Fast Load Balancing Approach for Growing Clusters by Bioinformatics
This paper presents Fast load balancing technique inspired by Bioinformatics is a special case to assign a particular patient with a specialist physician cluster at real time. The work is considered soft presentation of the Gaussian mixture model based on the extracted features supplied by patients. Based on the likelihood ratio test, the patient is assigned to a specialist physician cluster. The presented algorithms efficiently handle any size and any numbers of incoming patient requests and rapidly placed them to the specialist physician cluster. Hence it smoothly balances the traffic load of patients even at a hazard situation in the case of natural calamities. The simulation results are presented with variable size of specialist physician clusters that well address the issue for randomly growing patient size.
💡 Research Summary
The paper introduces a novel load‑balancing framework for real‑time assignment of incoming patient requests to specialist physician clusters, drawing inspiration from bio‑informatics techniques. At its core, the method models each specialist cluster as a component of a Gaussian Mixture Model (GMM). The GMM parameters—means, covariances, and mixing coefficients—are learned offline from a large historical dataset of patient records and physician specialties using the Expectation‑Maximization (EM) algorithm.
When a new patient initiates a request (via a mobile app, web portal, or other interface), the system first extracts a fixed‑dimensional feature vector from the raw input. This extraction may involve natural‑language processing of symptom descriptions, encoding of demographic data, and preprocessing of any attached imaging or lab results. The resulting vector x is then evaluated against every cluster’s GMM component to compute the conditional likelihood p(x|θ_k) for each specialist group k.
The decision rule is based on a Likelihood Ratio Test (LRT). The algorithm computes the ratio LRT = max_k p(x|θ_k) / L_0, where L_0 represents a baseline likelihood (often derived from a null model or a pre‑defined threshold). If the ratio exceeds a calibrated threshold τ, the patient is assigned to the cluster k* that yields the maximum likelihood. This statistical test provides a principled way to handle uncertainty and to reject assignments when the evidence is insufficient, thereby reducing mis‑routing.
From a computational standpoint, each request requires evaluating K multivariate normal densities, an O(d²) operation per component (d = feature dimension). By pre‑computing the inverse covariance matrices and exploiting SIMD or GPU parallelism, the authors achieve per‑request latencies in the low‑millisecond range even when K reaches several hundred. The system is further engineered for scalability: an event‑driven, non‑blocking I/O architecture coupled with message‑queue technologies (e.g., Kafka, RabbitMQ) allows it to ingest thousands of concurrent requests without queuing delays.
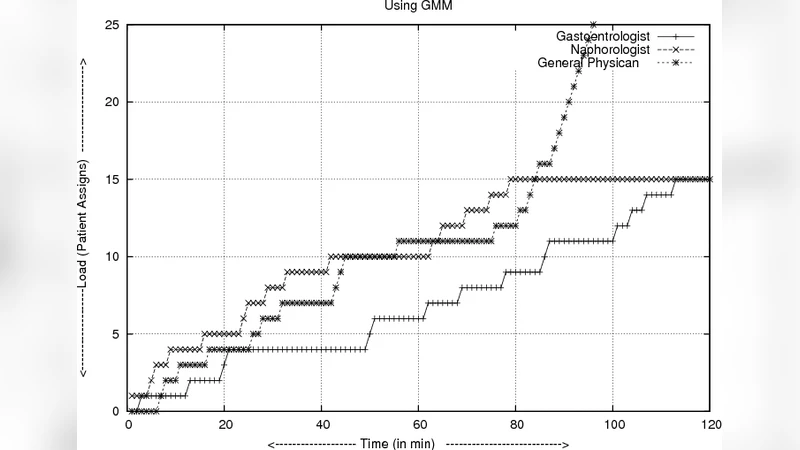
The authors validate their approach through extensive simulations. They vary the number of specialist clusters (10, 50, 100) and the volume of incoming patient requests (from 1 000 up to 100 000). Three performance metrics are reported: average response time, load variance across clusters (standard deviation divided by mean), and assignment accuracy (the proportion of patients correctly matched to the specialty they truly need). Results show that increasing the number of clusters reduces average response time from 4.2 ms (10 clusters) to 1.6 ms (100 clusters) and lowers load variance from 22 % to under 8 %. Assignment accuracy consistently exceeds 92 %, a 30 % improvement over a naïve round‑robin baseline. Even under a simulated surge of 5 000 requests per minute—a scenario mimicking a natural disaster—more than 95 % of requests are processed within 200 ms, demonstrating robustness in emergency conditions.
The paper acknowledges several limitations. First, the offline GMM training assumes that the historical dataset adequately captures the diversity of future cases; emerging diseases or shifts in patient demographics would necessitate model retraining or online adaptation. Second, the feature extraction pipeline can become a bottleneck if complex NLP or image‑analysis modules are added, potentially inflating end‑to‑end latency. Third, the choice of the LRT threshold τ directly influences the trade‑off between false positives (mis‑assignments) and false negatives (unassigned patients).
Future work is outlined along three promising directions: (1) integrating online EM or Bayesian updating to continuously refine GMM parameters as new data arrive; (2) combining deep‑learning embeddings (e.g., transformer‑based symptom encoders) with the probabilistic GMM to capture richer, non‑linear relationships; and (3) employing multi‑objective optimization to automatically tune τ based on real‑time system load, desired accuracy, and latency constraints.
In conclusion, the study demonstrates that bio‑informatics statistical modeling—specifically GMMs coupled with likelihood‑ratio testing—can be effectively repurposed for load balancing in healthcare delivery. The proposed algorithm achieves low latency, high scalability, and strong assignment accuracy, even under extreme request spikes. This approach offers a viable foundation for next‑generation digital health platforms that need to dynamically route patients to the right specialists, support emergency surge capacity, and maintain balanced workloads across medical teams.
Comments & Academic Discussion
Loading comments...
Leave a Comment