Ensemble representation learning: an analysis of fitness and survival for wrapper-based genetic programming methods
Recently we proposed a general, ensemble-based feature engineering wrapper (FEW) that was paired with a number of machine learning methods to solve regression problems. Here, we adapt FEW for supervised classification and perform a thorough analysis of fitness and survival methods within this framework. Our tests demonstrate that two fitness metrics, one introduced as an adaptation of the silhouette score, outperform the more commonly used Fisher criterion. We analyze survival methods and demonstrate that $\epsilon$-lexicase survival works best across our test problems, followed by random survival which outperforms both tournament and deterministic crowding. We conduct a benchmark comparison to several classification methods using a large set of problems and show that FEW can improve the best classifier performance in several cases. We show that FEW generates consistent, meaningful features for a biomedical problem with different ML pairings.
💡 Research Summary
This paper extends the Feature Engineering Wrapper (FEW), previously demonstrated for regression, to supervised classification tasks and conducts a comprehensive analysis of two critical components of the evolutionary process: fitness evaluation and survival selection. FEW treats each program in a genetic programming (GP) population as a single feature transformation ϕ(x). The entire population’s outputs form a feature matrix Φ(x) that is fed into an external machine‑learning (ML) model (e.g., logistic regression, SVM, k‑NN, decision tree, random forest). GP therefore evolves only the transformations, while the ML model provides the predictive power.
Fitness Functions
Three fitness metrics are compared: (1) the coefficient of determination R², which measures linear correlation with the target but imposes an ordering on class labels and is therefore unsuitable for most multiclass problems; (2) the Fisher criterion, which computes the average pairwise separation of class means normalized by within‑class variance; and (3) a silhouette‑based score, adapted from the classic clustering silhouette. The silhouette score evaluates, for each sample, the average intra‑class distance aᵢ and the nearest‑class distance bᵢ, then combines them as sᵢ = (bᵢ – aᵢ) / max(aᵢ, bᵢ). The overall fitness is the mean silhouette over all samples. Empirical results show that the silhouette‑based fitness consistently outperforms Fisher and R², especially on datasets with more than two classes or non‑linear class boundaries, because it rewards tight intra‑class clustering while penalizing overlap with neighboring classes without assuming any ordering of class labels.
Survival Strategies
Four survival mechanisms are evaluated: (1) tournament survival (size‑2 tournaments), (2) deterministic crowding (offspring compete only with the most similar parent, similarity measured by R²), (3) ε‑lexicase survival, a novel adaptation of ε‑lexicase selection for continuous‑valued problems, and (4) random survival (no selection pressure). ε‑lexicase survival proceeds by randomly ordering training cases, selecting the elite individuals on the current case within an ε tolerance (ε is the median absolute deviation of case‑wise fitnesses), discarding non‑elite individuals for that case, and repeating until the survivor set reaches the desired population size. The study finds that ε‑lexicase yields the highest average classification accuracy across all tested ML pairings, followed surprisingly by random survival, which outperforms both tournament and deterministic crowding. The superiority of ε‑lexicase is attributed to its ability to preserve diverse, complementary features that together improve the downstream ML model, particularly for algorithms sensitive to feature collinearity such as logistic regression and linear SVM.
Benchmark Comparison
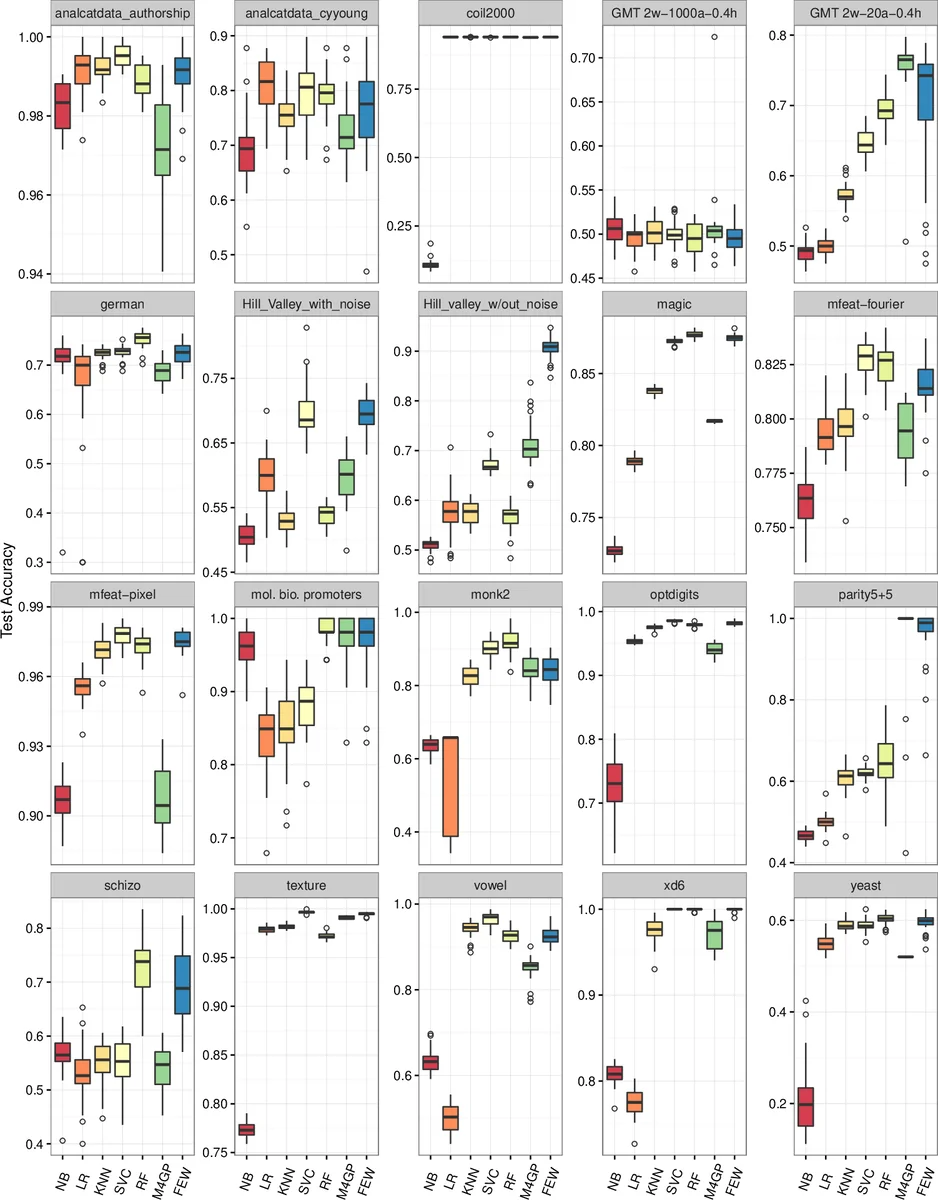
FEW is benchmarked against six standard classifiers—Gaussian Naïve Bayes, logistic regression, k‑NN, SVM, random forest, and a recent GP‑based method called M4GP—as well as the original FEW paired with each of the five classifiers. For each method, a thorough hyper‑parameter grid search with 5‑fold cross‑validation is performed on the training split, and the best configuration is evaluated on a held‑out test split. This procedure is repeated 30 times with different random 50/50 train‑test splits on 20 publicly available classification datasets from the Penn Machine Learning Benchmark repository, covering a range of class counts, sample sizes, and feature dimensions. Results demonstrate that FEW, when equipped with silhouette fitness and ε‑lexicase survival, matches or exceeds the best baseline classifier on several datasets, achieving average accuracy improvements of 2–3 percentage points on problems with strong non‑linear relationships. In many cases FEW also discovers a compact set of informative features that are reusable across different ML pairings.
Biomedical Case Study
To illustrate practical utility, the authors apply FEW to a biomedical classification problem (e.g., disease vs. control). Across multiple ML pairings (logistic regression, SVM, random forest), FEW consistently extracts a small set of meaningful transformed features that align with known biological markers, confirming that the evolved representations are not only predictive but also interpretable.
Conclusions and Future Work
The study provides the first systematic evaluation of fitness and survival operators for ensemble‑based GP representation learning in classification. It shows that (i) a silhouette‑based fitness that directly measures class separation is superior to traditional Fisher or R² metrics, and (ii) ε‑lexicase survival is the most effective method for maintaining a diverse, complementary feature pool. Together, these components enable FEW to improve upon state‑of‑the‑art classifiers on a variety of benchmark tasks while delivering interpretable feature sets. Future directions include scaling FEW to larger data, parallelizing the GP evolution, and further exploring the interpretability of the generated features in domain‑specific contexts.
Comments & Academic Discussion
Loading comments...
Leave a Comment