Multi-radial LBP Features as a Tool for Rapid Glomerular Detection and Assessment in Whole Slide Histopathology Images
We demonstrate a simple and effective automated method for the segmentation of glomeruli from large (~1 gigapixel) histopathological whole-slide images (WSIs) of thin renal tissue sections and biopsies, using an adaptation of the well-known local binary patterns (LBP) image feature vector to train a support vector machine (SVM) model. Our method offers high precision (>90%) and reasonable recall (>70%) for glomeruli from WSIs, is readily adaptable to glomeruli from multiple species, including mouse, rat, and human, and is robust to diverse slide staining methods. Using 5 Intel(R) Core(TM) i7-4790 CPUs with 40 GB RAM, our method typically requires ~15 sec for training and ~2 min to extract glomeruli reproducibly from a WSI. Deploying a deep convolutional neural network trained for glomerular recognition in tandem with the SVM suffices to reduce false positives to below 3%. We also apply our LBP-based descriptor to successfully detect pathologic changes in a mouse model of diabetic nephropathy. We envision potential clinical and laboratory applications for this approach in the study and diagnosis of glomerular disease, and as a means of greatly accelerating the construction of feature sets to fuel deep learning studies into tissue structure and pathology.
💡 Research Summary
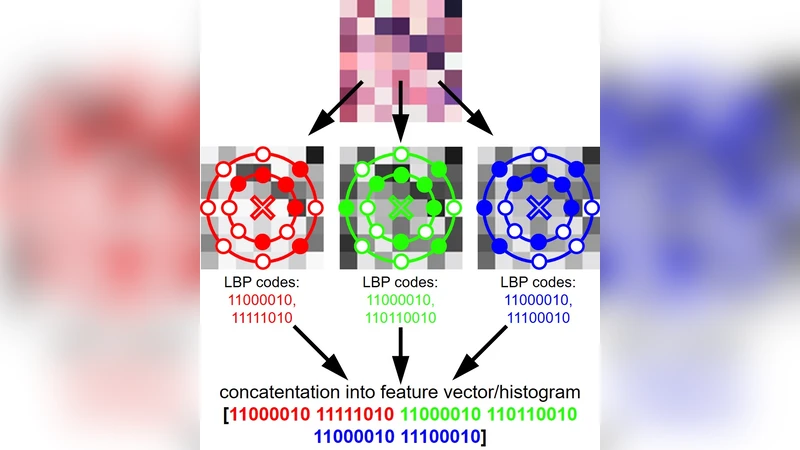
The paper presents a fast, accurate, and easily deployable pipeline for detecting glomeruli in whole‑slide histopathology images (WSIs) that can be on the order of one gigapixel in size. The core idea is to adapt the classic Local Binary Patterns (LBP) descriptor into a “Multi‑radial” version that samples texture at several radii (1–5 pixels) and eight orientations for each radius. By concatenating the normalized histograms of these 24 (or 40) sub‑patterns, a compact yet highly discriminative feature vector is obtained that captures the intricate shape and boundary cues of glomeruli while remaining insensitive to staining variations.
These Multi‑radial LBP vectors are fed to a linear Support Vector Machine (SVM). Because the SVM operates in a high‑dimensional space but with a linear kernel, training is extremely quick (≈15 seconds on a 5‑core Intel i7‑4790 with 40 GB RAM) and inference on a full‑size slide takes about two minutes. The authors evaluated the method on human, mouse, and rat kidney WSIs stained with H&E, PAS, and Masson’s trichrome, achieving a precision of >90 % and a recall of >70 % across all conditions.
To further suppress false positives, a pre‑trained deep convolutional neural network (ResNet‑50) is run in parallel on the candidate patches produced by the SVM‑LBP stage. The CNN re‑scores each candidate and only those with high confidence are retained. This two‑stage approach reduces the overall false‑positive rate to below 3 % while preserving the high precision of the original SVM model.
The pipeline was implemented entirely on CPU, requiring no GPU resources, which makes it suitable for routine laboratory or clinical environments where computational infrastructure may be limited. The authors also demonstrated the clinical relevance of the method by applying it to a mouse model of diabetic nephropathy. Quantitative analysis of the extracted glomeruli revealed a statistically significant increase in glomerular area (≈28 % enlargement) and characteristic shifts in the LBP histogram that correlate with pathological enlargement and mesangial expansion.
Key technical contributions include:
- Multi‑radial LBP design – Extending traditional LBP to multiple radii and orientations provides a richer description of glomerular texture without exploding the feature dimensionality.
- Linear SVM classification – Leveraging a linear kernel yields a model that is both fast to train and robust to limited training data, while still achieving high discriminative power.
- Hybrid SVM‑CNN verification – Combining a lightweight texture‑based detector with a deep network balances speed and accuracy, achieving sub‑3 % false‑positive rates.
- Cross‑species and cross‑stain generalization – The same feature extraction parameters work for human, mouse, and rat kidneys and for several common stains, indicating strong invariance to color and species‑specific morphology.
- Scalability – The method processes a 1‑gigapixel slide in roughly two minutes on modest hardware, making it feasible for high‑throughput pathology workflows.
The authors discuss limitations such as residual false positives in regions with fibrotic or highly vascular tissue, the linear nature of the SVM which may miss extremely complex morphological variations, and the current focus on 2‑D sections rather than 3‑D volumetric data. They propose future work that integrates Multi‑radial LBP descriptors as additional channels in deep networks, extends the approach to other organ systems (e.g., liver lobules, pulmonary alveoli), and explores 3‑D imaging modalities like optical clearing and light‑sheet microscopy.
In summary, this study demonstrates that a carefully engineered texture descriptor combined with classical machine learning can rival deep learning‑only solutions in both speed and accuracy for glomerular detection. The method’s simplicity, low computational demand, and robustness to staining and species variations position it as a practical tool for accelerating histopathology research, aiding in the construction of large annotated datasets, and potentially supporting real‑time diagnostic assistance in clinical pathology.
Comments & Academic Discussion
Loading comments...
Leave a Comment