📝 Original Info
- Title: Macro-molecular data storage with petabyte/cm^3 density, highly parallel read/write operations, and genuine 3D storage capability
- ArXiv ID: 1709.03596
- Date: 2017-09-14
- Authors: Researchers from original ArXiv paper
📝 Abstract
Digital information can be encoded in the building-block sequence of macro-molecules, such as RNA and single-stranded DNA. Methods of "writing" and "reading" macromolecular strands are currently available, but they are slow and expensive. In an ideal molecular data storage system, routine operations such as write, read, erase, store, and transfer must be done reliably and at high speed within an integrated chip. As a first step toward demonstrating the feasibility of this concept, we report preliminary results of DNA readout experiments conducted in miniaturized chambers that are scalable to even smaller dimensions. We show that translocation of a single-stranded DNA molecule (consisting of 50 adenosine bases followed by 100 cytosine bases) through an ion-channel yields a characteristic signal that is attributable to the 2-segment structure of the molecule. We also examine the dependence of the rate and speed of molecular translocation on the adjustable parameters of the experiment.
💡 Deep Analysis
Deep Dive into Macro-molecular data storage with petabyte/cm^3 density, highly parallel read/write operations, and genuine 3D storage capability.
Digital information can be encoded in the building-block sequence of macro-molecules, such as RNA and single-stranded DNA. Methods of “writing” and “reading” macromolecular strands are currently available, but they are slow and expensive. In an ideal molecular data storage system, routine operations such as write, read, erase, store, and transfer must be done reliably and at high speed within an integrated chip. As a first step toward demonstrating the feasibility of this concept, we report preliminary results of DNA readout experiments conducted in miniaturized chambers that are scalable to even smaller dimensions. We show that translocation of a single-stranded DNA molecule (consisting of 50 adenosine bases followed by 100 cytosine bases) through an ion-channel yields a characteristic signal that is attributable to the 2-segment structure of the molecule. We also examine the dependence of the rate and speed of molecular translocation on the adjustable parameters of the experiment.
📄 Full Content
1
Macro-molecular data storage with petabyte/cm3 density, highly
parallel read/write operations, and genuine 3D storage capability
Masud Mansuripur and Pramod Khulbe
Optical Sciences Center, University of Arizona, Tucson, Arizona 85721, masud@u.arizona.edu
[Published in Optical Data Storage 2004, edited by B.V.K. Vijaya Kumar and Hiromichi Kobori;
Proceedings of SPIE 5380, pp272-282 (2004). doi: 10.1117/12.562434]
Abstract. Digital information can be encoded in the building-block sequence of macro-
molecules, such as RNA and single-stranded DNA. Methods of “writing” and “reading”
macromolecular strands are currently available, but they are slow and expensive. In an ideal
molecular data storage system, routine operations such as write, read, erase, store, and transfer
must be done reliably and at high speed within an integrated chip. As a first step toward
demonstrating the feasibility of this concept, we report preliminary results of DNA readout
experiments conducted in miniaturized chambers that are scalable to even smaller dimensions.
We show that translocation of a single-stranded DNA molecule (consisting of 50 adenosine
bases followed by 100 cytosine bases) through an ion-channel yields a characteristic signal that
is attributable to the 2-segment structure of the molecule. We also examine the dependence of
the rate and speed of molecular translocation on the adjustable parameters of the experiment.
- Introduction. Many of the traditional problems in disk and tape data storage can be overcome
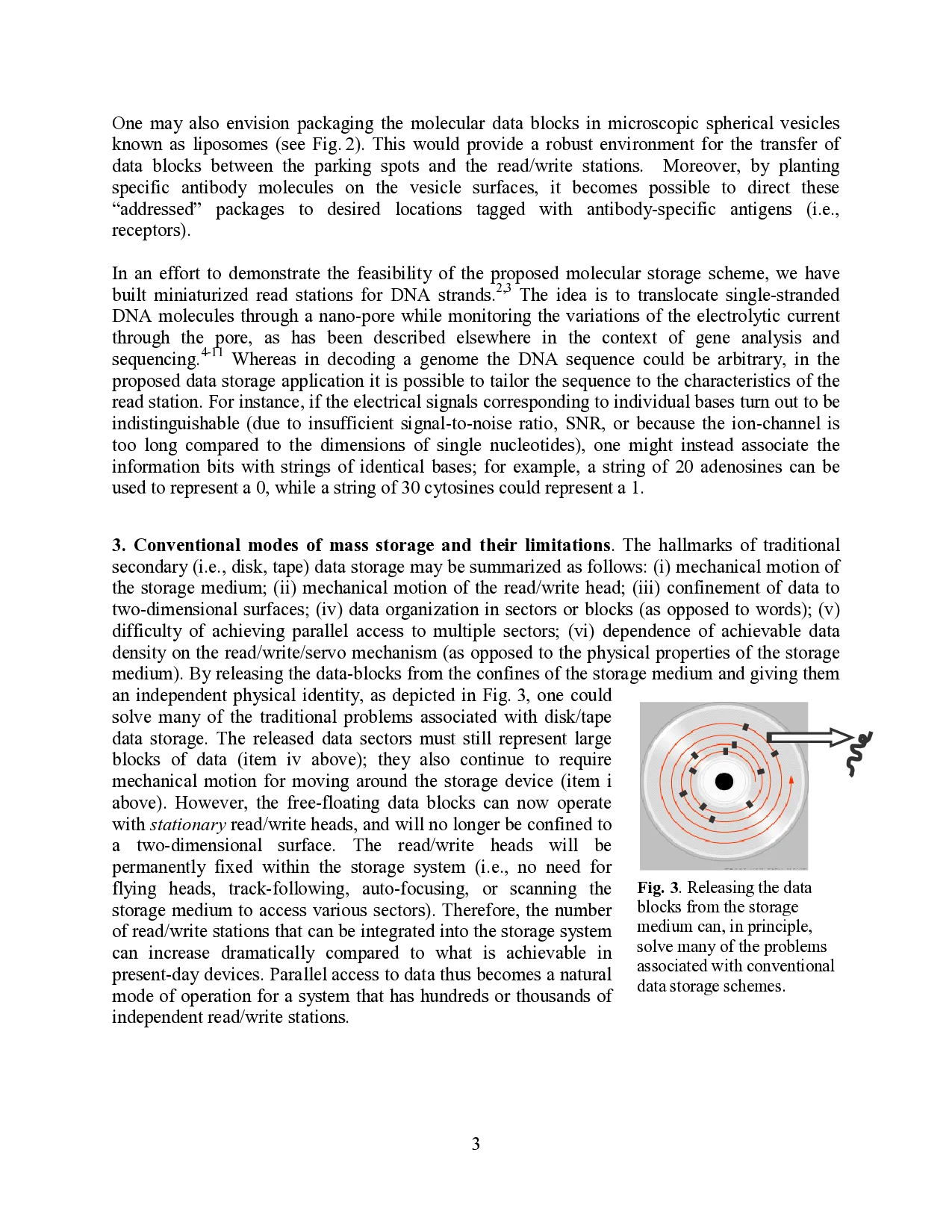
if data-blocks were to be released from the confines of a disk (or tape) and allowed to float freely
between read/write stations (i.e., heads) and permanent “parking spots.” The heads and parking
spots thus become fixed structures within an integrated chip, while the macro-molecular data
blocks themselves become the (mobile) storage media. In this scheme, a large number of
read/write heads could operate in parallel, the heads and parking spots would be constructed
(layer upon layer) in a truly 3-dimensional fashion, and individual nanometer-sized molecules –
strung together in a flexible macromolecular chain – would be used to represent the 0’s and 1’s
of binary information. In this paper we discuss the potential advantages of this alternative
scheme for (secondary) data storage and, to demonstrate the feasibility of the concept, we present
results of experiments based on DNA molecules that travel within micro-fluidic chambers.
In principle, the four nucleotides of DNA can be used to represent a 2-bit sequence (A = 00,
C = 01, G = 10, T = 11), although practical considerations may impose certain restrictions on the
specific sequences that can be used to encode the information. A data storage device built around
this concept must have the ability to (i) create macromolecules with any desired sequence of
building blocks, i.e., write or encode the digital information into macromolecular strands; (ii)
analyze and decode the sequence of a previously created macromolecule, i.e., read the recorded
information; (iii) provide an automated and reliable mechanism for transferring the
macromolecules between the read station, the write station, and designated locations (parking
spots) for storing each such macro-molecule. Although methods of writing and reading
macromolecular strands are currently available (e.g., arbitrary sequences of oligonucleotides can
be synthesized, and DNA sequences can be deciphered), these methods require large machines
and are slow and expensive. In an ideal molecular data storage system, routine operations such as
write, read, erase, store, and transfer must be carried out within an integrated chip, reliably and at
high speed.
2
- Device architecture. As a possible alternative to present-day mass data storage devices (e.g.,
magnetic and optical disks and tapes), we envision a system in which data blocks are encoded
into macromolecules constructed from two or more distinct bases, say, x and y; the bases can be
strung together in arbitrary order such as xxyxyyxy… xyx to represent binary sequences of user-
data (x = 0, y = 1).1,2 The macromolecular data blocks must be created in a write station,
transferred to parking spots for temporary storage, and brought to a read station for decoding
and readout. The erase operation is as simple as discarding a data block and allocating its
parking spot to another macromolecule. (In principle, discarded molecules can be recycled after
being broken down to their constituent elements.) The parking spots and read/write stations
depicted in Fig. 1, for example, are µ-fluidic chambers connected via µ-channels and µ-valves
(not shown) that enable automatic access through an electronic addressing scheme.2 With the
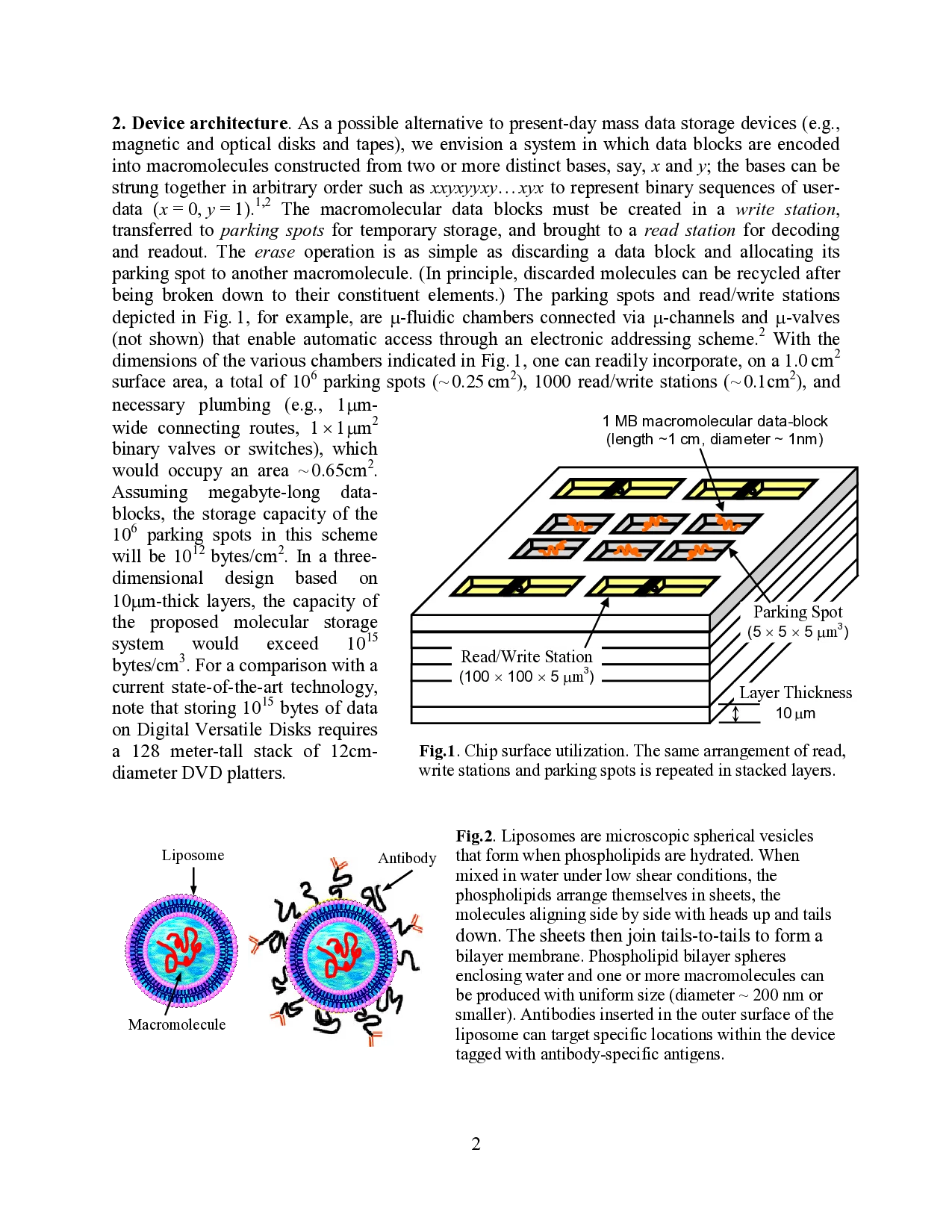
dimensions of the various chambers indicated in Fig. 1, one can readily incorporate, on a 1.0 cm2
surface area, a total of 106 parking spots (~ 0.25 cm2), 1000 read/write stations (~ 0.1cm2), and
…(Full text truncated)…
📸 Image Gallery
Reference
This content is AI-processed based on ArXiv data.