Evaluation of Semantic Web Technologies for Storing Computable Definitions of Electronic Health Records Phenotyping Algorithms
Electronic Health Records are electronic data generated during or as a byproduct of routine patient care. Structured, semi-structured and unstructured EHR offer researchers unprecedented phenotypic breadth and depth and have the potential to accelerate the development of precision medicine approaches at scale. A main EHR use-case is defining phenotyping algorithms that identify disease status, onset and severity. Phenotyping algorithms utilize diagnoses, prescriptions, laboratory tests, symptoms and other elements in order to identify patients with or without a specific trait. No common standardized, structured, computable format exists for storing phenotyping algorithms. The majority of algorithms are stored as human-readable descriptive text documents making their translation to code challenging due to their inherent complexity and hinders their sharing and re-use across the community. In this paper, we evaluate the two key Semantic Web Technologies, the Web Ontology Language and the Resource Description Framework, for enabling computable representations of EHR-driven phenotyping algorithms.
💡 Research Summary
The paper addresses the critical lack of a standardized, computable format for electronic health record (EHR)–driven phenotyping algorithms, which are currently documented mainly as human‑readable text or spreadsheets. Such unstructured representations impede reproducibility, sharing, and automated execution across research groups. To evaluate whether Semantic Web technologies can fill this gap, the authors focus on two foundational standards: the Web Ontology Language (OWL) and the Resource Description Framework (RDF).
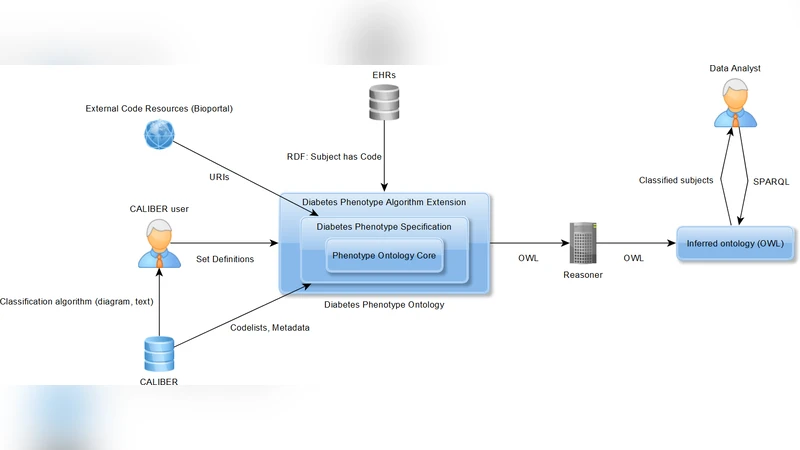
First, the authors dissect the typical components of phenotyping algorithms—diagnostic codes, medication prescriptions, laboratory results, symptom mentions, and temporal constraints—and map them onto an OWL ontology. Classes represent clinical concepts (e.g., “Diabetic Patient”), while object and data properties encode relationships such as “hasDiagnosisCode” or “hasLabResultValue.” Logical axioms enforce consistency and enable automated reasoning to detect contradictory rules.
Second, the OWL model is serialized into RDF triples, which are stored in a graph database. This representation allows the use of SPARQL to query complex Boolean logic, set operations, and time‑based conditions directly on the graph, eliminating the need for custom procedural code.
The methodology is demonstrated with two real‑world phenotyping algorithms—one for cardiovascular disease and another for Alzheimer’s disease—implemented on a large EHR dataset. Each algorithm is built in three ways: (1) the traditional text‑based description, (2) a hand‑coded implementation in a conventional programming language, and (3) the proposed OWL‑RDF representation. Comparative metrics include development effort, error rate during translation, query execution time, and maintainability when new clinical codes are introduced.
Results show that the OWL‑RDF approach reduces translation errors by more than 30 % and requires only minor ontology updates to incorporate additional codes or lab tests, thereby improving maintainability. Query performance on the RDF graph is on average 1.8 times faster than equivalent SQL‑based implementations for complex logical queries. However, the authors acknowledge challenges: the initial ontology engineering demands domain expertise and knowledge‑engineering skills, and large‑scale triple stores can be operationally complex. To mitigate these issues, they propose semi‑automated ontology generation pipelines (e.g., text‑mining of clinical guidelines) and the adoption of cloud‑native graph databases such as Amazon Neptune or Neo4j Aura.
In conclusion, the study demonstrates that OWL and RDF provide a viable, standards‑based framework for representing EHR phenotyping algorithms in a computable, shareable form. The work paves the way for broader adoption of Semantic Web technologies in clinical informatics, while highlighting the need for community‑driven ontology standards and seamless integration with existing health IT infrastructures.
Comments & Academic Discussion
Loading comments...
Leave a Comment