Adapting a Formal Model Theory to Applications in Augmented Personalized Medicine
The goal of this paper is to advance an extensible theory of living systems using an approach to biomathematics and biocomputation that suitably addresses self-organized, self-referential and anticipatory systems with multi-temporal multi-agents. Our first step is to provide foundations for modelling of emergent and evolving dynamic multi-level organic complexes and their sustentative processes in artificial and natural life systems. Main applications are in life sciences, medicine, ecology and astrobiology, as well as robotics, industrial automation and man-machine interface. Since 2011 over 100 scientists from a number of disciplines have been exploring a substantial set of theoretical frameworks for a comprehensive theory of life known as Integral Biomathics. That effort identified the need for a robust core model of organisms as dynamic wholes, using advanced and adequately computable mathematics. The work described here for that core combines the advantages of a situation and context aware multivalent computational logic for active self-organizing networks, Wandering Logic Intelligence (WLI), and a multi-scale dynamic category theory, Memory Evolutive Systems (MES), hence WLIMES. This is presented to the modeller via a formal augmented reality language as a first step towards practical modelling and simulation of multi-level living systems. Initial work focuses on the design and implementation of this visual language and calculus (VLC) and its graphical user interface. The results will be integrated within the current methodology and practices of theoretical biology and (personalized) medicine to deepen and to enhance the holistic understanding of life.
💡 Research Summary
**
The paper puts forward a comprehensive theoretical framework aimed at modeling living systems as dynamic wholes, an effort that the authors term Integral Biomathics. Recognizing that traditional biological and medical models are largely static and hierarchical, the authors argue that a new mathematics and computation paradigm is required to capture the self‑organizing, self‑referential, and anticipatory nature of multi‑temporal, multi‑agent systems. To this end they combine two previously developed formalisms: Wandering Logic Intelligence (WLI) and Memory Evolutive Systems (MES).
WLI is a multivalent logical system that continuously adapts its inference rules to the current situation and context. Unlike classical propositional or predicate logic, which assumes fixed rule sets, WLI allows the logical “rules” themselves to wander, re‑configure, and evolve as the network’s environment changes. This property makes WLI especially suitable for representing the non‑linear, non‑deterministic dynamics observed in cellular signaling networks, immune responses, and other biological processes that exhibit feedback loops and emergent behavior.
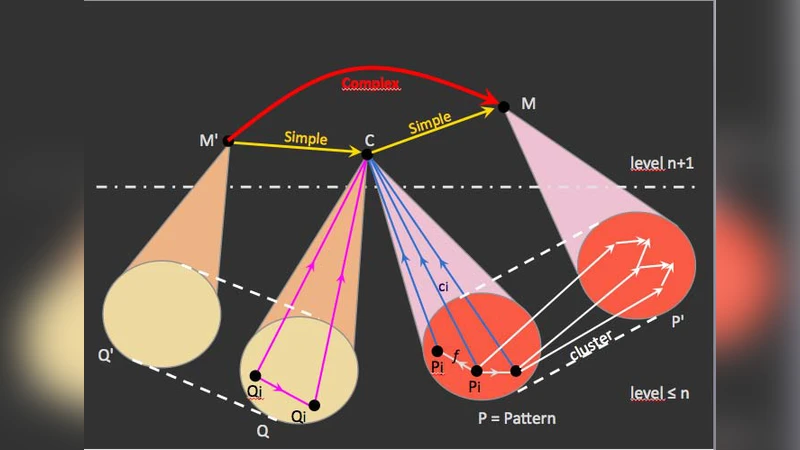
MES, on the other hand, is a category‑theoretic construction that models objects and morphisms across multiple scales (e.g., molecules, cells, tissues, organs, whole organisms). Its distinctive feature is a “memory” structure that records the history of morphism transformations, thereby enabling the representation of structural re‑organization over time. In MES, each scale is a categorical object, and the relationships between scales are expressed as functors that can themselves evolve, providing a mathematically rigorous way to describe hierarchical emergence and de‑emergence.
The authors’ central contribution is the synthesis of these two approaches into a unified framework they call WLIMES. In WLIMES, the context‑aware logical dynamics of WLI are mapped onto the multi‑scale categorical architecture of MES. Consequently, every situational change triggers a corresponding transformation of the categorical morphisms, allowing the model to simultaneously track “situation‑context‑time” dimensions. This integration yields a formalism capable of representing both the local, context‑dependent decision making of individual agents and the global, hierarchical evolution of the whole system.
To make WLIMES usable for scientists and clinicians, the paper introduces a Visual Language and Calculus (VLC). VLC is a graphical programming environment where nodes, edges, and icons directly correspond to the formal elements of WLIMES. Users construct models by dragging and linking visual components rather than writing complex equations. The authors further embed VLC in an augmented‑reality (AR) interface, enabling users to project models into physical space, manipulate parameters with gestures or voice commands, and observe real‑time simulations. This immersive interface is intended to lower the barrier to entry for non‑technical stakeholders, fostering collaborative model building and hypothesis testing.
A primary application domain highlighted is personalized (or “augmented”) medicine. Patient‑specific omics data (genomics, transcriptomics, proteomics, metabolomics) are mapped onto the MES hierarchy, creating a patient‑specific categorical structure. Therapeutic interventions—such as drug administration, gene editing, or lifestyle changes—are encoded as WLI‑driven situational updates. By running simulations on the WLIMES model, clinicians can predict how the patient’s biological network will reorganize over time, anticipate adverse feedback loops, and optimize treatment schedules. This approach promises to capture the non‑linear, time‑dependent feedback that conventional statistical or machine‑learning models often miss.
Beyond biomedicine, the authors discuss potential extensions to robotics, industrial automation, and human‑machine interfaces. In multi‑robot systems, each robot could run its own WLI module to interpret local sensor data, while the collective behavior would be coordinated through a shared MES structure, enabling adaptive, hierarchical collaboration that reacts to environmental changes in real time.
In conclusion, the paper presents WLIMES and its visual‑AR implementation as a novel, mathematically rigorous yet practically accessible platform for modeling complex adaptive systems. By uniting situation‑aware logic with multi‑scale categorical memory, the framework offers a way to encode and simulate the emergent, anticipatory dynamics that characterize living organisms and engineered adaptive systems alike. The authors outline future work involving validation with clinical datasets and robotic testbeds, aiming to demonstrate that WLIMES can serve as a cornerstone for a truly integrative theory of life and its technological analogues.
Comments & Academic Discussion
Loading comments...
Leave a Comment