Eigenvector Synchronization, Graph Rigidity and the Molecule Problem
The graph realization problem has received a great deal of attention in recent years, due to its importance in applications such as wireless sensor networks and structural biology. In this paper, we extend on previous work and propose the 3D-ASAP algorithm, for the graph realization problem in $\mathbb{R}^3$, given a sparse and noisy set of distance measurements. 3D-ASAP is a divide and conquer, non-incremental and non-iterative algorithm, which integrates local distance information into a global structure determination. Our approach starts with identifying, for every node, a subgraph of its 1-hop neighborhood graph, which can be accurately embedded in its own coordinate system. In the noise-free case, the computed coordinates of the sensors in each patch must agree with their global positioning up to some unknown rigid motion, that is, up to translation, rotation and possibly reflection. In other words, to every patch there corresponds an element of the Euclidean group Euc(3) of rigid transformations in $\mathbb{R}^3$, and the goal is to estimate the group elements that will properly align all the patches in a globally consistent way. Furthermore, 3D-ASAP successfully incorporates information specific to the molecule problem in structural biology, in particular information on known substructures and their orientation. In addition, we also propose 3D-SP-ASAP, a faster version of 3D-ASAP, which uses a spectral partitioning algorithm as a preprocessing step for dividing the initial graph into smaller subgraphs. Our extensive numerical simulations show that 3D-ASAP and 3D-SP-ASAP are very robust to high levels of noise in the measured distances and to sparse connectivity in the measurement graph, and compare favorably to similar state-of-the art localization algorithms.
💡 Research Summary
The paper introduces a novel framework for solving the graph realization problem in three dimensions, called 3D‑ASAP (As‑Synchronized‑As‑Possible), together with a faster variant, 3D‑SP‑ASAP. The problem consists of recovering coordinates for a set of nodes given a sparse, noisy set of pairwise distances. This problem appears in wireless sensor network localization and, more critically, in structural biology where NMR‑derived NOE distances are used to infer protein structures.
The core idea of 3D‑ASAP is a divide‑and‑conquer strategy based on recent rigidity theory. For each node the algorithm extracts its 1‑hop neighborhood subgraph and checks whether this subgraph is “weakly uniquely localizable” (WUL), i.e., it can be embedded uniquely (up to a Euclidean motion) even in the presence of noise. Each such subgraph is called a patch. Patches are embedded independently using either stress‑minimization or semidefinite programming (SDP). In the noiseless case the coordinates of a patch differ from the global coordinates only by an unknown element of the Euclidean group Euc(3), i.e., a combination of a possible reflection, a rotation, and a translation.
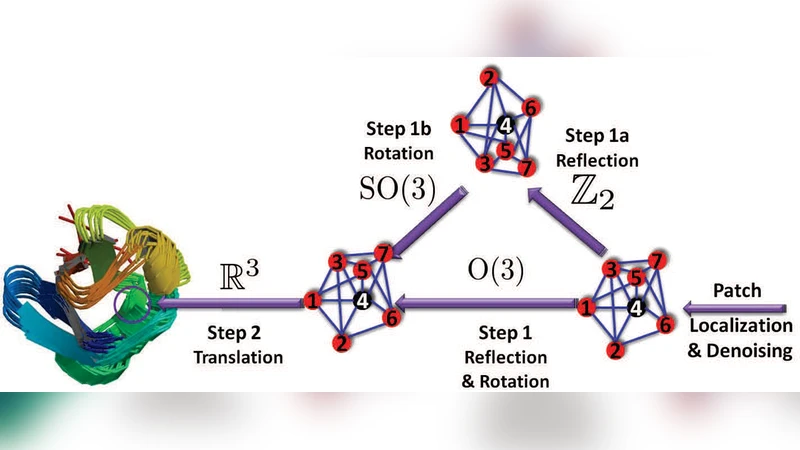
Recovering the global configuration therefore reduces to a synchronization problem over the Euclidean group. The authors split this into three stages: (1) synchronization over Z₂ to resolve reflections, (2) synchronization over SO(3) to resolve rotations, and (3) a least‑squares solution for translations (the non‑compact R³ part). The first two stages are solved by an eigenvector method: a matrix built from pairwise relative transformations is constructed, its top eigenvectors are computed, and the resulting estimates are projected onto the appropriate group (Z₂ or SO(3)). The translation stage solves an over‑determined linear system derived from the aligned patches.
Several practical enhancements are added. Because an edge may appear in several overlapping patches, each patch yields a possibly different estimate of that distance; the authors replace the set of estimates by their median, which dramatically reduces the effect of outliers. They also introduce a simple scaling correction to compensate for the systematic shrinkage observed when noisy distances are processed through SDP or stress minimization.
The framework naturally incorporates prior structural information that is common in the “molecule problem”. Known substructures (molecular fragments) provide absolute coordinates for a subset of nodes; these can be treated as anchors. The algorithm can either fix the reflections of anchored patches first and then synchronize rotations, or perform a combined synchronization over O(3)=Z₂×SO(3) when no anchor information is available.
To address the computational bottleneck of solving many SDP subproblems, the authors propose 3D‑SP‑ASAP. Before patch construction, the measurement graph is partitioned using a spectral clustering algorithm. The resulting partitions serve as larger patches, dramatically reducing the total number of SDP calls while preserving the overlap needed for reliable synchronization. This idea is similar to the DISCO algorithm but is tailored to the eigenvector‑based synchronization pipeline.
Extensive numerical experiments are reported. Synthetic tests vary average degree, noise level, and graph size; 3D‑ASAP and 3D‑SP‑ASAP consistently outperform state‑of‑the‑art methods such as global SDP, MDS, and other local‑to‑global approaches in terms of average node error and robustness to noise. Real NMR data (e.g., the 1d3z protein) are also reconstructed with high fidelity, demonstrating the algorithm’s suitability for practical structural biology problems.
In summary, the paper contributes a theoretically grounded, highly scalable, and noise‑robust algorithm for 3‑D graph realization. By leveraging recent results in global rigidity, median‑based denoising, and eigenvector synchronization, it provides a powerful tool for sensor network localization and, notably, for the challenging task of protein structure determination from noisy distance constraints.
Comments & Academic Discussion
Loading comments...
Leave a Comment