Neighborhood-Based Label Propagation in Large Protein Graphs
Understanding protein function is one of the keys to understanding life at the molecular level. It is also important in several scenarios including human disease and drug discovery. In this age of rapid and affordable biological sequencing, the number of sequences accumulating in databases is rising with an increasing rate. This presents many challenges for biologists and computer scientists alike. In order to make sense of this huge quantity of data, these sequences should be annotated with functional properties. UniProtKB consists of two components: i) the UniProtKB/Swiss-Prot database containing protein sequences with reliable information manually reviewed by expert bio-curators and ii) the UniProtKB/TrEMBL database that is used for storing and processing the unknown sequences. Hence, for all proteins we have available the sequence along with few more information such as the taxon and some structural domains. Pairwise similarity can be defined and computed on proteins based on such attributes. Other important attributes, while present for proteins in Swiss-Prot, are often missing for proteins in TrEMBL, such as their function and cellular localization. The enormous number of protein sequences now in TrEMBL calls for rapid procedures to annotate them automatically. In this work, we present DistNBLP, a novel Distributed Neighborhood-Based Label Propagation approach for large-scale annotation of proteins. To do this, the functional annotations of reviewed proteins are used to predict those of non-reviewed proteins using label propagation on a graph representation of the protein database. DistNBLP is built on top of the “akka” toolkit for building resilient distributed message-driven applications.
💡 Research Summary
The paper addresses the pressing need to annotate the ever‑growing number of protein sequences stored in UniProtKB/TrEMBL, which lack functional information such as Gene Ontology (GO) terms, cellular localization, or disease relevance. Manual curation, as performed for the Swiss‑Prot subset, cannot keep pace with the influx of data generated by high‑throughput sequencing technologies. To bridge this gap, the authors propose DistNBLP (Distributed Neighborhood‑Based Label Propagation), a scalable framework that propagates functional labels from reviewed (Swiss‑Prot) proteins to unreviewed (TrEMBL) proteins using a graph‑based representation of the entire UniProtKB database.
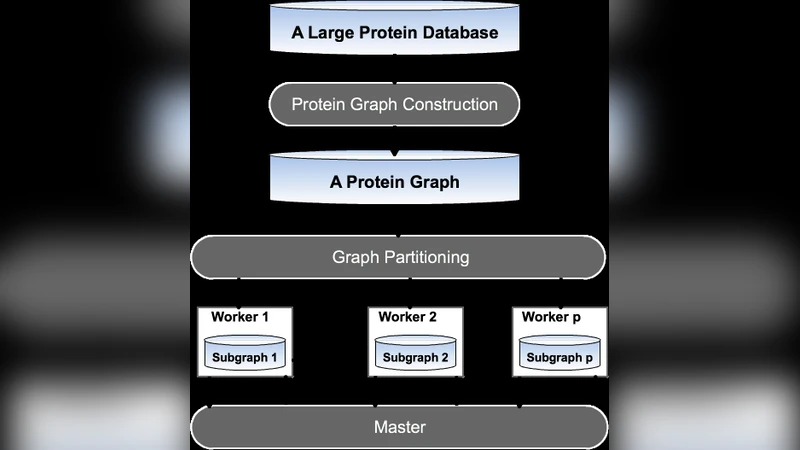
Graph Construction
Each protein is represented as a node. Edges are created between proteins that share significant similarity according to three complementary criteria: (1) sequence similarity measured by BLAST‑P scores, (2) domain architecture similarity derived from Pfam/HMMER matches, and (3) taxonomic proximity based on the least common ancestor in the NCBI taxonomy tree. The three similarity scores are normalized to the interval
Comments & Academic Discussion
Loading comments...
Leave a Comment