Enhancement of Epidemiological Models for Dengue Fever Based on Twitter Data
Epidemiological early warning systems for dengue fever rely on up-to-date epidemiological data to forecast future incidence. However, epidemiological data typically requires time to be available, due to the application of time-consuming laboratorial tests. This implies that epidemiological models need to issue predictions with larger antecedence, making their task even more difficult. On the other hand, online platforms, such as Twitter or Google, allow us to obtain samples of users’ interaction in near real-time and can be used as sensors to monitor current incidence. In this work, we propose a framework to exploit online data sources to mitigate the lack of up-to-date epidemiological data by obtaining estimates of current incidence, which are then explored by traditional epidemiological models. We show that the proposed framework obtains more accurate predictions than alternative approaches, with statistically better results for delays greater or equal to 4 weeks.
💡 Research Summary
The paper addresses a critical limitation of dengue fever early warning systems (EEWS): the latency of official epidemiological data, which typically becomes available several weeks after the actual infection date due to laboratory confirmation and reporting processes. This delay (denoted γ) forces predictive models to issue forecasts with a larger antecedence (β + γ), thereby reducing accuracy because recent incidence is a strong predictor of near‑future cases.
To mitigate this problem, the authors propose a three‑component framework that leverages real‑time Twitter data as a proxy for the missing recent epidemiological observations. First, they collect geolocated tweets from January 2011 to October 2016 that contain the keywords “dengue”, “aedes”, or “aegypti”. The tweets are aggregated by city and week, producing a time series of dengue‑related tweet counts for each of 213 Brazilian municipalities (selected because they have populations >100 k and experienced at least medium incidence during the study period).
The framework proceeds as follows:
- Estimation of delayed epidemiological data – A Gaussian Process (GP) regression model learns the relationship between weekly tweet volumes (online covariate) and the logged dengue incidence rate (DIR) from periods where both are available. The GP provides both point estimates and predictive variance for the missing weeks t − γ … t.
- Reliability assessment – The variance of each GP estimate is compared against a pre‑defined threshold. Only estimates with sufficiently low uncertainty are admitted to the training set; uncertain estimates are discarded to avoid contaminating the model with noise.
- Traditional EEWS forecasting – Using the possibly augmented training set (original confirmed DIR up to t − γ plus reliable GP‑estimated DIR for t − γ … t), a second GP model forecasts DIR at future time t + β. This forecasting GP employs a composite covariance function k = k_loc + k_qp, where k_loc is a Matérn 5/2 kernel capturing short‑term temporal smoothness, and k_qp is a quasi‑periodic kernel (product of a Matérn and a periodic component) that encodes the strong annual seasonality observed in dengue data. Hyper‑parameters are learned by maximizing the log‑likelihood.
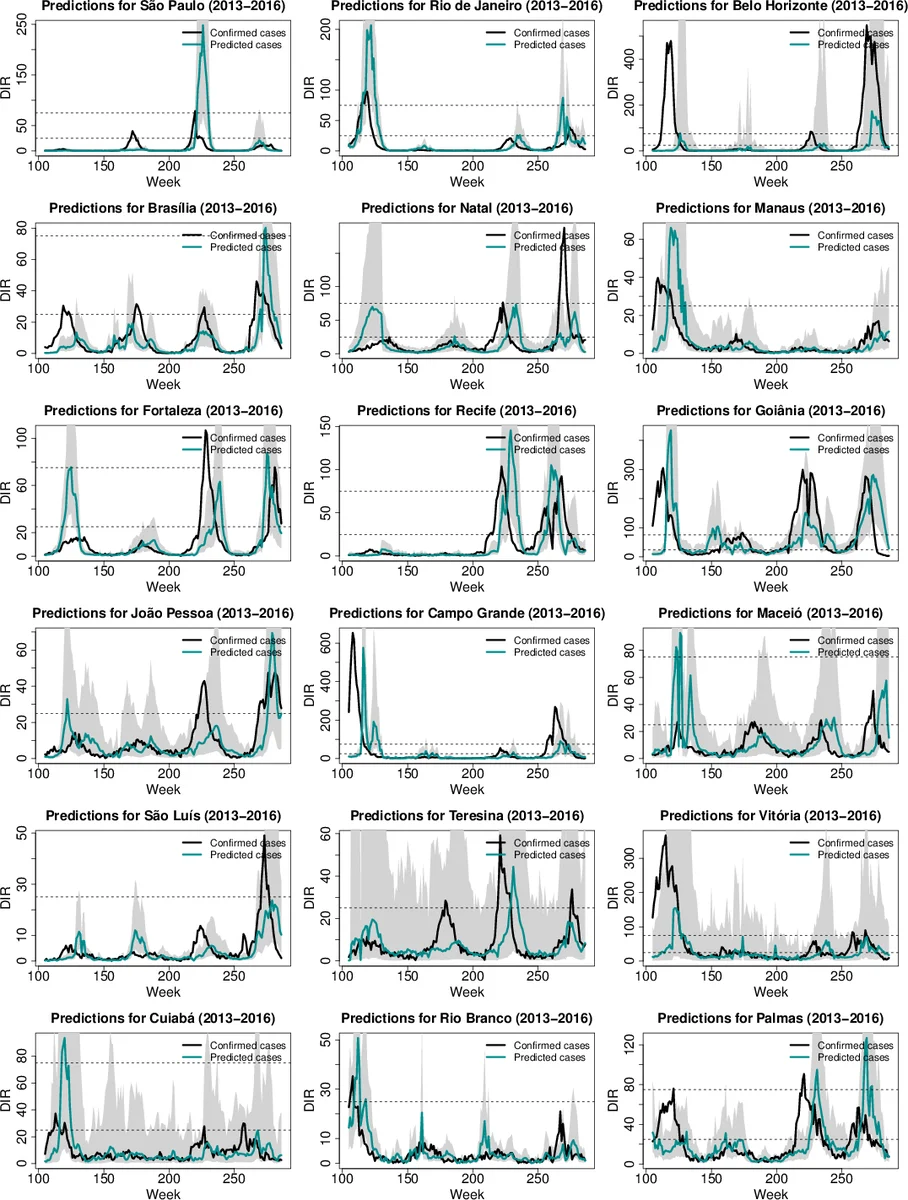
Experimental evaluation compares three setups: (a) the naïve “increased antecedence” approach that ignores any online data, (b) a baseline linear/polynomial model using lagged Twitter counts, and (c) the proposed framework. The authors test reporting delays γ of 2, 4, and 6 weeks while keeping the forecast horizon β at 2 weeks. Performance metrics include Mean Absolute Error (MAE), Root Mean Square Error (RMSE), and Pearson correlation. Results show that when γ ≥ 4 weeks, the proposed method reduces MAE and RMSE by roughly 10–15 % relative to the naïve approach, and improves correlation from ~0.78 to ~0.84. Statistical significance (p < 0.05) is reported. Moreover, the framework automatically refrains from using Twitter estimates in cities where tweet volume is sparse, thereby preserving baseline performance rather than degrading it.
The study contributes (i) a generalizable pipeline for converting real‑time social‑media signals into reliable estimates of delayed epidemiological data, (ii) an uncertainty‑driven gating mechanism that safeguards model training, and (iii) a seamless integration of these estimates into a sophisticated spatio‑temporal GP forecasting model. Limitations include potential demographic bias of Twitter users, noise introduced by irrelevant tweets, and reliance on a single keyword set. Future work is suggested to incorporate multiple online sources (Google Trends, Wikipedia page views), explore deep‑learning based text classification for richer signal extraction, and develop Bayesian hierarchical models that propagate uncertainty throughout the entire pipeline.
In summary, by augmenting traditional dengue EEWS with Twitter‑derived now‑casts of recent incidence, the authors demonstrate a statistically significant improvement in forecast accuracy, especially under realistic reporting delays of four weeks or more, while maintaining robustness in data‑scarce settings.
Comments & Academic Discussion
Loading comments...
Leave a Comment