Patient-Driven Privacy Control through Generalized Distillation
The introduction of data analytics into medicine has changed the nature of patient treatment. In this, patients are asked to disclose personal information such as genetic markers, lifestyle habits, and clinical history. This data is then used by stat…
Authors: Z. Berkay Celik, David Lopez-Paz, Patrick McDaniel
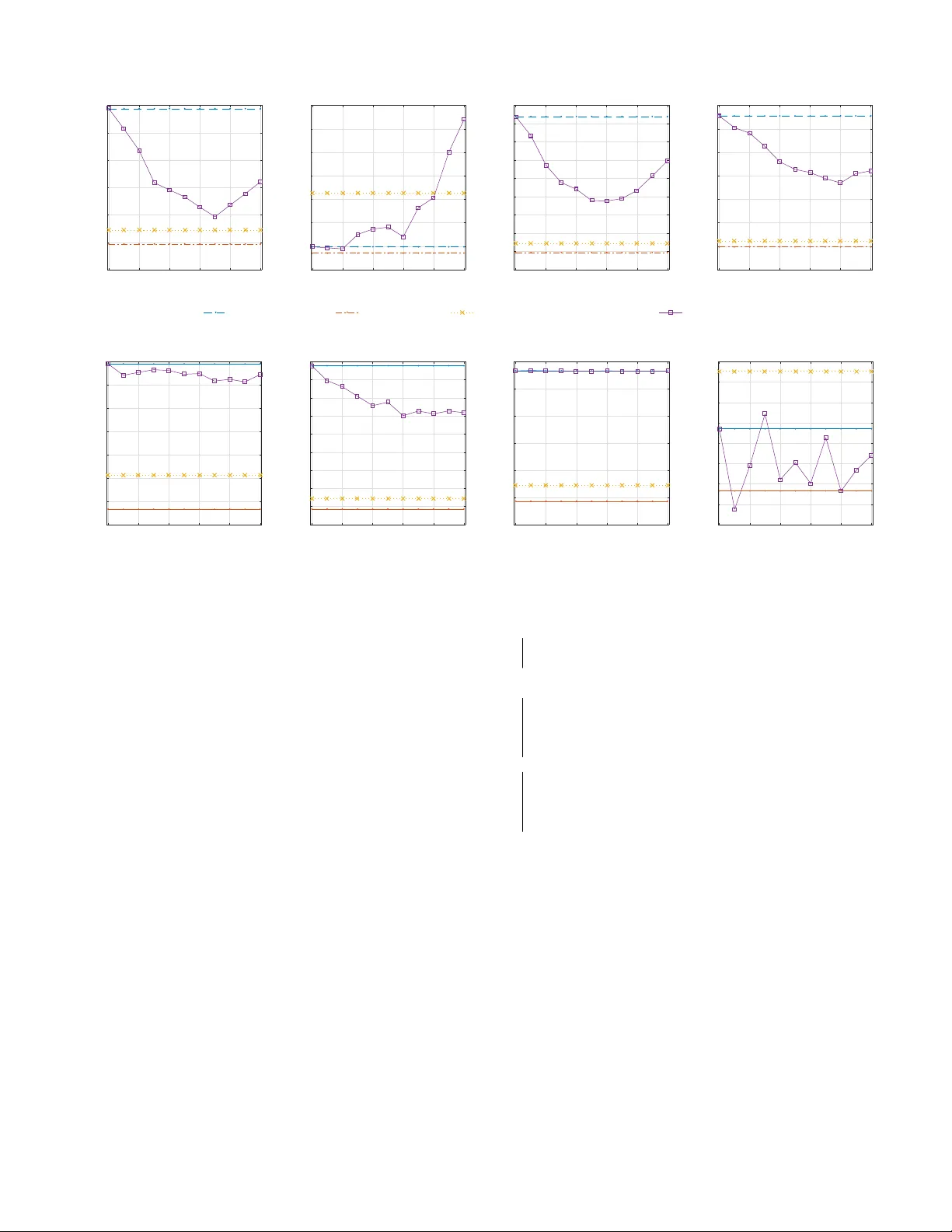
P atient-Dri v en Pri v ac y Control through Generalized Distillation Z. Berkay Celik 1 , David Lopez-P az 2 , and Patrick McDaniel 1 1 SIIS Laboratory , Department of CSE, The Pennsylv ania State University { zbc102, mcdaniel } @cse.psu.edu 2 Facebook AI Research dlp@fb.com Accepted to the IEEE Symposium on Priv acy-A ware Computing (IEEE P A C) , 2017. W ashington DC, USA. Abstract —The introduction of data analytics into medicine has changed the nature of patient tr eatment. In this, patients are asked to disclose personal information such as genetic markers, lifestyle habits, and clinical history . This data is then used by statistical models to predict personalized treatments. However , due to privacy concerns, patients often desire to withhold sensitive information. This self-censorship can impede proper diagnosis and treatment, which may lead to serious health complications and even death over time. In this paper , we present privacy distillation, a mechanism which allows patients to control the type and amount of inf ormation they wish to disclose to the healthcare pr oviders for use in statistical models. Meanwhile, it retains the accuracy of models that ha ve access to all patient data under a sufficient but not full set of privacy-r elev ant information. W e validate privacy distillation using a corpus of patients prescribed to warfarin for a personalized dosage. W e use a deep neural network to implement privacy distillation for training and making dose predictions. W e find that privacy distillation with sufficient privacy-rele vant information i) retains accuracy almost as good as ha ving all patient data (only 3% worse), and ii) is effective at pre venting errors that introduce health-related risks (only 3.9% worse under - or over -prescriptions). I . I N T RO D U C T I O N Data analytics has introduced a sea change in modern healthcare systems. In this, there are many advances in computation that hav e the potential to impact society positiv ely on analytically driv en personalized medicine [43]. This effort has lead to Precision Medicine Initiative [21], announced in 2015. This initiativ e aims to pioneer data-driven healthcare to treat and prevent disease, based on variability in patient data such as genes, en vironment, and lifestyle. Personalized medicine relies on building statistical mod- els able to predict personalized treatments, tailored to the characteristics of specific patients. In turn, developing such statistical models call for large amounts of patient data. Ho we ver , in practice, patients often prefer not to divulge certain kinds of data due to priv acy concerns. For instance, social stigma and discrimination concerns of patients suffering from mental disorders and sexually transmitted disease, embarrassing themselves in the presence of their physicians, lack of trust to the medical providers are one of the many reasons [2], [5], [19], [51]. In short, personalized medicine is based on statistical models that call for large bodies of patient data, but patients hav e different levels of willingness to expose their data 1 . T o illustrate this dilemma, consider the problem of per- sonalized warfarin dosing . W arfarin is an oral anticoagulant prescribed to prevent blood clotting. The International W arfarin Pharmacogenetics Consortium (IWPC) has built a statistical model to predict personalized warfarin dose [30]. The IWPC model requires sev eral types of patient data, including genetic markers, clinical history , and demographics. Remarkably , the IWPC model outperforms current standard clinical approaches, which rely on fixed-dose prescriptions. Howe ver , a practitioner requires all patients to disclose their complete information to predict treatments. For instance, a patient not filling out a particular medication intake status through a medical history form can not get precise treatment. The previous example illustrates one main shortcoming of current statistical healthcare models: all patients must provide all of their data, or prediction of treatment is impossible with patients withholding their sensitiv e data [48]. T o amend this problem, data imputation is often applied in healthcare for “fill- ing the missing” data using the information provided by public (av ailable) data [47]. Howe ver , imputation incurs cumbersome endeav or: in warfarin example, imputing medical history of patients is more complicated than predicting the warfarin dose. This may incur biased estimates under man y sensiti ve data if the relationship between missing and av ailable data is intricate [42]. More importantly , imputation is impossible if the missing data is statistically independent of the av ailable data [10]. Therefore, data-dri ven healthcare is in need for a general purpose solution to the problem of making treatments with priv acy concerns of patients. W e propose one solution to this problem, named priv acy distillation. Privacy Distillation- Priv acy distillation addresses the priv acy concerns of patients for the data used in healthcare statistical models. W e define patient data that compose features from genetic and clinical data. Genetic data include genes or DNA sequences with a kno wn location on a chromosome that its variability plays a major role in treatments. Clinical data includes inputs from medication history , psychological profiles, dietary habits, sexual preferences, demographics, and so on. 1 W e note that information withholding that we address in this paper and obscuring (patients lying about their sensitiv e information) are two different critical problems identified in healthcare. W e are currently working on obscurity problem which is algorithmically and conceptually more complex, as it is strictly harder than the withholding problem. n o n - r e d ac te d * r e d ac te d * X 1 ,* X 2* X 3 , X 4 … ... X 1 , X 2 , X 3 , X 4 ... … Pa t i e n t s d i scl o si n g a l l y ... ... T a rg e t X 3 ,* X 4 * L e ar n i n g * A l g o r i th m * y ... ... f ( X 1 , X 2 , X 3 , X 4 ) L e ar n i n g * A l g o r i th m * f (X 3 , X 4 ) ŷ Privacy distillation process for patient profile redacting X 1 and X 2 T raining time a b c f (X 3 , X 4 ) d ý * T est time A patient comes for treatment dis9ll* Fig. 1: Priv acy distillation scheme: The distilled model f ( x 3 , x 4 ) preserves the privac y of patients at treatment, since it only requires access to their non-redacted data ( x 3 , x 4 ) . Howev er, the distilled model was trained to imitate the predictions of a full model f ( x 1 , x 2 , x 3 , x 4 ) , which had access to the redacted data ( x 1 , x 2 ) of other patients and clinical studies. Through this transference of knowledge, the distilled model provides with more accurate treatments than models using only the non-redacted data ( x 3 , x 4 ) and treatments ´ y are nearly as accurate as models assuming access to the complete patient data ( x 1 , x 2 , x 3 , x 4 ). The intuition behind using both types of data is that they are useful for facilitating predictions of the correct treatments. In this setting, we define redacted featur es as those that patients withhold due to priv acy concerns. Priv acy distillation works as follows (see Figure 1): At training time, priv acy distillation creates a nov el model that retains the accuracy in the absence of redacted features. This is achiev ed by improving the model training by transferring knowledge from redacted features av ailable across other patients and clinical studies ( a ). At test time, a patient is asked to provide a set of clinical and genetic features for medical treatment through doctor-patient communication, surve ys and so on ( b ). The patient chooses to redact features she deems priv acy-sensitiv e ( c ). The redacted features of a patient are matched with a precomputed patient pr ofile that defines different patient disclosure behaviors among patients, and corresponding distilled model then makes the predictions on non-redacted features ( d ). If a patient desires a profile that has not been precomputed, we adapt two solutions (see Section III-A ). The goal of this effort is to develop an algorithm such that the accuracy under the redacted patient data is sufficiently close to that of complete data, and any errors in the model should not induce sev ere medical side effects. W e highlight that our approach is a generalizable solution, we here focus its application to personalized medicine. In this paper , we implement pri v acy distillation using a deep neural network (DNN) and ev aluate its effecti veness in the problem of warfarin dose prediction. W e compare privacy distillation with two baselines. The first baseline is partially- r edacted , a model built on public (non-redacted) features. The second baseline is the non-redacted , which is the current state- of-the-art linear model trained on complete (redacted and non-redacted) patient data. W e evaluate priv acy distillation by studying the trade-off between the amount of information disclosed by patients and dose accuracy; then we analyze dose errors introducing health-related risks. In this, we make the following contributions: • W e introduce priv acy distillation, a mechanism, that allows patients to control their sensitive information used in health- care statistical models. Priv acy distillation of fers an accuracy similar to the non-redacted model across various patient profiles; it increases the error of the non-redacted model by only 3% and outperforms the accuracy of the partially- redacted model by 13.4%. • W e show that accuracy loss in healthcare models leads to medical side effects. In the case of warfarin dose predictions, errors cause stroke, embolism, internal bleeding, and e ven mortality . Priv acy distillation provides personalized treatments that are within safety windo w of a state-of-the-art linear model for 96.1% of the patients; whereas partially- redacted models lead to 83.2%. • W e improve the accuracy of the current state-of-the-art warfarin dose linear model by 2.7% with the use of deep neural networks. I I . T E C H N I C A L P R E L I M I NA R I E S W e introduce the essentials of predictiv e models necessary to understand healthcare statistical models. W e then provide an overvie w of generalized distillation which our priv acy distillation mechanism builds on. A. Healthcar e Statistical Models Statistical models are functions f : X → Y that aim at predicting targ ets y ∈ Y giv en some explanatory features x ∈ X . In the following, we consider real-valued targets y ∈ R and real vector -v alued features x ∈ R d . Statistical models are built using a dataset containing pairs of features and targets, denoted by D = { ( x i , y i ) } n i = 1 , and a loss function ` : Y × Y → [ 0 , ∞ ) . The loss function penalizes de viations between true targets and predictions. Learning is then searching for the statistical model f minimizing the av erage loss: L ( D , f ) = 1 n n ∑ i = 1 ` ( f ( x i ) , y i ) . (1) For instance, in simple linear least-squares regression, the statistical model is linear , i.e. f ( x ) = α > x + β , and the loss function is the squared loss ` ( f ( x ) , y ) = ( f ( x ) − y ) 2 . Howe ver , to model complex nonlinear relationships between features and targets, we will not be using linear statistical models, but deep neural networks (DNN) [37]. In the domain of healthcare, the applications of statisti- cal models are skyrocketing: examples include personalized medicine prescriptions, online crowdsourcing healthcare, dis- ease risk tests, and personality trait tests. In these applications, the features x often correspond to different features describing a patient (medical history , race, weight, etc.), and targets y correspond to a quantity of interest to improve the patient health (optimal dosage of a particular drug, recovery time, depression lev el, etc.). When deploying a statistical model in healthcare applications, there are three well-defined stages: i) data collection, ii) feature selection, and iii) model learning. First, data collection refers to the process of gathering relev ant diagnostic data from patients, into a dataset of feature-target pairs D = { ( x i , y i ) } n i = 1 . Second, feature selection refers to the process of removing unnecessary features or attributes from each of the vectors x i . Third, model learning refers to the process of learning a statistical model f from the data D . These three processes are repeated as needed (collecting data, selecting features, learning models) until a suf ficient accuracy is achiev ed. W e call the process of learning a statistical model training time . W e call the process of using a fully trained statistical model to make predictions test time . B. Generalized Distillation Model compression [7] or distillation [24] are techniques to reduce the size of statistical models. Model distillation compresses large models f large by training a small model f small ( x ) that imitates the predictions of the lar ge model f large ( x ) . Remarkably , model distillation is often able to compress models without incurring any loss in accuracy [24]. Put in math, distillation assumes that the large model f large has been learned by minimizing Equation 1, and proceeds to learn the small model f small by minimizing ( 1 − λ ) L ( { ( x i , y i ) } n i = 1 , f small ) + λ L ( { x i , s i } n i = 1 , f small ) , (2) where λ ∈ [ 0 , 1 ] is an imitation parameter trading-off ho w much does the small model imitate the big model, versus directly learning the data. In (2) , a second dataset is introduced with targets s i = f large ( x i ) / T ; these are the softened predictions made by the large model. Here, T > 0 is a temperatur e parameter scaling the predictions of the large model 2 . Lopez-Paz et al. recently introduced generalized distillation , an extension of model distillation used to compress models built on a set of features into models built on a different set of features [39]. Generalized distillation is one specific instance of learning using privile ged information [13], [52], [53], a learning paradigm assuming that some of the features used to train a statistical model will not be av ailable at test time. More formally , generalized distillation assumes that a statistical 2 The temperature parameter was recently interpreted as a defense mechanism to adversarial data perturbations [44] and as a mechanism to increase the detection accuracy of security sensitiv e applications [11]. model will be trained on some data { ( x i , x ? i , y i ) } n i = 1 , and trained model will be tested on some data { x j } n + m j = n + 1 . Therefore, the set of features { x ? i } n i = 1 is av ailable at training but not at test time. Howe ver , these features may contain important information that would lead to statistical models of higher accuracy . For example, consider the case where x i is the image of a biopsy , y i ∈ {− 1 , + 1 } specifies if the tissue shown in the biopsy shows cancer and x ? i is the medical report of an oncologist. It is reasonable to assume that the medical report x ? i contains useful information to classify biopsy images x i , but such information will be unav ailable at test time. Generalized distillation [39] tackles the problem of learning without unav ailable data at test time as follo ws. First, it trains a model f ( x ) on the feature-target set { x ? i , y i } n i = 1 by minimizing Equation (1) . Second, it trains a second model f ( x ) by minimizing Equation (2) , where s i = f large ( x ? i ) / T . Therefore, it allows dev eloping objectives able to incorporate such sources of information into predictiv e models, without requiring them at test time. In the sequel, the features not av ailable at test time will correspond to undisclosed pri v ate features. Using these features, we will formulate a new v ariant of generalized distillation in a regression setting through patients’ pri v acy behaviors: we propose to use kno wledge extracted from redacted features of a patient across other databases and clinical studies to improv e model training when patients redact a different set of priv acy-sensitiv e information. I I I . P A T I E N T - D R I V E N P R I V A C Y In the introduction, we stated that patients desire to control data they wish to expose for use in healthcare statistical models. Using the language from Section II-A , in order to use a trained model f ( x ) for treatment, we must have values for all the entries comprising the feature vector x . Howe ver , in the healthcare domain, the vectors x often describe different types of patient data, and some of these entries may refer to genomic and clinical privac y . Therefore, patients may have different lev els of willingness to disclose such data. W e give examples of disclosure behavior of the patients in Appendix A. T o make predictions under redacted inputs, one may consider data imputation to fill the blanks with estimated values. Y et, imputation is impractical because it is either hard to apply under high dimensional redacted (missing) inputs or impossible if redacted inputs are statistically independent of the public (av ailable) inputs. A revie w of this literature is given in [14]. W e now start with technical prerequisites required for dev eloping a model that supports patient-driven priv acy . W e then de velop a mechanism for healthcare statistical models complying with the prerequisites, named privacy distillation . Patient-driv en privacy prer equisites- W e outline the charac- teristics required for a model to implement an effecti ve and complete patient-driv en mechanism: 1) Feature selection- Given a large pool of patient data; selection of relev ant clinical and genomic features is required for a model to improv e the accuracy . D a t a s e t & Ac q u i s i , o n & P a# e n t' g e n o m i c ' an d ' c l i n i c al ' i n f o r m a# o n ' Fe a t u r e & S e l e c , o n & S e l e c t' r e l e v an t' c l i n i c al ' f e atu r e s ' Pa ,en t &Cl u st eri n g& C l u s te r ' p a# e n t' p r o fi l e s ' b as e d ' o n ' r e d ac # o n ' b e h av i o r ' Le ar ning&thr ough&Ge ne r aliz e d&D is, lla, on& B u i l d ' a' d i s # l l e d ' m o d e l ' p e r ' p a# e n t' p r o fi l e ' F e atu r e s ' r e q u i r e d ' f o r ' p r e d i c # o n s ' A ' p a# e n t' r e d ac ts ' s o m e ' f e atu r e s ' Matc h ' w i th ' a' p r e c o m p u te d ' p a# e n t' p r o fi l e ' Mak e ' p r e d i c # o n s ' w i th ' p r e c o m p u te d ' dis#lle d'm ode l' B u i l d ' a' n e w ' m o d e l ' o r ' as s i g n ' a' p r o fi l e ' m ax i m al l y ' o v e r l ap p i n g ' n o n - r e d ac te d ' f e atu r e s ' Ye s' No ' Pred i c# o n s ' Genera,ng&Dis,lled&Models&at&Training&Time& Using&Dis,lled&Models&for&Treatment& Fig. 2: Privac y distillation process: (Left) how to learn distilled models based on redaction behaviors, (Right) how to use the distilled model to predict personalized treatments for new patients. 2) Control over type and amount of features- The model should consider patients’ desire to redact information that is deemed priv acy-sensiti ve. 3) Disclosure behavior of patients- Understanding the disclo- sure behavior of patients on redacted features is necessary to build efficient models. 4) Maintaining accuracy- The model under sufficient redacted features should retain the accuracy as almost as good as having complete patient data. 5) Patient safety- The prediction errors should not introduce any serious health-related risks. The rest of this section describes how we address these characteristics with priv acy distillation. A. Privacy Distillation Priv acy distillation is a mechanism to build healthcare statistical models that allow patients to define their own non- redacted and redacted features while providing them with safe and accurate personalized treatments. It comprises two main stages (see Figure 2). The first stage trains a statistical model that includes four steps: data acquisition, feature selection, patient clustering and learning via generalized distillation (Section III-A 1). The second stage addresses how to use the distilled model to predict personalized treatments for new patients. It includes two steps: assigning patients to a profile and making predictions (Section III-A2). Motivating Example- W e introduce the priv acy distillation through a simple example. Suppose we wish to treat a patient suffering from Post-T raumatic Stress Disorder (PTSD) with a patient profile “no sexual assault features”. A patient profile describes a set of patients providing all their features except their background about sexual assault. Pri v acy distillation proceeds as follows. As a first step, priv acy distillation defines a set of patient profiles. Each patient profile describes a dif ferent priv acy behavior . In the following, let us denote all the patient features by ( x 1 , x 2 , x 3 , x 4 ) , and ( x 1 , x 2 ) the sexual assault features. Thus, the patient profile “no sexual assault features” considers the features ( x 1 , x 2 ) as redacted, and the features ( x 3 , x 4 ) as non-redacted. Once defined, priv acy distillation builds a model for each patient profile. First, priv acy distillation builds a model ˆ y = f ( x 1 , x 2 , x 3 , x 4 ) that uses all the features av ailable across other patients and clinical studies to estimate the personalized treatment ˆ y . In particular , this model is built to predict the true clinical treatments y av ailable in some database. Second, priv acy distillation builds a model ˜ y = f ( x 3 , x 4 ) , this time only on non-redacted features. Howe ver , the model ˜ y = f ( x 3 , x 4 ) is built to learn the true clinical treatments y av ailable in our database, as well as to imitate the treatments ˆ y = f ( x 1 , x 2 , x 3 , x 4 ) pr edicted by the model that uses all the featur es . Finally , when a new patient belonging to the “no sexual assault features” profile comes for treatment, we can use the model ˜ y = f ( x 3 , x 4 ) to provide a personalized treatment, while preserving the desired lev el of priv acy . 1) T raining Distilled Models: This section presents the four steps of training distilled models: data acquisition, feature selection, patient clustering and learning distilled models. Data Acquisition- W e collect a set of feature-target pairs D = { ( x i , y i ) } n i = 1 , where x i ∈ R d is a vector of d features (both clinical and genetic) describing the i -th patient, and y i ∈ R is the target defined for a patient. In our experiments, we use real data collected on patients prescribed to warfarin from multiple medical institutions (see Section IV -B). Featur e selection- The raw patient data D may include multiple irrelev ant features by means of questionnaires, health history , lifestyle choices, etc.. These irrelev ant features contain no information about the problem of interest, and make the analysis more complicated due to the curses of high- dimensional data [29]. T o alleviate this issue, we preprocess the raw data D by discarding those features irrelev ant to the problem. In particular , we use Backwar d Attribute Elimination (B AE) [35], [38] algorithm which is found in pre vious studies to be highly effecti ve in the dataset we use. The B AE algorithm starts by building d − 1 statistical models, where the i -th statistical model is built using all the features except the i -th one, for all i = 1 , . . . , d . Next, the BAE algorithm chooses the model with the highest accuracy and decides that the feature that was excluded from that model is irrelev ant to the problem. The B AE algorithm proceeds then to build d − 2 statistical models, in search for the next irrelev ant feature. This process is repeated while the desired prediction accuracy is retained. W e perform B AE on clinical features since genetic features are often verified by experts. In the following, we denote by D the dataset obtained after data collection and feature selection. Patient Clustering- W e cluster our data D into K different patient profiles . Each patient profile describes a different disclosur e behavior : what features are understood as public, and what features are understood as priv acy-sensiti ve. The patient profiles are decided by analyzing the different disclosing behavior patterns found in the dataset. The k -th patient profile is a subset of D is defined as follows: D k = { ( x k i , x k , ∗ i , y k i ) } n i = 1 . In the previous, the vector ( x k i , x k , ∗ i , y k i ) contains the features of the i -th patient from the k -th patient profile. More specifically , x k i are non-redacted features, x k , ∗ i are redacted features, and y k i are targets. W e note that we find the disclosure behavior of users by examining the dataset. Howe ver , this process can be generalized to a user study to collect their priv acy behaviors. This might allow better identification of disclosure behavior of patients; thus the number precomputed patient profiles. W e defer conducting a user study to future work. Learning Distilled Models- W e no w introduce learning dis- tilled models, which is a technique we propose that improves model training under redacted features. Distilled models are adapted from generalized distillation procedure introduced in Section II-B , to suit our goal of improving model accuracy in the face of patients redacting priv ate features. Our intuition is that not all patients redact the same data; thus, non-redacted data can be acquired from other patients or controlled clinical studies. This is verified through analysis of patient data collected from multiple databases in Section IV -B . Therefore, we transfer the knowledge acquired from non- redacted data of other patients when a patient redacts data. W e tackle the problem of learning with redacted information as follows. First, we train a privileged model f priv ( x ) on the feature-target set { x ? i , y i } n i = 1 . Second, we train a distilled model f dist ( x ) . More precisely , given patient profiles, we build one model per patient profile, to obtain K distilled statistical models f 1 , . . . , f K . The objectiv e of the k -th distilled model is minimized as follows: predict data z }| { ( 1 − λ ) L ( { ( x k i , y i ) } n i = 1 , f k ) + imitate priv ate model z }| { λ L ( { x k i , f k , ∗ ( x k , ∗ i ) } n i = 1 , f k ) , (3) where the model f k , ∗ is learned on the set { ( x k , ∗ i , y k i ) } n k i = 1 . This, we call f k , ∗ a privile ged model , since it had access to the redacted features at training time. Therefore, the distilled model learns by simultaneously imitating the privile ged predictions of the pri vileged model and learning the tar gets of the original data. The objectiv e is independent of the learning algorithm; thus it can be minimized using arbitrary models (see Section V). Generalized distillation may ha ve a major problem when used for regression. This is because of the objectiv e for- mulation through a temperature parameter: L ( f ( x i ) , y i ) + λ L ( f ( x i ) , f ( x ∗ i ) / T ) . Consider a temperature parameter for T = 50, this would be L ( f ( x i ) , y i ) + λ L ( f ( x i ) , f ( x ∗ ) / 50 ) which is almost equal to: L ( f ( x i ) , y i ) + λ L ( f ( x i ) , 0 ) . Using the mean square error , the error is computed: ( f ( x i ) − y i ) 2 + λ f ( x i ) 2 . That is, it penalizes the large predictions, and drives predictions to zero; thus the use of temperature parameter may increase the error of the distilled model while treating patients (i.e., underes- timates the predictions). Therefore, we formulate the objecti ve 3 with use of imitation parameter λ in contrast to having both imitation and temperature parameters. In our formulation, the imitation parameter λ controls the trade-off between priv acy and accuracy . For λ ≈ 0 , the objectiv e (3) approaches the previously introduced standard objectiv e (1) (Section II-A ), which amounts to learning a model f k solely on non-redacted features. Ho we ver , as λ → 1 , the objectiv e (3) transfers the kno wledge acquired by the privile ged model f ∗ , k into the model f k . The intuition is that whenev er pri vileged model f k , ∗ makes a prediction error at patient x k i , the model f k should forget about getting that patient right, and focus on the rest of patients in that patient profile. These teachings from the privileged model do, in many cases, significantly help learning process of f k . T o conclude, we balance the priv acy and accuracy by using redacted features provided by external patients (i.e., not in that profile) when learning the model but ignoring such features when providing treatment to the patients. This is achieved by complementing the training of the model by allowing the transference of kno wledge about redacted attributes across patients and clinical studies. As a consequence, models trained with priv acy distillation do not require all data from patients and aims to retain the accuracy under redacted features close to that of complete features. 2) Using Distilled Models for T r eatment: At test time, a new patient x 0 decides what features to disclose, and what features to redact. W e then assign the patient x 0 to the profile k that maximally overlaps with respect to which features are disclosed (non-redacted features). Then, the distilled model f k associated with that profile is used to make a prediction. If none of the K patient profiles properly describes the disclosure behavior of the patient x 0 , we can adopt one out of two solutions to maximize the accuracy of her treatment. First, we can build a new patient profile D k + 1 and statistical model f k + 1 which mimics the disclosure behavior of the new patient. This solution would translate into a small computational ov erhead 3 at test time. Second, we can find an already built profile which discloses e very feature that the ne w patient discloses, but is more conservati ve about other features. This solution would translate into a potential loss of accuracy since not all of the disclosed features by the new patient would be used for treatment at test time. 3 In our experiments; we train 5K patient samples with 65 features on 2.6GHz 2-core Intel i5 processor with 8GB RAM less than one minute including optimal parameter search. Attribute Patient sample Demographic Race Black, Black or African American Ethnicity Both not Hispanic or Latino Age 50 - 59 (binned age in years) Gender male Background (clinical history) Height (cm) 161.29 W eight (kg) 86.1 Comorbidities no cardiomyopathy , no hyperlipidemia, no hypertension Medications not aspirin, not simvastatin, ... Macrolide antibiotics Y es V alve Replacement No Diabetes Not present Herbal medications, Y es vitamins, supplements Genotypic Cyp2C9 genotypes *3 VKORC1 SNP rs9923231 A/G Phenotypic Current Smoker Y es T ABLE I: Data used for personalized warf arin dose prediction. I V . E V A L U A T I O N W e now ev aluate priv acy distillation at the problem of warfarin dosing; wherein data is collected from a broad patient population to determine the proper personalized dose. W e start with basic clinical information essential to understanding warfarin, as well as revie w the current state-of-the-art models used to predict its personalized dosage (Section IV -A ). W e then describe experimental setup, including the data acquisition, the selection of features, the definition of patient profiles, the baseline models against which we will compare, and the ev aluation metrics that we care about (Section IV -B ). W e ev aluate the performance of priv acy distillation on the tension between priv acy and dose predictions under various patient profiles (Section IV -C ). Finally , we explore the tension between pri v acy and more tangible utility of patient health safety (Section IV -D). Our key findings are as follows: • Pri v acy distillation provides only 3% less accurate person- alized treatments than the state-of-the-art linear model that hav e access to the complete patient data, and 13.4% more accurate than the models that ignore pri v acy-sensiti ve data. • A loss in dose accuracy in healthcare models introduces ov er -and under-prescriptions that lead to stroke, embolism, and bleeding. Priv acy distillation increases the under- and ov er -prescriptions by 3.9% of the patients over the state-of- the-art linear model, whereas models constructed without patients’ sensitive features (i.e., public features) lead to 16.8% of the patients. • By using deep neural networks, we improve the accuracy of the state-of-the-art model by 2.7%. A. W arfarin Dose Algorithm W arfarin, known by the brand name Coumadin, is a widely prescribed (over 20 million times each year in the United States) oral anticoagulant used to prevent blood clots from ." ." ." Patient Profiles ." ." ." redacted' non-redacted' T raining Distilled Models Model 1 Patient Profiles Model n Fig. 3: Priv acy distillation process for pri v acy behaviors observed in multiple dataset. forming mostly in the heart and lungs. Unfortunately , wrong dosages of warfarin may lead to serious and even fatal adverse consequences. On the one hand, under-prescriptions of warfarin will render the treatment useless. On the other hand, over - prescriptions of warfarin may cause hemorrhagic strokes and major bleeding [1]. Current practice suggests fixing the initial dose to 5 or 10 milligrams per day [49]. Then, patients undertake regular blood tests to measure how long it takes for blood clots to form. This measure is referred to as the International Normalized Ratio (INR). The subsequent warfarin dosages are then designed to keep the patient’ s INR constant and within the desired lev els. The International W arfarin Pharmacogenetics Consortium (IWPC) performed a study on a large and div erse patient pop- ulation prescribed to warfarin, for the purpose of determining rules for an accurate personalized warfarin dosage [30]. The study concluded one of the largest and most comprehensive datasets to date for ev aluating personalized warfarin. A long line of work culminated on a linear pharmacogenetic model that outperformed all previous approaches to personalized warfarin dosage prediction 4 . The model combines genotypic, demo- graphic, phenotypic, and background patient information [30], [31], [46]. The genotypic information concerns genes VKORC1 and CYP2C9. While the VKORC1 gene encodes the target enzyme of warfarin (vitamin K), the CYP2C9 gene has an effect on warfarin metabolization [18]. The demographic and phenotypic information concern various patient characteristics and habits, such as their smoking status. The background information in volv es the clinical history of a patient, including medications that may interfere their INR. All of this information highlights the variability across patients; thus the necessity of personalized treatment [46]. B. Experimental Setup In the following, we describe the steps used to set up our experiment for warfarin dose prediction. This section parallels the steps described in Section III. Data Acquisition- W e use previously introduced IWPC dataset [30]. The dataset is collected from 21 medical organiza- tions from nine countries and four continents. The y are located in T aiwan, Japan, K orea, Singapore, Sweden, Israel, Brazil, United Kingdom, and the United States. The data collection 4 The algorithm has been given to doctors and other clinicians for predicting ideal dose of warfarin. 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Attribute index Demographic Background Phenotypic Genotypic 1 " 2 " 5 " 3 " 4 " 6 " 7 " 8 Fig. 4: An exploration of the features categories and ranking of the each feature. (Refer to the text for feature definitions.) from multiple sources allows to analyze the priv acy behavior of patients and to use the non-redacted features in model training required for priv acy distillation. For instance, when a patient redacts a set of features, these features are found out to be non-redacted in the dataset of other organizations. This process is illustrated in Figure 3 and explained throughout this section. The dataset includes clinical and genotypic data of patients. T able I presents an example of features collected from a patient. W e split the clinical data into three categories of six demographic, 24 background, one phenotypic feature. W e represent two genotypic features with genetic variants of CYP2C9 as combinations of *1, *2, *3 and VK OCR1 with single-nucleotide rs9923231 of G/G, A/G or A/A genotypes. W e split the data into two cohorts; the training cohort is used to define patient profiles and to learn the distilled models, and the validation cohort is used to ev aluate the accuracy of the distilled models and the errors introducing health-related risks. W e conv ert the categorical variables into numeric factors and standardize features by removing the mean and scaling to unit variance. Overall, the cohorts used in experiments consist of 1221 training and 656 validation samples in which each patient is described by 33 features of four categories, and has warfarin dose as a target. Featur e Selection- W e apply BFE algorithm using least- squares regression (as in reference paper [30]) to infer the relev ant features for warfarin dosing. W e illustrate the 33 selected features and their categories in Figure 4. The x-axis shows the feature numbers of each category and y-axis is the name of these categories. The most relev ant features are numbered in circles and found to be the primary predictors: Demographic feature of 4 race; genotypic features of 2 Cyp2C9 and 3 VK ORC1; background features of 1 weight, 5 use of rifampicin (an antibiotic to treat or prev ent a several types of bacterial infections), 6 medications (list of medicines taken), and 7 use of Cordarone (used for treatment of irregular heartbeats), and phenotypic feature of 8 current smoker status. Patient Profiles- W e define three patient profiles based on what information patients disclose, with respect to the four categories of demographic, background, phenotypic, and geno- typic information (see T able II). First, public patients disclose Patient data categories Legend: 3 non-redacted, 7 redacted Demographic Background Phenotypic Genotypic Public patient Pharmacogenetic algorithm 3 3 3 3 W ith all except demographic 7 3 3 3 Closed W ith all except background 3 7 3 3 patient W ith all except phenotypic 3 3 7 3 W ith all except genotypic 3 3 3 7 Demographic except others 3 7 7 7 Strict Background except others 7 3 7 7 patient Phenotypic except others 7 7 3 7 Genotypic except others 7 7 7 3 T ABLE II: Patient profiles used in experiments. the information relating to the four categories. Second, closed patients disclose the information relating to three out of the four categories. Third, strict patients disclose the information relating to only one out of the four categories. For instance, a closed patient refers to “with all except genotypic” does not expose genotypic features of VKORC1 and CYP2C9, yet discloses the others, and a strict patient refers to “genotypic except others” exposes genotypic features, but nothing else. Privacy Distillation Implementation- W e implement priv acy distillation using a deep neural network (DNN) with a hidden layer of 32 rectified linear units each, using the Keras libary [15] running the Theano backend [8]. W e use the Adam optimizer to minimize mean squared error loss function [34]. This setup performance is consistent with more complex architectures that hav e ev aluated with the warfarin dataset and allows us to implement network training as well as the distilled models efficiently . In our experiments, we show the results of pri v acy distillation with an imitation parameter values of λ ∈ [0,1]. Non-redacted demographic background phenotypic genotypic 0 20 20 30 40 50 60 70 80 90 100 110 120 130 140 mean error (MAPE) (%) Linear model DNN model closed patient profiles Fig. 5: W arfarin dose prediction performance measured against linear pharmacogenetic and deep neural network models on a subset of dataset. (Strict patient profiles are similar to closed patient profiles.) Evaluation Metrics- W e ev aluate the accuracy of models using Mean Absolute Error (MAE) and Mean Absolute Percentage Error (MAPE). These metrics are easily interpreted statistics which is a measure of how far predicted values are away from clinically-deduced warfarin dose and accuracy as a percentage of the error . They are widely used in prediction of real-valued outputs in medical treatments [30]. The lower values of MAE, the closer estimates to the actual dose, indicate better quality of treatment. For each experiment, we report the warfarin dosage 0 0.2 0.4 0.6 0.8 1 imitation parameter ( λ ) 10.5 11 11.5 12 12.5 13 13.5 mean absolute error (mae) (a) with all except demographic partially-redacted (DNN) non-redacted (DNN) non-redacted (pharmacogenetic algorithm) distilled model (DNN) 0 0.2 0.4 0.6 0.8 1 imitation parameter ( λ ) 10.9 11 11.1 11.2 11.3 11.4 11.5 11.6 mean absolute error (mae) (b) with all except background 0 0.2 0.4 0.6 0.8 1 imitation parameter ( λ ) 10.5 11 11.5 12 12.5 13 13.5 14 14.5 15 mean absolute error (mae) (c) with all except phenotypic 0 0.2 0.4 0.6 0.8 1 imitation parameter ( λ ) 10 11 12 13 14 15 16 17 mean absolute error (mae) (d) with all except genotypic Closed patient profiles 0 0.2 0.4 0.6 0.8 1 imitation parameter ( λ ) 10.8 11 11.2 11.4 11.6 11.8 12 12.2 mean absolute error (mae) (e) demographic except others 0 0.2 0.4 0.6 0.8 1 imitation parameter ( λ ) 10.5 11 11.5 12 12.5 13 13.5 14 14.5 15 mean absolute error (mae) (f) background except others 0 0.2 0.4 0.6 0.8 1 imitation parameter ( λ ) 10.5 11 11.5 12 12.5 13 13.5 mean absolute error (mae) (g) phenotypic except others 0 0.2 0.4 0.6 0.8 1 imitation parameter ( λ ) 10.85 10.9 10.95 11 11.05 11.1 11.15 11.2 11.25 mean absolute error (mae) (h) genotypic except others Strict patient profiles Fig. 6: Comparison of priv acy distillation with non-redacted and partially-redacted models. Std. dev . of ten runs falls in on av erage ± 1.7 (see Section IV -D for medical consequences of models to make dosing decisions). MAE and MAPE when a veraged ov er ten random training- validation cohort splits. Models for Comparison- W e e v aluate the accuracy of priv acy distillation under the patient profiles against two baselines: P artially-redacted , which is a model trained only on the information disclosed by the patients, and non-r edacted model is trained on the complete patient data. T o build a non-redacted model, we reimplement the pharmacogenetic algorithm in Python by direct translation from the authors’ implementa- tion [30]. Furthermore, we implement a deep neural network model with the same architecture used in priv acy distillation. W e find that DNN model gives on a verage 2.7% more accurate dose predictions than the current non-redacted linear model. W e go on and compare the model performance on partially-redacted models; similar improvements hold both for closed and strict patient profiles (see Figure 5). T o make fair comparisons, we use DNNs to implement the three models: priv acy distillation, partially-redacted and non-redacted. C. Results on Dose Accuracy W e e v aluate the accuracy of pri v acy distillation against partially-redacted (dashed line), non-redacted linear pharma- cogenetic algorithm (dotted line), and non-redacted model of DNN (dashed-dotted line). All experiments are performed on pre viously described patient profiles. T able III summarizes the av erage results of errors and Figure 6 presents the MAE of priv acy distillation with an imitation parameter λ ∈ [0,1]. Non-redacted model MAE MAPE (%) Public Pharmacogenetic algorithm 11.2 39.3 patient Our implementation (DNN) 10.9 36.6 Partially-redacted (DNN) Distilled (DNN) MAE MAPE (%) MAE MAPE (%) W ith all except demographic 13.3 44.6 11.4 39.7 Closed With all except background 11.0 37.1 11.0 37.0 patient W ith all except phenotypic 14.8 50.4 12.3 43.5 W ith all except genotypic 16.3 56.5 13.5 46.8 Demographic except others 12.2 41.8 12.0 40.8 Strict Background except others 14.3 49.1 13.3 45.5 patient Phenotypic except others 13.3 48.5 13.3 48.5 Genotypic except others 11.1 37.3 10.9 36.6 T ABLE III: Prediction performance of models for warfarin dose under patient profiles. W e note that partially redacted models implemented with pharmacogenetic algorithm performs on average 13.4% worse than pri v acy distillation. W e first ev aluate the effecti veness of priv acy distillation on closed patient profiles. As shown in Figure 6(a-d), privacy distillation with a car efully chosen λ outperforms the partially- r edacted model, and achieves a similar accuracy to the non- r edacted model. For instance, illustrated in Figure 6(a), “with all except demographic” is a profile in which a patient does not disclose her demographic features. Assume for now that warfarin dose prediction is possible only using non-redacted features of patients by using the partially-redacted model at test time. The error yields MAE of 13.3; Howe ver , if a patient provides complete data, the error yields 10.9. This indicates 8% less accurate dose predictions than the non-redacted model. As we detail in Section IV -D , this difference significantly increases various medical side effects. No w , we make warfarin dose predictions by distilled models. Returning to Figure 6(a), when a patient redacts the demo- graphic information, the distilled model yields MAE of 11.4. This is exactly 3.1% mean absolute error less than the non- redacted model compared to 8% of the partially-redacted model. As seen in Figure 6(b-c), the results of other closed patient profiles are similar , and both retain dose errors similar to the non-redacted model, and always better than the partially- redacted model. It is important to note that the impact of using models implemented with DNN over linear models is significant. As shown in Figure 6(b), a patient with only partially-redacted features giv es better results than the original linear method with all features. In addition, we find that when a patient does not disclose genotypic features, the increase in distilled model errors is more observ able than other patient profiles (Figure 6(d)) as these features are the ke y variables for warfarin dose predictions (see Section III-A1). A similar analysis on the strict patient profiles confirms the tensions between accuracy and the amount of information disclosed by patients (Figure 6(e-h)). The benefit of distilled models similar to the closed patient profiles. Howe ver , we make two additional important remarks here. First, patients within the “phenotypic except other” would solely disclose their “smoking status”. This results in a very simplistic model with limited information, and yields higher errors than other patient profiles (Figure 6(g)). Second, the importance of genotypic features becomes more apparent when using deep neural networks, instead of the original linear pharmacogenetic algorithm (Figure 6(h)). That is, the genetic information of patients better generalizes the nonlinear DNN distilled model with the patients used in our experiments and assigns slightly better dosages than the non-redacted model at λ =0.1. Overall, the benefit of priv acy-distillation is consistent across all patient profiles. On average, priv acy-distillation offers dose predictions 5.7% and 3.9% less accurate than those provided by the non-redacted DNN and state-of-the-art linear pharmacogenetic model. Howe ver , it is necessary to study the impact of errors introducing health risks as we discuss next. D. Results on P atient Health In this section, we ask the essential question: Does making dose predictions with priv acy distillation introduce health- related risks? T o answer this question, we present a study to ev aluate the clinical relev ance of dose errors when patients redact features. The study aims at analyzing the dose errors that are inside and outside of the warfarin safety window , and the medical side effects of under- or over prescriptions. Overview- W e design a study to validate the clinical value of priv acy distillation on warfarin dose. The study aims at ev aluating the dose errors of partially-redacted, non-redacted models and pri v acy distillation. W e consider weekly dosage errors for each patient because using weekly values eliminates Valida&on)cohort:)656)pa&ents)from)Interna&onal) Warfarin)Pharmacogene&c)Consor&um)Dataset) Pharmacogene&c)) algorithm) Pa&ent-defined) model) Privacy)) dis&lla&on) )))Clinical)relevance) Enrollment) Ini&al)dose)predic&on))) Weekly)predic&ons) Magnitude)of)errors)associated)in)pa&ent)health) ))20%) ))Underes&ma&on) Overes&ma&on) )))))Safety)window) Fig. 7: Overvie w of study designed to ev aluate the clinical v alue of pri v acy distillation. Details of clinical relev ance and definition of safety windo w are giv en Section IV -D1. the errors posed by the initial (daily) dose. For instance, a small initial dose error may incur more health risks ov er time (see discussion for details). W e follow the design of the clinical relev ance implemented in [30] which is to date the lar gest completed warfarin analysis. The authors assess the pharmacogenetic algorithm over clinical and fixed-dose algorithms, yet we measure the model efficacy on making dose predictions. W e define two arms to get the weekly dosages: 1) Pharmacogenetic: Use of a DNN implementation of the pharmacogenetic algorithm with complete patient data. 2) Priv ate: Use of priv acy distillation and partially-redacted models. These models are identical to the pharmacogenetic arm; howe ver , we only replace the pharmacogenetic algo- rithm with priv acy distillation and partially-redacted model under different patient profiles. Recall that pharmacogenetic algorithm (non-redacted model) is the current best-known algorithm for initial dose prediction ov er clinical and fixed-dose approach. 1) P atient Safety: T o ev aluate the impact of errors that introduce health-related risks, we draw patient samples from v alidation cohort and assign them weekly warfarin dosages with the use of pharmacogenetic and priv acy arms (see Figure 7). W e then measure the errors between the estimated dose and clinically-deduced ground truths. These errors define ho w good arms are on the prediction of dosage for a particular patient. The direction of the error comes with a positive or negati ve sign that points out the under or ov er-prescribes. W e use this value for finding the side effects as detailed next. Safety window- W e calculate the percentage of patients whose predicted dose of warfarin is within a therapeutic safety windo w . The weekly dose is accepted in the safety window if an estimated dose falls within 20% of its corresponding clinically- deduced ground truth. This is because the v alue defines the dif ference of 1 mg per day relati ve to the fixed starting dose of Non-redacted model Legend U (%) SW (%) O (%) U: Underestimation Public Pharmacogenetic algorithm 21.8 40.9 5 37.3 SW : Safety Windo w patient Our implementation (DNN) 24.9 42.3 33.8 O: Overestimation Partially-r edacted model (DNN) Privacy distillation (DNN) U (%) SW (%) O (%) U (%) SW (%) O (%) W ith all except demographic 28.2 36.1 35.7 22.8 40.4 36.8 Closed W ith all except background 23.5 42.0 34.5 23.4 42.3 34.3 patient W ith all except phenotypic 30.9 33.0 36.1 23.3 36.5 40.2 W ith all except genotypic 31.3 30.1 38.6 27.7 33.2 39.1 Demographic except others 24.0 37.4 38.6 25.1 37.8 37.1 Strict Background except others 28.9 32.4 38.7 27.5 34.6 37.9 patient Phenotypic except others 26.8 31.0 42.2 26.2 31.7 42.1 Genotypic except others 20.5 39.2 40.3 20.1 39.8 40.1 T ABLE IV: Percentage of patients in validation cohort that are within the safety windo w , underestimated or ov erestimated against weekly dosages of clinically-deduced dosages. Std. dev . of ten independent runs falls in on average ± 2.5. Partially redacted models implemented with pharmacogenetic algorithm perform on average 24.1% of the patients in the safety window . 5 mg per day which is often accepted as clinically relev ant [30], [33]. The deviations of both directions fall outside of the safety window . Specifically , any dose prediction is an ov erestimation that is at least 20% higher than the actual dose and is an underestimation that is at least 20% lower than the actual dose. Health risks- T o identify the side effects of warfarin, we follo w U.S. Food and Drug Administration (FD A) warfarin medical guide for the most common side effects of ov erestimation and underestimation [20]. T aking high-dose warfarin causes INR to be high (thin blood) which causes higher risks of having intracranial and extracranial bleeding. T aking low doses causes INR to be low (thick blood), and warfarin does not protect patients from dev eloping a blood clot, and it causes embolism and stroke. In both cases, extremely high or low doses may lead to death. W e note that certain amount of dosing errors, in reality , do not immediately cause these adverse effects, as factors such as commonly used medications and foods with vitamin K also play a role in blood clotting. 2) Clinical Relevance: W e now ev aluate the health-related risks of making dose decisions under patient profiles with partially-redacted, non-redacted and priv acy distillation models. T able IV shows the percentage of patients within the dose safety window , under-prescriptions, and over -prescriptions. Remarkably , privacy-distillation yields nearly-optimal weekly dose err ors, when compar ed to non-r edacted DNN implemen- tation of a pharmacogenetic algorithm that assumes access to complete patient information . It yields on av erage 5.3% fewer patients in the safety window . For instance, “with all except demographic” closed patient profile gi ves 22.8% under- prescribes, 40.4% in the safety windo w and 36.8% over - prescribes. This value is only 1.9% less than the non-redacted model and 4.3% more than the results of the partially-redacted model. It is interesting to report that the direction of the error changes. The percentage of overestimated patients is increased from 33.8% to 36.8% and underestimated patients is decreased from 24.9% to 22.8%. Analyzing the results more in detail, we find that some patient profiles yield a lower percentage of patients in the safety windo w compared to other profiles. “W ith all except genotypic” and “Phenotypic except others” patient profiles yield 33.2% and 31.7% of the patients in the safety window compared to the 42.3% of public patients. As discussed in Section IV -C , the high amount of information disclosed by patients such as using only phenotypic feature or disclosure of important v ariables such as genotypic features causes loss of information in model learning. Thus, over - and under-prescriptions become more apparent than other patient profiles. Howe ver , pri vacy distillation always performs better treatments than those of partially-redacted models. T o conclude, pri v acy distillation gives nearly optimal dosage error across all patient profiles. It offers 5.3% and 3.9% fewer patients in the safety window than those provided by the non-redacted DNN and state-of-the-art linear pharmacogenetic model. In turn, it is effecti ve at pre venting errors that introduce health-related risks. V . D I S C U S S I O N W e presented privac y distillation in a regression setting. Howe ver , its objecti ve can be formulated to other statistical models such as supervised, semi-supervised, transfer and univ ersum learning [39]. In these, two important factors need to be addressed to maximize the accuracy: Model selection and parameter tuning. For the former , the objectiv e function can be minimized using arbitrary models, as pri v acy distillation is a model free mechanism. For the latter , while minimizing the objecti ve, the imitation parameter λ should be chosen carefully for finding the optimal distilled model. It is important to note that our study on predicting warfarin dosages makes the evaluation of priv acy distillation different than those solely based on raw accuracy . Ho we ver , there exist more comprehensiv e studies such as clinical trials to ev aluate 5 Note: W e remark that our results are similar to those found in reference paper [30]. These results represent the current best-known pharmacogenetic method for predicting initial warfarin dose, and often giv es more accurate dose prescriptions than current clinical practices as difficulty in establishing an initial dose of warfarin varies by a factor of 10 among patients [30], [32]. long-term health risks. For instance, it is common to observe warfarin dose by subsequently titrating to 90 days by using pharmacokinetic/pharmacodynamics (PK/PD) models, as dose titration after one-week period may change the dosages and pose varying health risks [3]. Howe ver , long-term studies require observation of INR responses because INR lev els are not stable during this period. The other crucial point for an ev aluation of errors is the models for risk estimation. There exist advanced models to estimate an adverse ev ent occurring in a specific period or as a result of a specific situation. For instance, varying INR lev els of a patient can be used to estimate the correlation between stroke and bleeding event [50]. These models define more detailed health risks through various measures of the blood clot. Howe ver , the input/output of the model components interacts with detailed clinical patient information for precise estimation. Priv acy distillation can be easily integrated with more exhausti ve clinical trials under these models. V I . R E L A T E D W O R K Privacy-sensiti ve Information- Clarke have categorized the types of priv acy and outlined specific protections [16]. The categories include the pri v acy of the person, priv acy of personal behavior , the priv acy of personal data, and priv acy of personal communication. Pertaining to these categories, recent studies hav e analyzed the individuals’ behavior of information disclosure both in online and offline contexts[4], [41]. The common point of the studies is individuals’ context-dependent preferences and the subjectivity on the personal matters. W e solve these problems with one of our implementation goal of making predictions a v ailable under v arying number of redacted inputs of patients (i.e., patient profiles). The Law and Medical Privacy- The goal of medical priv acy is keeping information about a patient confidential. This in volves both conv ersational discretion and medical record security . The Health Insurance Portability and Accountability Act (HIP AA) priv acy and security rules [25], [45] is the baseline law that protects patient medical information. There are also recent laws dealing with genetic information priv acy [28]. The goal of these regulations is to establish the rights individuals hav e concerning their health information. Pri vacy distillation formalizes this goal and addresses the patient control ov er priv acy-sensiti ve information under these regulations. Healthcare Statistical Models- In recent years, researchers hav e extensi vely in vestigated healthcare models. Futoma et al. used random forests and DNNs for predicting early hospital readmissions [23]. V olm et al. developed in vitr o tests to predict tumors’ drug responsiveness for cancer treatment [54]. Other researchers in pharmacogenomics predicted the dose of the medicines and their responses on patients [6], [9]. These models can be equipped with priv acy distillation when a set of features is identified as priv acy-sensiti ve. In addition, priv acy distillation handles all stages of modeling, including feature selection and redaction behavior of patients. Privacy Threats to Patient Information- Priv acy threats target obtaining sensitiv e patient information from models or databases. Homer et al. used the public allele frequencies to infer the possibility of the participation of an individual in a genotype database [27]. W ang et al. showed that statistics on genetics and diseases could be used to identify individuals [55]. Fredrikson et. al. introduced a model in version attack and used same warfarin dataset to predict a patient’ s genetic markers from some clinical data and warfarin dosages [22]. These approaches consider dif ferent utility/priv acy metrics, as they assume that patients disclose all of their information and attempt to infer sensiti ve information by studying the relationships between inputs and outputs of the model. Defense of Privacy Leaks- As a response to the priv acy threats, researchers suggest prevention and remedies to such breaches [12], [17], [40]. These approaches are limited to protecting the priv acy of users on inputs required for training or classification. W e view our efforts in this paper to be complementary to much these. Priv acy distillation can be easily integrated as a user-driv en mechanism to strike a balance between accuracy and redacted inputs of users. V I I . C O N C L U S I O N W e proposed a patient-driv en priv acy mechanism named pri v acy distillation. It is a learning meta-algorithm to construct accurate healthcare statistical models that allo w patients to con- trol their priv acy-sensitiv e data. W e ev aluated the effectiv eness of priv acy distillation on pharmacogenetic modeling of person- alized warfarin dose. In our experiments, priv acy distillation outperformed the state-of-the-art healthcare statistical models that ignore priv acy-sensitiv e data. By reusing knowledge about pri v acy-sensiti ve information across patients, pri vacy distillation showed nearly optimal warfarin prescriptions, competing with the idealized models that assume access to complete patient data. W e analyzed the impact of the accuracy gains provided by priv acy distillation on patient health; this showed a significant reduction in warfarin under- and over -prescriptions. Such accurate predictions can translate into less health-related risks of embolism, strokes, or bleeding. This work is the first effort at developing models under redacted users inputs. The capacity afforded by this approach will allow us to make accurate predictions in a wide array of applications requiring priv ate inputs, such as the ones found in medicine, law , forensics, and social networks. In the future, we will explore a wide range of en vironments and ev aluate the ability of priv acy distillation to promote its effecti veness. V I I I . A C K N O W L E D G M E N T Research was sponsored by the Army Research Laboratory and was accomplished under Cooperative Agreement Number W911NF-13-2-0045 (ARL Cyber Security CRA). The views and conclusions contained in this document are those of the authors and should not be interpreted as representing the official policies, either expressed or implied, of the Army Research Laboratory or the U.S. Government. The U.S. Government is authorized to reproduce and distribute reprints for Government purposes notwithstanding any copyright notation here on. R E F E R E N C E S [1] N. S. Abraham et al. Comparativ e risk of gastrointestinal bleeding with dabigatran, ri varoxaban, and warfarin: population based cohort study . British Medical Journal , 2015. [2] A. Acquisti and J. Grossklags. Privac y and rationality in individual decision making. IEEE Security & Privacy , 2005. [3] J. L. Anderson et al. Randomized trial of genotype-guided vs standard warfarin dosing in patients initiating oral anticoagulation. Circulation , 2007. [4] A. Appari and M. E. Johnson. Information security and priv acy in healthcare: current state of research. International journal of Internet and enterprise management , 2010. [5] P . S. Appelbaum. Privac y in psychiatric treatment: threats and responses. F ocus , 2003. [6] M. J. Arranz et al. Pharmacogenetics of the efficacy of antipsychotic drugs in schizophrenia. In Genetic Influences on Response to Drug T reatment for Major Psychiatric Disorders , 2016. [7] J. Ba and R. Caruana. Do deep nets really need to be deep? In Advances in neural information processing systems , 2014. [8] J. Bergstra et al. Theano: A cpu and gpu math compiler in python. In Python for Scientific Computing Conference (SciPy) , 2010. [9] J. Biernacka et al. The international SSRI pharmacogenomics consortium: a genome-wide association study of antidepressant treatment response. Natur e T ranslational psychiatry , 2015. [10] J. R. Carpenter and M. G. Kenward. Missing data in clinical trials-a practical guide . John Wile y & Sons, 2008. [11] Z. B. Celik et al. Extending detection with forensic information. arXiv:1603.09638 , 2016. [12] Z. B. Celik et al. Curie: Policy-based secure data exchange. arXiv:1702.08342 , 2017. [13] Z. B. Celik et al. Feature cultiv ation in privile ged information-augmented detection. In International W orkshop on Security And Privacy Analytics (in vited paper) . A CM, 2017. [14] J. Chen, X. Liu, and S. Lyu. Boosting with side information. In Asian Confer ence on Computer V ision , 2012. [15] F . Chollet. Keras: Theano-based deep learning library . Code: https://github .com/fchollet , 2015. [16] R. Clarke. Information technology and dataveillance. ACM Communica- tions , 1988. [17] F . K. Dankar and K. El Emam. The application of differential pri vac y to health data. In ACM ICDT , 2012. [18] L. Dean. W arfarin therap y and the genotypes c yp2c9 and vkorc1. National Center , Biotechnology Information , 2013. [19] Standards for pri vac y of individually identifiable health information; final rule, 2000. [20] Medication guide, caumadin (warfarin sodium). http://www .fda.gov. [Online; accessed 11-January-2017]. [21] The presicion medicine initiative. https://obamawhitehouse.archiv es.gov/ precision- medicine. [Online; accessed 11-February-2017]. [22] M. Fredrikson et al. Priv acy in pharmacogenetics: An end-to-end case study of personalized warfarin dosing. In USENIX Security , 2014. [23] J. Futoma, J. Morris, and J. Lucas. A comparison of models for predicting early hospital readmissions. J ournal of biomedical informatics , 2015. [24] G. Hinton, O. V inyals, and J. Dean. Distilling the knowledge in a neural network. arXiv pr eprint arXiv:1503.02531 , 2015. [25] Hipaa privac y rule and public health. Centers for Disease Control and Prev ention, 2003. [26] T . C. Holdeman. In visible wounds of war: Psychological and cognitive injuries, their consequences, and services to assist recovery . Psychiatric Services , 2009. [27] N. Homer et al. Resolving indi viduals contributing trace amounts of dna to highly complex mixtures using high-density snp genotyping microarrays. PLoS Genet , 2008. [28] K. L. Hudson et al. Keeping pace with the times-the genetic information nondiscrimination act of 2008. New England Journal of Medicine , 2008. [29] M. Hund et al. Analysis of patient groups and immunization results based on subspace clustering. In International Conference on Brain Informatics and Health , 2015. [30] International W arfarin Pharmacogenetics Consortium. Estimation of the warfarin dose with clinical and pharmacogenetic data. The New England journal of medicine , 2009. [31] F . Kamali and H. W ynne. Pharmacogenetics of warfarin. Annual revie w of medicine , 2010. [32] S. E. Kasner et al. W arfarin dosing algorithms and the need for human intervention. The American journal of medicine , 2016. [33] S. E. Kimmel et al. A pharmacogenetic versus a clinical algorithm for warfarin dosing. New England Journal of Medicine , 2013. [34] D. Kingma and J. Ba. Adam: A method for stochastic optimization. arXiv preprint arXiv:1412.6980 , 2014. [35] R. K ohavi and G. H. John. Wrappers for feature subset selection. Artificial intelligence , 1997. [36] L. K oontz et al. Information Privacy in the Evolving Healthcar e En vir onment . 2013. [37] Y . LeCun, Y . Bengio, and G. Hinton. Deep learning. Natur e , 2015. [38] J. Li, K. Cheng, et al. Feature selection: A data perspective, 2016. [39] D. Lopez-Paz, L. Bottou, B. Sch ¨ olkopf, and V . V apnik. Unifying distillation and pri vileged information. International Conference on Learning Representations (ICLR) , 2016. [40] P . J. McLaren et al. Privac y-preserving genomic testing in the clinic: a model using HIV treatment. Genetics in Medicine , 2016. [41] M. Naveed et al. Priv acy in the genomic era. Computing Surveys , 2015. [42] C. D. Newgard and R. J. Lewis. Missing data: how to best account for what is not known. JAMA , 2015. [43] B. Obama. United states health care reform: progress to date and next steps. JAMA , 2016. [44] N. Papernot, P . McDaniel, X. W u, S. Jha, and A. Swami. Distillation as a defense to adversarial perturbations against deep neural networks. IEEE Security and Privacy , 2016. [45] Health insurance portability and accountability , 1996. [46] A. H. Ramirez et al. Predicting warfarin dosage in european-americans and african-americans using dna samples linked to an electronic health record. Pharmacogenomics , 2012. [47] P . H. Rezvan, K. J. Lee, and J. A. Simpson. The rise of multiple imputation: a review of the reporting and implementation of the method in medical research. BMC medical r esear ch methodology , 2015. [48] M. Saar-Tsechansky and F . Provost. Handling missing values when applying classification models. machine learning resear ch , 2007. [49] A. J. Shepherd et al. What is the best starting dose of warfarin when initiating therapy? Evidence Based Practice , 2015. [50] S. V . Sorensen et al. Cost-effecti veness of warfarin: trial versus real-world stroke prevention in atrial fibrillation. Heart Journal , 2009. [51] R. T ourangeau and T . Y an. Sensitive questions in surveys. Psychological Bulletin , 2007. [52] V . V apnik and R. Izmailov . Learning using privileged information: Similarity control and knowledge transfer . J ournal of Machine Learning Resear ch , 2015. [53] V . V apnik and A. V ashist. A ne w learning paradigm: Learning using privile ged information. Neural Networks , 2009. [54] M. V olm and T . Efferth. Prediction of cancer drug resistance and implications for personalized medicine. F r ontiers in oncology , 2015. [55] R. W ang et al. Learning your identity and disease from research papers: information leaks in genome wide association study . In ACM Computer and communications security , 2009. A P P E N D I X A. Privacy Concerns of P atients W e provide some examples of patients redacting their data due to the priv acy concerns. W e refer the reader to K oontz [36] for a more comprehensiv e discussion of priv acy concerns of patients and their reasons. • U.S. Department of Health & Human Services (HHS) estimated that 2M Americans did not seek treatment for mental illness [19]. • Millions of young Americans that suffers from sexually transmitted diseases do not seek treatment [19]. • HHS estimated that 586K Americans did not seek earlier cancer treatment [19]. • The Rand Corporation found that 150K soldiers suffering from Post-T raumatic Stress Disorder (PTSD) do not seek treatment [26]. Pri vac y concerns contribute to the highest rate of suicide among activ e duty soldiers in 30 years.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment