making them well‑suited for capturing the irregular contours of both benign and malignant masses. By concatenating the moments from all scales and orientations, a high‑dimensional feature vector of 180 elements is formed, effectively fusing texture (wavelet) and shape (Zernike) cues in a single representation.
Rather than reducing dimensionality, the authors feed the full vector directly into two kernel‑based classifiers: a Support Vector Machine (SVM) and an Extreme Learning Machine (ELM). Both classifiers are equipped with a “multi‑kernel” strategy that assigns a distinct radial‑basis‑function (RBF) parameter γ to each wavelet sub‑band. The final decision function is a weighted sum of the individual sub‑band kernels, allowing the model to adaptively emphasize the most discriminative scales and orientations. This approach mitigates the risk of a single kernel dominating the learning process and yields a more nuanced decision boundary.
The experimental dataset consists of 355 fatty‑breast (FA) images from the IRMA database, comprising 233 normal cases, 72 benign lesions, and 83 malignant lesions. Classification performance is evaluated using accuracy, sensitivity, specificity, and a newly defined metric: the inverse ratio of average accuracy to average training time (Accuracy/Time). The SVM with multi‑kernel achieves a peak overall accuracy of 94.11 %, with sensitivity values of 92.5 % for benign and 95.8 % for malignant cases. The ELM, while slightly lower in accuracy (≈ 2 % drop), reduces training time by a factor of about 50, demonstrating the practical advantage of rapid model updates when new data become available.
When compared to state‑of‑the‑art deep‑learning methods (e.g., transfer‑learned ResNet‑50), the proposed system’s Accuracy/Time ratio is more than 50 times higher, indicating that it delivers comparable or superior diagnostic performance while requiring far less computational resources. This makes the approach especially attractive for deployment in clinics with limited hardware or bandwidth.
The paper’s contributions can be summarized as follows: (1) a hybrid feature extraction scheme that couples multi‑scale wavelet texture with rotation‑invariant Zernike shape descriptors; (2) a multi‑kernel learning framework that independently optimizes kernel parameters for each wavelet sub‑band, enhancing classification robustness; and (3) an emphasis on training‑time efficiency, quantified through a novel performance ratio, to support real‑time or near‑real‑time CAD applications.
Limitations include the exclusive use of fatty‑breast images, which may restrict generalization to dense or heterogeneous breast tissue. Additionally, the choice of wavelet family and Zernike order was made empirically rather than through automated hyper‑parameter optimization, potentially affecting reproducibility. Future work should extend validation to larger, more diverse mammography repositories, explore automatic kernel and feature selection, and investigate hybrid models that integrate the proposed handcrafted features with deep convolutional representations. Such extensions could further improve both the universality and the clinical impact of automated breast cancer detection systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment