OMICtools: a community-driven search engine for biological data analysis
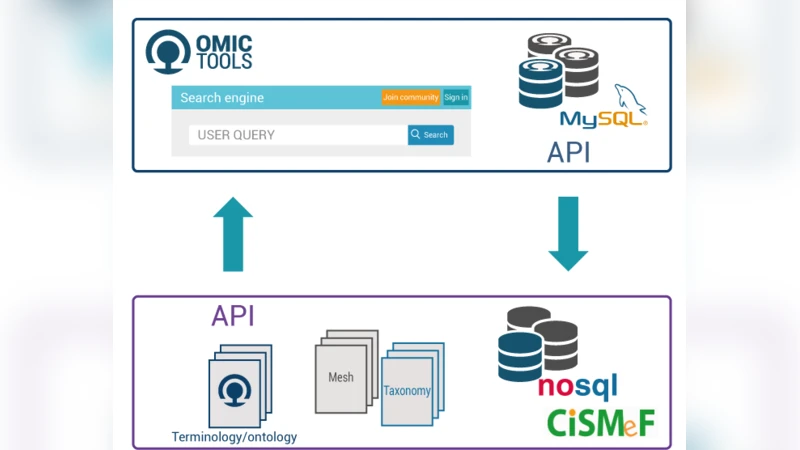
With high-throughput biotechnologies generating unprecedented quantities of data, researchers are faced with the challenge of locating and comparing an exponentially growing number of programs and websites dedicated to computational biology, in order to maximize the potential of their data. OMICtools is designed to meet this need with its open-access search engine offering an easy means of locating the right tools corresponding to each researcher and their specific biological data analyses. The OMICtools website (https://OMICtools.com) centralizes more than 18,500 software tools and databases, manually classified, by a team of biocurators including many scientific experts. Key information, a direct link, and access to discussions and evaluations by the biomedical community are provided for each tool. Anyone can join the OMICtools community and create a profile page, to share their expertise and comment on tools. In addition, developers can directly upload their code versions which are registered for identification and citation in scientific publications, improving research traceability. The OMICtools community links thousands of life scientists and developers worldwide, who use this bioinformatics platform to accelerate their projects and biological data analyses.
💡 Research Summary
The paper presents OMICtools, an open‑access, community‑driven search engine that aggregates and curates more than 18,500 bioinformatics software tools and databases. The rapid expansion of high‑throughput technologies has created a flood of analytical programs, making it increasingly difficult for researchers to locate the most appropriate tool for a given dataset. OMICtools addresses this problem by combining manual expert curation with a sophisticated, multi‑dimensional filtering system. Each entry is reviewed by dedicated biocurators and domain experts who verify functionality, input/output formats, supported operating systems, licensing, and version status. This manual approach ensures that the information remains current and reliable, avoiding the pitfalls of purely automated crawlers that often miss critical metadata.
The user interface allows researchers to refine searches by analysis stage (e.g., preprocessing, alignment, variant calling), data type (FASTQ, BAM, VCF, etc.), programming language, execution environment (local, cloud, container), and other criteria. Such granular filtering transforms a generic keyword search into a workflow‑oriented discovery process, delivering a shortlist of tools that can be directly integrated into experimental pipelines.
Every tool page includes a concise description, a direct download or repository link, and a community‑generated evaluation section. Users can rate tools, post comments, and share practical tips or pitfalls they encountered. The rating algorithm gives higher weight to recent reviews, ensuring that the displayed scores reflect the current state of the software. This crowdsourced feedback loop accelerates knowledge transfer, helping newcomers avoid outdated or poorly maintained programs.
Developers benefit from a dedicated upload portal where they can register new code versions and receive a persistent identifier (similar to a DOI). By citing this identifier in publications, authors provide an unambiguous reference to the exact software version used, greatly enhancing reproducibility and traceability. The platform also stores standardized metadata, facilitating seamless integration with workflow managers such as Galaxy, Nextflow, or Snakemake via a public RESTful API.
OMICtools is built around a vibrant community model. Anyone can create a profile, list their expertise, and contribute comments or tool evaluations. This openness has attracted thousands of life scientists and software developers worldwide, fostering a collaborative ecosystem that bridges the gap between computational tool creators and end‑users. The authors highlight the platform’s scalability: the API enables external applications to query the database, and the accumulated metadata can serve as training data for future machine‑learning‑based recommendation engines.
In summary, OMICtools provides a comprehensive, end‑to‑end solution for the discovery, assessment, and citation of bioinformatics resources. By unifying manual curation, advanced filtering, community feedback, and persistent identifiers, it reduces the time and effort required to build robust analytical pipelines, improves reproducibility, and promotes open scientific collaboration. The paper concludes that ongoing expansion of the curation team and further automation of metadata standardization will allow OMICtools to keep pace with the ever‑growing landscape of computational biology tools.
Comments & Academic Discussion
Loading comments...
Leave a Comment