HuGaDB: Human Gait Database for Activity Recognition from Wearable Inertial Sensor Networks
This paper presents a human gait data collection for analysis and activity recognition consisting of continues recordings of combined activities, such as walking, running, taking stairs up and down, sitting down, and so on; and the data recorded are segmented and annotated. Data were collected from a body sensor network consisting of six wearable inertial sensors (accelerometer and gyroscope) located on the right and left thighs, shins, and feet. Additionally, two electromyography sensors were used on the quadriceps (front thigh) to measure muscle activity. This database can be used not only for activity recognition but also for studying how activities are performed and how the parts of the legs move relative to each other. Therefore, the data can be used (a) to perform health-care-related studies, such as in walking rehabilitation or Parkinson’s disease recognition, (b) in virtual reality and gaming for simulating humanoid motion, or (c) for humanoid robotics to model humanoid walking. This dataset is the first of its kind which provides data about human gait in great detail. The database is available free of charge https://github.com/romanchereshnev/HuGaDB.
💡 Research Summary
The paper introduces HuGaDB, a publicly available human gait dataset collected with a body‑sensor network composed of six wearable inertial measurement units (IMUs) and two surface electromyography (EMG) sensors. The IMUs (MPU9250) are placed symmetrically on the right and left thighs, shins, and feet, while the EMG electrodes are attached to the rectus femoris muscles of both legs. Each IMU provides three‑axis acceleration (±2 g) and three‑axis angular velocity (±2000 °/s) at an average sampling rate of 56.35 kHz; the EMG channels sample at 1 kHz with 8‑bit resolution.
Data were recorded from 18 healthy young adults (4 female, 14 male, mean age 23.7 years, mean height 179 cm, mean weight 73 kg). Participants performed a natural sequence of activities—sitting, standing, walking, running, ascending/descending stairs, bicycling, elevator rides, and sitting in a car—without any artificial constraints or obstacles. The recordings are continuous; activity transitions are captured and later segmented with precise labels. In total, the collection comprises 2,111,962 samples, equivalent to roughly 10 hours of sensor data, covering 12 activity classes.
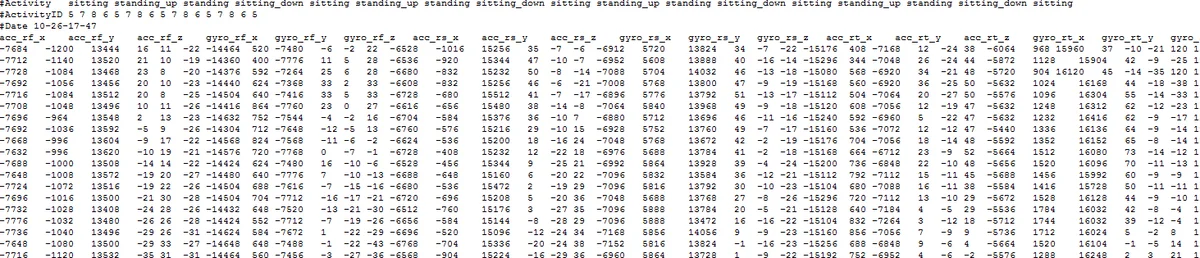
The dataset is stored as plain‑text, tab‑delimited files. Each file follows a naming convention “HGD_vX_ACT_PR_CNT.txt” where ACT denotes the activity identifier (or “VARIOUS” for mixed‑activity recordings), PR is the participant ID, and CNT is a repetition counter. A four‑line header lists the activities present, their numeric IDs, and the recording timestamp. The body contains 39 columns: 36 columns for the six IMUs (each providing three acceleration and three gyroscope axes), two columns for the EMG signals, and one column for the activity label. Raw values are kept in integer format (int16 for IMU, uint8 for EMG) to allow users to apply custom scaling or filtering. Example loading scripts for Python (NumPy) and MATLAB are provided, as well as a utility to import the data into an SQLite database.
A variance analysis shows low intra‑subject variability for short gait cycles but considerably higher inter‑subject variability, especially in the EMG channels, due to differences in skin conductivity, muscle morphology, and sensor placement. The authors argue that this variability underscores the need for robust evaluation protocols; they recommend a leave‑one‑subject‑out cross‑validation scheme to assess generalization to unseen users.
Compared with existing public datasets (e.g., PAMAP2, WARD, Daphnet Gait, CMU‑MMAC), HuGaDB’s novelty lies in (1) the dense placement of sensors on each leg segment, enabling detailed study of inter‑segment kinematics; (2) the simultaneous acquisition of EMG and inertial data, facilitating research on muscle‑movement coupling; and (3) the provision of continuous, annotated recordings that include transition periods between activities. These features make the dataset valuable for a broad range of applications:
- Healthcare and rehabilitation – detection of gait abnormalities, monitoring of Parkinson’s disease progression, and assessment of post‑surgical recovery.
- Virtual reality and gaming – realistic avatar motion synthesis by integrating acceleration data to reconstruct limb trajectories.
- Humanoid robotics – data‑driven modeling of human‑like walking patterns for control and learning algorithms.
- Multimodal sensor fusion research – combining video, inertial, and EMG modalities for richer activity recognition.
The authors acknowledge limitations: the data were collected mainly in indoor environments and a few outdoor scenarios (bicycling, car ride), lacking diverse terrains, weather conditions, or treadmill walking. All sensors are from a single manufacturer, so cross‑device calibration remains an open issue. Nevertheless, the dataset (≈455 MB) is freely downloadable from GitHub, and the authors provide scripts for preprocessing and database creation.
In summary, HuGaDB offers a high‑resolution, multi‑modal gait dataset with comprehensive activity labeling and inter‑segment coverage, filling a gap in the public domain and enabling more nuanced studies of human locomotion, activity recognition, and related biomedical and robotics applications.
Comments & Academic Discussion
Loading comments...
Leave a Comment