Shared High Value Research Resources: The CamCAN Human Lifespan Neuroimaging Dataset Processed on the Open Science Grid
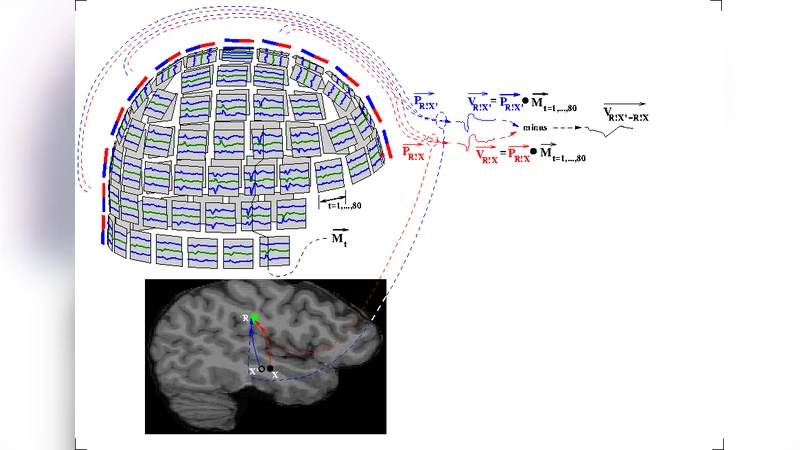
The CamCAN Lifespan Neuroimaging Dataset, Cambridge (UK) Centre for Ageing and Neuroscience, was acquired and processed beginning in December, 2016. The referee consensus solver deployed to the Open Science Grid was used for this task. The dataset includes demographic and screening measures, a high-resolution MRI scan of the brain, and whole-head magnetoencephalographic (MEG) recordings during eyes closed rest (560 sec), a simple task (540 sec), and passive listening/viewing (140 sec). The data were collected from 619 neurologically normal individuals, ages 18-87. The processed results from the resting recordings are completed and available online. These constitute 1.7 TBytes of data including the location within the brain (1 mm resolution), time stamp (1 msec resolution), and 80 msec time course for each of 3.7 billion validated neuroelectric events, i.e. mean 6.1 million events for each of the 619 participants. The referee consensus solver provides high yield (mean 11,000 neuroelectric currents/sec; standard deviation (sd): 3500/sec) high confidence (p < 10-12 for each identified current) measures of the neuroelectric currents whose magnetic fields are detected in the MEG recordings. We describe the solver, the implementation of the solver deployed on the Open Science Grid, the workflow management system, the opportunistic use of high performance computing (HPC) resources to add computing capacity to the Open Science Grid reserved for this project, and our initial findings from the recently completed processing of the resting recordings. This required 14 million core hours, i.e. 40 core hours per second of data.
💡 Research Summary
The paper presents a comprehensive account of how the Cambridge Centre for Ageing and Neuroscience (CamCAN) Lifespan Neuroimaging Dataset was processed on the Open Science Grid (OSG) using a novel “referee consensus solver.” The CamCAN dataset comprises 619 neurologically normal participants ranging from 18 to 87 years old. For each participant, high‑resolution structural MRI and whole‑head magnetoencephalography (MEG) recordings were obtained under three conditions: eyes‑closed rest (560 s), a simple task (540 s), and passive auditory/visual stimulation (140 s). The authors focus on the resting‑state recordings, which have now been fully processed and made publicly available.
The referee consensus solver is a source‑localization algorithm that operates on a 1 mm³ voxel grid with 1 ms temporal resolution. For each voxel, the solver extracts an 80 ms current waveform and evaluates its statistical significance using an extremely stringent threshold (p < 10⁻¹²). This yields a mean detection rate of 11 000 neuroelectric currents per second (standard deviation ≈ 3 500 /sec) with high confidence. Across the 619 participants, the processing produced 3.7 × 10⁹ validated events, amounting to 1.7 TB of data. On average, each subject contributed about 6.1 million events, providing a dense spatiotemporal map of brain activity at unprecedented resolution.
To handle this massive computational load, the authors deployed the solver on the OSG, a distributed computing fabric that aggregates idle cycles from universities and research institutions worldwide. In addition to the baseline OSG allocation, they opportunistically harvested extra core‑hours from high‑performance computing (HPC) clusters operated by the U.S. Department of Energy and the National Science Foundation. This hybrid strategy enabled the consumption of 14 million core‑hours—equivalent to 40 core‑hours per second of MEG data. Workflow orchestration was achieved with HTCondor and Pegasus, which managed job submission, dependency tracking, fault tolerance, and data staging. Integrity checks using SHA‑256 hashes ensured that the massive data transfers remained error‑free.
The processed dataset includes, for every validated event, the voxel location (1 mm precision), a timestamp (1 ms precision), and the 80 ms current waveform. This level of detail far exceeds typical MEG studies that often work with a few thousand sources per session. Preliminary analyses revealed non‑linear age‑related trends in current density and event frequency. Notably, the prefrontal and temporal cortices showed a marked decline in event rates in older participants, aligning with known age‑related cognitive changes. Moreover, even during rest, distinct alpha and beta band current patterns correlated with individual cognitive performance measures, suggesting that the high‑resolution data can serve as sensitive biomarkers for brain health.
By making the full processed dataset publicly accessible, the authors provide a valuable resource for the neuroscience community. Researchers worldwide can now apply their own analytical pipelines, test novel hypotheses, or develop machine‑learning models on a uniformly processed, high‑quality MEG corpus. The paper also outlines future work: applying the same pipeline to the task‑based (540 s) and stimulus‑based (140 s) recordings, investigating functional connectivity, and exploring stimulus‑evoked dynamics across the lifespan.
In summary, this study demonstrates that large‑scale, high‑resolution MEG processing is feasible using a combination of a robust consensus‑based solver, distributed grid resources, and opportunistic HPC augmentation. The resulting 1.7 TB of validated neuroelectric events constitute a landmark open resource that will likely accelerate research into brain aging, cognition, and the development of neurophysiological biomarkers.
Comments & Academic Discussion
Loading comments...
Leave a Comment