Probabilistic generation of random networks taking into account information on motifs occurrence
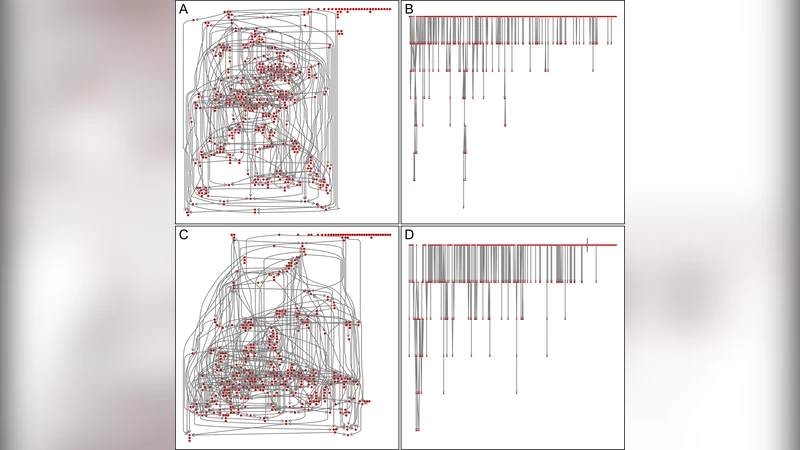
Because of the huge number of graphs possible even with a small number of nodes, inference on network structure is known to be a challenging problem. Generating large random directed graphs with prescribed probabilities of occurrences of some meaningful patterns (motifs) is also difficult. We show how to generate such random graphs according to a formal probabilistic representation, using fast Markov chain Monte Carlo methods to sample them. As an illustration, we generate realistic graphs with several hundred nodes mimicking a gene transcription interaction network in Escherichia coli.
💡 Research Summary
The paper addresses the challenging problem of generating large random directed graphs that not only match a prescribed edge density but also reproduce higher‑order structural features such as degree distributions and motif frequencies. Building on the classic Erdős‑Rényi model, the authors introduce a hierarchical probabilistic framework consisting of three independent priors. The first prior assigns a Bernoulli probability pᵢⱼ to each possible directed edge, allowing the incorporation of expert knowledge or experimental evidence on specific interactions. The second prior encodes a global degree distribution, modeled as a power‑law P(d) ∝ d⁻ᵞ, where the exponent γ can be tuned to reflect scale‑free behavior observed in many biological networks. The third prior captures the occurrence of three‑node motifs—specifically feed‑forward loops (FFL) and feedback loops (FBL)—by using a Beta‑Binomial distribution with parameters (u, v). This choice reflects uncertainty about the exact proportion of each motif while still enforcing an expected ratio derived from empirical observations.
These three components are multiplied to form a total graph prior P_total(G), which is proportional to the product of the edge‑level Bernoulli terms, the degree‑based factor, and the motif‑based Beta‑Binomial term. Because the normalizing constant is intractable, the authors rely on Markov chain Monte Carlo (MCMC) sampling to draw graphs from this distribution. They implement a Metropolis‑Hastings algorithm that proposes toggling a single entry of the adjacency matrix at each step, computes the ratio of the total prior probabilities for the proposed and current graphs, and accepts the move with the usual probability. By iterating this process billions of times, the chain converges to the target distribution. Convergence is assessed using the Gelman‑Rubin diagnostic, which consistently indicates satisfactory mixing after roughly one billion iterations.
The methodology is demonstrated on the transcriptional regulatory network of Escherichia coli, consisting of 423 operons (nodes) and 578 directed regulatory interactions (edges), representing a sparse graph (≈0.32 % of all possible edges). Two experimental settings are explored. In the first, all edges receive a uniform “vague” prior equal to the observed edge density (0.0032), the degree exponent γ is set to 1.7 (matching the empirical power‑law tail), and the motif prior parameters are chosen as u = 2, v = 50, yielding an expected feedback‑loop proportion of about 2 %. In the second setting, edges reported in the curated RegulonDB database are given a high probability (0.95) while all other potential edges receive a low probability (0.00016), preserving the same expected total edge count. For each scenario, three independent MCMC chains of two billion iterations each are run on a modest Intel Core 2 Duo workstation. Sampling a 423‑node graph takes roughly two minutes, and the algorithm scales approximately with the square of the number of nodes, making it feasible for networks of a thousand nodes or more.
Results show that the generated graphs faithfully reproduce the target degree distribution, the exact number of feed‑forward loops (≈42), and the near‑absence of feedback loops, closely matching the real E. coli network. Minor discrepancies appear only for extremely rare events (frequency < 1/10 000), where Monte Carlo variance becomes noticeable. The authors emphasize that their contribution lies not only in the formal probabilistic formulation that unifies edge‑level, degree‑level, and motif‑level information, but also in the practical, fast MCMC implementation that can serve as a prior generator for Bayesian network inference or as a test‑bed for simulation studies. They suggest extensions to larger motifs, signed edges, and integration with downstream Bayesian learning pipelines, which would allow simultaneous incorporation of prior structural knowledge and data‑driven likelihoods.
Comments & Academic Discussion
Loading comments...
Leave a Comment