Low-dose cryo electron ptychography via non-convex Bayesian optimization
Electron ptychography has seen a recent surge of interest for phase sensitive imaging at atomic or near-atomic resolution. However, applications are so far mainly limited to radiation-hard samples because the required doses are too high for imaging b…
Authors: Philipp Michael Pelz, Wen Xuan Qiu, Robert B"ucker
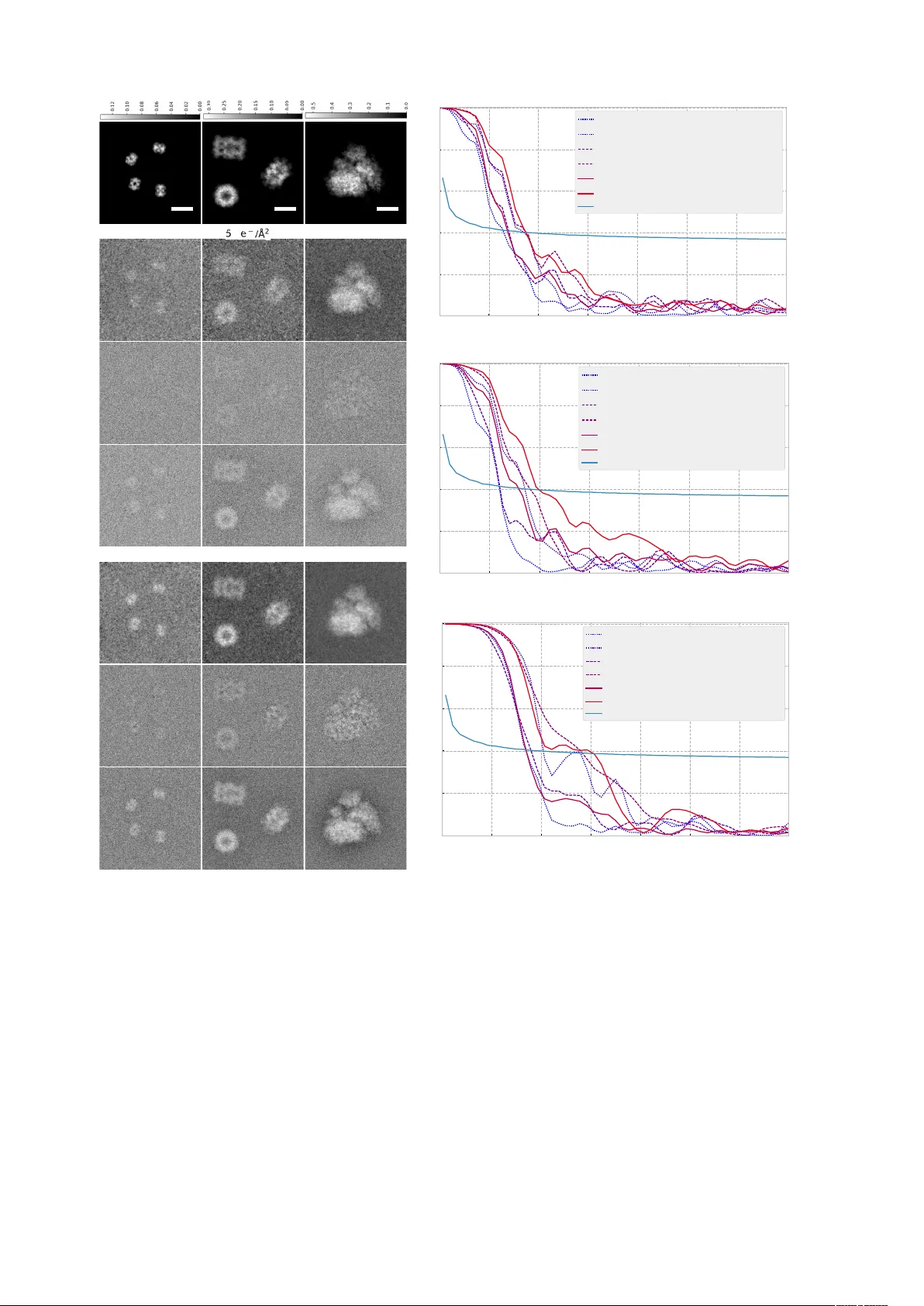
Lo w-dose cry o electr on ptyc hograph y via non-con ve x Ba y esian optimization Philipp Michael P elz 1,2,3,* , W en Xuan Qiu 4 , Robert Buecker 1,2 , Guenther Kassier 1,2 , and R.J. Dwa yne Miller 1,2,3,4 1 Max Planck Institute f or the Structure and Dynamics of Matter , 22761 Hamburg, Ger many 2 Center f or F ree Electron Laser Science , Luruper Chaussee 149, 22761 Hamburg, Germany 3 Depar tment of Physics , Univ ersity of Hamburg, Hamb urg 22761, Germany 4 Depar tments of Chemistr y and Physics , Univ ersity of T oronto , 80 St. George Street, T oronto M5S 1H6, Canada * philipp .pelz@mpsd.mpg.de ABSTRA CT Electron ptychograph y has seen a recent surge of interest f or phase sensitiv e imaging at atomic or near-atomic resolution. Ho we v er , applications are so f ar mainly limited to radiation-hard samples because the required doses are too high for imaging biological samples at high resolution. W e propose the use of non-conv e x, Bay esian optimization to ov ercome this prob lem and reduce the dose required f or successful reconstruction by two orders of magnitude compared to previous experiments. W e suggest to use this method for imaging single biological macromolecules at cr yogenic temperatures and demonstrate 2D single-par ticle reconstructions from simulated data with a resolution of 7 . 9 ˚ A at a dose of 20 e − / ˚ A 2 . When av eraging over only 15 low-dose datasets, a resolution of 4 ˚ A is possible for large macromolecular complex es. With its independence from microscope transf er function, direct recov ery of phase contrast and better scaling of signal-to-noise ratio, cr yo-electron ptychograph y may become a promising alternative to Zer nike phase-contrast microscopy . Introduction The adv ent of direct electron detectors has led to a resolution re volution in the field of cryo-electron microscopy in the last few years, producing three-dimensional atomic potential maps of biological macromolecules of a fe w 100 kDa with a resolution better than 3 . 5 ˚ A [ 1 , 2 ], so that indi vidual amino acid side-chains can be resolv ed. An important part in this re volution is ne w image processing algorithms based on a Bayesian approach, which infer important parameters without user intervention [ 3 ] and correct beam induced motion [ 4 ]. Howe v er , se veral challenges remain to be ov ercome to routinely reach 3 ˚ A resolution also for small complex es [ 5 , 6 ]: Firstly , beam-induced specimen charging and subsequent motion currently still render the high resolution information of the first fe w frames of a high repetition rate mo vie recorded with a direct electron detector unusable [ 7 ], because the motion is too f ast to efficiently correct for it. Secondly , the Detecti v e Quantum Ef ficiency (DQE) of detectors can still be improv ed at high spatial frequencies [ 6 , 8 ]. Thirdly , the contrast of single images can still be improv ed to enable reconstructions with fewer particles and increase the throughput [ 8 ]. The last of these challenges has recently been addressed with a ne w phase plate model [ 9 , 10 ], which is comparati v ely simple to use and provides e xcellent contrast at lo w spatial frequencies. In addition to this hardware-based approach to achiev e linear phase contrast in the measured amplitudes, discovered by Zernike in the 1930s [ 11 ], it is also possible to algorithmically retriev e the phase information from a set of coherent diffraction measurements. One such technique, commonly known as ptyc hogr aphy or scanning coher ent dif fractive micr oscopy [ 12 ], is becoming increasingly popular in the field of materials science due to experimental robustness and the possibility to obtain quantitativ e phase contrast with an essentially unlimited field of view [ 13 , 14 ]. The use of ptychography for imaging radiation sensiti ve samples with electrons at high resolution is ho we v er precluded so far by its high dose requirements. W e show here ho w the use of non-con ve x Bayesian optimization to solve the ptychographic phase retriev al problem fulfills the dose requirements for imaging biological macromolecules and mak es it possible to obtain 2D images from single particles with sub-nanometer resolution. After a short introduction into the technique, we will also mention how ptychograph y offers impro vements for the other tw o challenges discussed abov e. Despite being first proposed as a solution to the phase problem for electrons [ 15 , 16 ], ptychography has seen its biggest success in X-ray imaging, due to the less stringent sample requirements and the experimental need for lensless imaging techniques. Recent developments include the introduction of iterativ e algorithms to enable the reconstruction of datasets collected with an out-of focus probe [ 17 , 18 ], which decreases the memory requirements of the method dramatically; the algorithmic corrections of experimental dif ficulties such as unknown scan positions 1 [ 19 , 20 , 21 ], partial coherence [ 22 ], probe movement during exposure [ 23 , 24 ], intensity fluctuations during the scan [ 22 , 25 ] and reconstruction of background noise [ 13 , 25 ]. In recent years, some of these advances hav e been applied in the context of electron microscopy and yielded atomic resolution reconstructions of low-atomic number materials [ 26 , 27 , 28 ] and quantitati v e phase information [ 13 ]. Fig. 1 shows the experimental set-up for an out-of focus ptychography experiment. A ptychographic dataset is collected by scanning a spatially confined, coherent beam, subsequently called ’probe’, over the specimen and recording far-field diffraction patterns at a series of positions such that the illuminated regions of neighboring positions ov erlap. The diffraction limited resolution of the final image is gi ven by the maximum angle subtended by the detector r d = λ ∗ ∆ z N pix / 2 ∗ d pix , where N pix is the number of detector pixels , d pix is the detector pixel size, λ is the wa velength. Given the set of positions and a realistic forw ard model which describes the formation of the dif fraction pattern, the complex-v alued transmission function which describes the atomic properties of the specimen [ 29 ], can be retriev ed by solving a non-con ve x in v erse problem. The electron dose used for successful reconstruction has Figure 1. Experimental geometry in ptychography . The coherent electron wav e function ψ ( ~ r ) illuminates sev eral regions of the sample, characterized by the transmission function T ( ~ r ) . For e very position, a 2D dif fraction pattern is recorded in the far field. The sample thickness t can be neglected for biological macromolecules in the reconstruction at the resolutions presented in this paper . been abov e 1 × 10 3 e − / ˚ A 2 so far , limiting its usability to radiation-hard specimens. T able 1 lists recently published electron ptychography experiments and the used doses. The lowest dose was reported in [ 30 ], which used an estimated 3 . 33 × 10 3 e − / ˚ A 2 at a resolution of 58 . 4 pm , resulting in a dose of 1 . 1 × 10 3 e − / pixel and achieving a line resolution of 2 . 3 ˚ A , demonstrated by resolving the lattice spacing of gold nanoparticles. W e demonstrate here via simulations that it is possible to reduce the dose by almost a factor of 100 to reach the dose range required for imaging biological macromolecules. The problem of beam-induced sample movement has already been addressed before the de velopment of f ast direct detectors. Scanning with small spots of sev eral 10 nm in size over a vitrified sample has shown to reduce beam induced specimen movement [ 31 , 32 , 33 ] in real-space imaging and it has been noted that the remaining movement may be due to radiation damage, not sample charging [ 31 ]. Ptychography naturally operates with a confined beam, thus minimizing the area where char ge can b uild up, such that the mo vement should be reduced compared to the illumination of large areas in cryo-EM. Additionally we note that due to ptychography being a scanning technique, fast acquisition and rastering is instrumental to achie ve high throughput. This and the sampling requirements giv en by the experimental setup allo w to operate the detector with very large ef fecti ve pix el sizes, such that the DQE and MTF are at near-perfect v alues. 2/ 14 Results Image formation of cry o-TEM and cryo-ptyc hography W e perform multislice simulations of three dif ferent biological macromolecules with molecular weights ranging from 64 kDa to 4 MDa . W e choose the 64 kDa hemoglobin [ 34 ], the 706 kDa 20S proteasome from yeast [ 2 ], and the 4 MDa human ribosome [ 35 ].W e create atomic potential maps using the Matlab code InSilicoT em [ 36 ] with a thickness of 70 nm at an electron ener gy of 300 keV . W e use the isolated atom superposition approximation, without solving the Poisson-Boltzmann equations for the interaction between the molecule and the ions. W e also do not model the amorphousness of the solvent, which w as performed in [ 36 ] using molecular dynamics simulations, but was seen to hav e a negligible effect at v ery low doses. As described in [ 36 ], we model the imaginary part of the potential via the inelastic mean free path, creating a minimal transmission contrast between the vitreous ice and the protein. Using these potential maps, we simulate a ptychograph y experiment by cropping three-dimensional slices from this potential at se veral positions and propagate a coherent incoming wa ve through the potential slices using the methods described in [ 37 ] in custom code. W e model the detector properties as in [ 36 ], by multiplying the Fourier transform of the exit-wa v e intensity with p DQE ( ~ q ) before applying shot noise. Subsequently we con volv e with the noise transfer function of the detector to yield the final intensity . A notable difference in the simulations and in practice is the f act that for cryo-EM usually no binning is applied to maximize the imaged area and increase throughput. Therefore, the also high spatial-frequency re gions with low values of the DQE and NTF are used for image formation. For ptychography on the other hand the detector can be heavily binned until the scanning beam still fits into the real-space patch gi ven by r d · N pix . W e therefore apply a 14x binning to the dif fraction patterns and crop to a size of 256 × 256 , i.e. we only use the detector up to 6 . 7 % Nyquist frequency . The real-space patch co vered by the detector is then 51 nm at for a r d = 2 ˚ A , more than necessary to fit a 10 nm size beam. This leads to a near-constant DQE and a near unity NTF , so that there is no necessity to include them in the ptychography reconstructions, while we still include them in the simulation of the dif fraction data. W e note, howe ver that a con v olution with a detector transfer function can be modeled with a partially coherent beam if necessary , as demonstrated in [ 38 , 39 ]. W e choose the Gatan K2 Summit as the detector for our simulations because it has the highest published DQE and MTF values at lo w spatial frequencies at 300 keV [ 40 ]. W e note that other direct detection cameras with f aster readout may be more suitable for ptychographic scanning e xperiments [ 41 , 42 ], but characteristics for these cameras at 300 keV are either not published or worse than the K2 Summit. Additionally , the use of the K2 Summit for both ptychography and phase-contrast TEM simulations simplifies a direct comparison between the two methods. The final model for the formation of the intensity on the detector is then I 0 ( ~ q ) = | F [ ψ exi t ( ~ r )] | 2 (1) for the diffraction pattern and I 0 ( ~ q ) = | F − 1 [ F [ ψ exi t ( ~ r )] · CTF ( ~ q )] | 2 (2) for the cryo-EM image. The intensity after detection is modeled as I ( ~ q ) = F − 1 h F h Poisson F − 1 h F [ I 0 ( ~ q )] · p DQE ( ~ q ) ii · NTF ( ~ q ) i , (3) where NTF is the noise transfer function of the detector [ 43 ]. Single-particle reconstructions Fig. 2 shows a comparison of lo w-dose ptychography reconstructions with currently used methods for single-particle imaging with electrons: defocus-based cryo-EM and Zernike phase contrast cryo-EM with a V olta phase-plate. W e choose ex emplary doses of 5 e − / ˚ A 2 as the typical threshold where the highest resolution details are destro yed [ 44 ] and 20 e − / ˚ A 2 as a typical dose at which experiments are performed. W e have reversed the contrast in the cryo-EM images to simplify the visual comparison with the ptychography reconstructions. It can be seen that ptychography clearly produces the best images for the larger particles at both doses, while it giv es roughly the same results as Zernik e phase-contrast for v ery small particles like hemoglobin. T o quantitativ ely assess the image quality , we hav e computed both the Fourier Shell Correlation (FRC) [ 45 ] and the average signal-to-noise ratio (SNR) in Fig. 2 h)-i). As ground truth for the cryo-EM images we use the electron counts in a noiseless, aberration-free image. W e choose here the 1-bit criterion as a resolution threshold, as the point where the SNR drops to 1 [ 45 ]. W ith this criterion we achiev e resolutions of 14 . 6 ˚ A and 11 . 5 ˚ A for hemoglobin at 5 e − / ˚ A 2 and 20 e − / ˚ A 2 . For 20S proteasome 12 . 7 ˚ A and 10 . 1 ˚ A ; and for human ribosome 11 . 5 ˚ A and 7 . 9 ˚ A respectiv ely at doses of 5 e − / ˚ A 2 and 20 e − / ˚ A 2 . Looking at the a verage SNR v alues, our ptychograph y approach seems to ha ve the biggest adv antages for particles with molecular weight of several 100 kDa and larger . Here, the improv ements over con ventional methods are 8 . 8 dB for 20S proteasome at 20 e − / ˚ A 2 and 7 . 2 dB for human ribosome at 20 e − / ˚ A 2 , almost an order of magnitude improvement. For the ribosome, the signal is strong enough that the oscillating contrast transfer of the defocus-based method is nicely visible in the FRC. 3/ 14 hemoglobin 20S proteasome human ribosome ground truth (1a) (2a) (3a) (1b) (2b) (3b) (3c) (3d) (3e) (3f) (3g) (2c) (2d) (2e) (2f) (2g) (1c) (1e) (1d) (1f) 1g) 2 0 e − / Å 2 phase-plate TEM defocused TEM ptychography phase-plate TEM defocused TEM ptychography 20S proteasome human ribosome 0 . 0 0 0 . 0 5 0 . 1 0 0 . 1 5 0 . 2 0 0 . 2 5 0 . 3 0 q [ Å − 1 ] 0 . 0 0 . 2 0 . 4 0 . 6 0 . 8 1 . 0 F RC d e f o c u s e d TEM @ 5 e − / Å 2 , SNR = - 8 . 9 d B d e f o c u s e d TEM @ 2 0 e − / Å 2 , SNR = - 3 . 8 d B p h a s e -p l a t e TEM @ 5 e − / Å 2 , SNR = - 9 . 4 d B p h a s e -p l a t e TEM @ 2 0 e − / Å 2 , SNR = - 3 . 3 d B p t y c h o g r a p h y @ 5 e − / Å 2 , SNR = - 0 . 6 d B p t y c h o g r a p h y @ 2 0 e − / Å 2 , SNR = 3 .9 d B 1 -b i t c r i t e r i o n (j) 0 . 0 0 0 . 0 5 0 . 1 0 0 . 1 5 0 . 2 0 0 . 2 5 0 . 3 0 q [ Å − 1 ] 0 . 0 0 . 2 0 . 4 0 . 6 0 . 8 1 . 0 F RC d e f o c u s e d TEM @ 5 e − / Å 2 , SNR = - 1 1 .3 d B d e f o c u s e d TEM @ 2 0 e − / Å 2 , SNR = - 5 . 8 d B p h a s e -p l a t e TEM @ 5 e − / Å 2 , SNR = - 1 1 .5 d B p h a s e -p l a t e TEM @ 2 0 e − / Å 2 , SNR = - 5 . 5 d B p t y c h o g r a p h y @ 5 e − / Å 2 , SNR = - 1 0 .3 d B p t y c h o g r a p h y @ 2 0 e − / Å 2 , SNR = 3 .3 d B 1 -b i t c r i t e r i o n (i) 0 . 0 0 0 .0 5 0 . 1 0 0 .1 5 0 . 2 0 0 .2 5 0 . 3 0 q [ Å − 1 ] 0 . 0 0 . 2 0 . 4 0 . 6 0 . 8 1 . 0 F RC d e f o c u s e d TEM @ 5 e − / Å 2 , SNR = - 2 1 . 9 d B d e f o c u s e d TEM @ 2 0 e − / Å 2 , SNR = - 1 5 . 9 d B p h a s e -p l a t e TEM @ 5 e − / Å 2 , SNR = - 2 2 . 5 d B p h a s e -p l a t e TEM @ 2 0 e − / Å 2 , SNR = - 1 6 . 5 d B p t y c h o g r a p h y @ 5 e − / Å 2 , SNR = - 2 1 . 5 d B p t y c h o g r a p h y @ 2 0 e − / Å 2 , SNR = - 1 9 . 8 d B 1 -b i t c r i t e r i o n (h) hemoglobin Figure 2. Cryo-electron ptychography reconstructions from simulated data and simulated cryo-EM images for different doses and 3 macromolecules with gro wing molecular weights in columns 1-3. Row a): Phase of the transmission function, the ground truth for the ptychography reconstructions. The scale bar abov e the figures is in rad. Rows b) and e): ptychography reconstruction at doses of 5 e − / ˚ A 2 and 20 e − / ˚ A 2 . Rows c) and f): Simulated cryo-EM image with a defocus of 1 . 6 µm at a dose of 5 e − / ˚ A 2 and 20 e − / ˚ A 2 . Rows d) and g): Simulated cryo-EM image with a Zernike phase plate and a defocus of 50 nm at doses of 5 e − / ˚ A 2 and 20 e − / ˚ A 2 . Column ( 1 ) hemoglobin, column ( 2 ) 20S proteasome, column ( 3 ) human ribosome. The scale bar is 10 nm. Signal-to-noise ratio ov er spatial frequency for h) hemoglobin, i) 20S proteasome, j) human ribosome. Effect of avera ging In single-particle 3D cryo-EM, a large ensemble of 2D images is collected and then oriented and a veraged in three dimensions to improve the signal-to-noise ratio. A 3D reconstruction from ptychographic data is out of the scope of this paper , because dedicated algorithms need to be developed to achie ve optimal results. The reconstructed 2D images do not obe y the same noise statistics as cryo-EM micrographs and therefore the use of standard softw are would lead to suboptimal results. In the best case, a 3D model can be reconstructed directly from the raw dif fraction data, while coarse orientation alignment could be done in real space from 2D reconstructions shown here. 4/ 14 T o giv e a rough estimate ho w the resolution and SNR of our algorithm scales with av eraging multiple datasets, we perform here an computer experiment and av erage over reconstructions of 15 datasets where the particles are in identical orientation. W e choose an average of 15 because there is no significant improvement when averaging more datasets. Fig. 3 a)-f) shows images of the averaged reconstructions of our three samples, at 5 e − / ˚ A 2 and 20 e − / ˚ A 2 respectiv ely . Comparing the SNR v alues with the single-particle reconstructions, we see for almost all doses and samples more than an order of magnitude increase in SNR. For hemoglobin, the a v erage SNR increases from − 16 . 8 dB to − 0 . 7 dB at 5 e − / ˚ A 2 and from − 14 dB to 3 . 2 dB at 20 e − / ˚ A 2 . For the 20S proteasome the SNR increases from − 5 . 5 dB to 10 dB at 5 e − / ˚ A 2 and from 6 . 2 dB to 12 dB at 20 e − / ˚ A 2 . For the human ribosome the SNR increases from − 0 . 6 dB to 13 . 7 dB at 5 e − / ˚ A 2 and from 3 . 9 dB to 16 . 1 dB at 20 e − / ˚ A 2 . All in all a veraging of reconstructions giv es lar ger improv ements for very lo w doses. W e also FRC curves for the a veraged reconstructions of tw o independently created data sets to giv e a resolution estimate. W e use here the 1/2-bit resolution threshold discussed in [ 45 ], which gi ves a slightly more conserv ati ve estimate than the 0.143-criterion commonly used in av eraged reconstructions for cryo-EM. With averaging, a resolution of 8 ˚ A is achiev ed for hemoglobin, 4 . 4 ˚ A is achiev ed for 20S proteasome and 3 . 9 ˚ A for human ribosome at a dose of 20 e − / ˚ A 2 . a b c d e f 0 . 0 0 0. 0 5 0 . 1 0 0. 1 5 0 . 2 0 0. 2 5 0 . 3 0 q [Å 1 ] 0 . 0 0 . 2 0 . 4 0 . 6 0 . 8 1 . 0 FR C 0 .1 4 3 1 3 . 2 Å 1 0 . 4 Å 8 . 0 Å 1 5 · 5 e / Å 2 , SNR = - 0 . 7 d B 1 5 · 1 0 e / Å 2 , SNR = 1 .8 d B 1 5 · 2 0 e / Å 2 , SNR = 3 .2 d B 1 / 2 - b i t t h r e s h o l d g 0 . 0 0 0 .0 5 0 . 1 0 0 .1 5 0 . 2 0 0 . 2 5 0 . 3 0 q [Å 1 ] 0 . 0 0 . 2 0 . 4 0 . 6 0 . 8 1 . 0 FR C 0 .1 4 3 9 . 0 Å 6 . 9 Å 4 . 4 Å 1 5 · 5 e / Å 2 , SNR = 1 0 .2 d B 1 5 · 1 0 e / Å 2 , SNR = 1 2 .5 d B 1 5 · 2 0 e / Å 2 , SNR = 1 2 .4 d B 1 / 2 - b i t t h r e s h o l d h hemoglobin 20S proteasome human ribosome 2 0 e / Å 2 0 . 0 0 0 .0 5 0 . 1 0 0 .1 5 0 . 2 0 0 .2 5 0 . 3 0 q [Å 1 ] 0 . 0 0 . 2 0 . 4 0 . 6 0 . 8 1 . 0 FR C 0 .1 4 3 6 . 2 Å 4 . 7 Å 4 . 1 Å 1 5 · 5 e / Å 2 , SNR = 1 3 .7 d B 1 5 · 1 0 e / Å 2 , SNR = 1 4 .8 d B 1 5 · 2 0 e / Å 2 , SNR = 1 6 . 1 d B 1 / 2 - b i t t h r es h o l d i Figure 3. A verage ov er 15 ptychographic reconstructions from independent data sets for a) hemoglobin with 5 e − / ˚ A 2 , b) hemoglobin with 20 e − / ˚ A 2 , c) proteasome 20S with 5 e − / ˚ A 2 , d) proteasome 20S with 20 e − / ˚ A 2 , e) human ribosome with 5 e − / ˚ A 2 , f) human ribosome with 20 e − / ˚ A 2 . FRC of av eraged reconstructions from independent data sets for g) hemoglobin, h) proteasome 20S i) human ribosome. Probe and dose dependence It is well kno wn that the phase profile of the ptychographic probe can hea vily influence the reconstruction quality [ 46 , 47 , 48 , 49 , 50 ]. Here we numerically test three dif ferent probes, depicted in Fig. 4 , and their influence on the 5/ 14 reconstruction SNR at lo w and high doses: 1) a standard defocused probe with defocus aberration of 400 nm, 2) a defocused Fresnel Zone Plate (FZP), and 3) a randomized probe generated by a holographic phase plate and a focusing lens. Fig. 4 depicts these probe in real and Fourier space and typical dif fraction patterns at infinite dose and low dose. The FZP was recently suggested as a phase modulator for bright-field STEM [ 51 ], because its simple phase modulation allows analytical retriev al of linear phase contrast. Howe ver , diffracti v e optics typically have imperfections due to the manufacturing process which introduce errors and dose inef ficienc y if the phase modulation is obtained by a simple fitting procedure. Iterativ e ptychography algorithms allo w for the simultaneous retrie v al of the probe wa ve function [ 17 , 18 ] and therefore of fers full flexibility in the design of the phase profile. Empirically , probes with a diffuse phase profile result in better reconstructions, therefore we test as a third probe a random illumination generated by a holographic phase plate and a focusing lens. Fig. 5 shows the SNR of the three proposed probes as a function of spatial frequenc y . It can be seen that at very low doses of 5 e − / ˚ A 2 , the randomized probe achiev es the best SNR. At high doses of 1380 e − / ˚ A 2 , the SNR of the randomized probe at very lo w frequencies is higher up to a factor of 1 × 10 4 than the defocused probe and the FZP , whereas the performance at high resolution is only slightly better . W e gi ve two qualitati ve e xplanations of this fact, b ut emphasize that a theory for optimal measurement design in ptychography and a practically feasible implementation of it is still outstanding and may drastically improv e upon the results presented here. 0 1 2 3 4 5 6 1a 2a 3a 3b 2b 1b 0 .0 0 .2 0 .4 0 .6 0 .8 1 .0 1c 3d 1d 2d 2c 3c Figure 4. Different probes e v aluated in this paper and corresponding diffraction patterns. Ro w 1: defocused beam with defocus aberration of 400nm, con ver gence half-angle 7 . 85 mrad ; row 2: defocused beam created by a Fresnel zone plate, 600 nm from focus; row 3: randomized beam, generated by a holographic phase plate and focused by a con ventional lens. Column a) beam in real space, at the sample position, scale bar is 10 nm ; column b) beam at the probe forming aperture, scale bar is 3 . 85 mrad ; column c) diffraction pattern of human ribosome at unlimited dose, normalized to the maximum intensity; column d) diffraction pattern for a scan with 589 acquisitions, at an electron dose of 20 e − / ˚ A 2 . The inset in 1a sho ws the color wheel that is used to represent amplitude and phase in columns a) and b). 6/ 14 0 .0 0 0 .0 5 0 .1 0 0 .1 5 0 .2 0 0 .2 5 0 .3 0 0 .3 5 q [Å 1 ] 10 3 10 2 10 1 10 0 10 1 10 2 10 3 10 4 10 5 10 6 10 7 10 8 10 9 S N R Rose c r it e r io n p t y c ho g r a p hy @ 5 e / Å 2 , PS NR = 1 9 .0 d B p t y c ho g r a p hy @ 2 0 e / Å 2 , PS NR = 2 4 .3 d B p t y c ho g r a p hy @ 8 0 e / Å 2 , PS NR = 2 9 .1 d B p t y c ho g r a p hy @ 3 2 0 e / Å 2 , PS NR = 3 3 .3 d B p t y c ho g r a p hy @ 1 3 8 0 e / Å 2 , PS NR = 3 5 .7 d B (c) 0 .0 0 0 .0 5 0 .1 0 0 .1 5 0 .2 0 0 .2 5 0 .3 0 0 .3 5 q [Å 1 ] 10 3 10 2 10 1 10 0 10 1 10 2 10 3 10 4 10 5 10 6 10 7 10 8 10 9 S N R Rose c r it e r io n p t y c ho g r a p hy @ 5 e / Å 2 , PS NR = 1 3 .8 d B p t y c ho g r a p hy @ 2 0 e / Å 2 , PS NR = 2 0 .1 d B p t y c ho g r a p hy @ 8 0 e / Å 2 , PS NR = 2 5 .9 d B p t y c ho g r a p hy @ 3 2 0 e / Å 2 , PS NR = 3 1 .2 d B p t y c ho g r a p hy @ 1 3 8 0 e / Å 2 , PS NR = 3 5 .3 d B (b) 0 .0 0 0 .0 5 0 .1 0 0 .1 5 0 .2 0 0 .2 5 0 .3 0 0 .3 5 q [Å 1 ] 10 3 10 2 10 1 10 0 10 1 10 2 10 3 10 4 10 5 10 6 10 7 10 8 10 9 SNR Rose c r it e r io n p t y c ho g r a p hy @ 5 e / Å 2 , PS NR = 9 . 7 d B p t y c ho g r a p hy @ 2 0 e / Å 2 , PS NR = 1 6 .5 d B p t y c ho g r a p hy @ 8 0 e / Å 2 , PS NR = 2 0 .9 d B p t y c ho g r a p hy @ 3 2 0 e / Å 2 , PS NR = 2 3 .6 d B p t y c ho g r a p hy @ 1 3 8 0 e / Å 2 , PS NR = 2 5 .9 d B (a) Figure 5. Signal to noise ratios of reconstructions of the human ribosome at different radiation doses using a) the defocused probe, b) the Fresnel zone plate, c) the randomized probe Methods Mathematical framew ork of Ptychograph y The two-dimensional complex transmission function of the object is discretized as a n 1 × n 2 matrix and denoted as T : D n 1 × n 2 r d → C , where r d > 0 is the dif fraction limited length scale. The object is illuminated by a small beam with known distribution, and discretized as a m 1 × m 2 matrix denoted as ψ : D M r d → C . For simplicity , in this paper we only consider the case n 1 = n 2 and m 1 = m 2 , i.e. a uniform discretization in both axes. In the experiment, the beam is mov ed ov er the sample to positions ~ r i and illuminates K > 1 subregions to obtain K dif fraction images. I i ( ~ q ) = | F [ ψ ( ~ r + ~ r i ) · T ( ~ r )] | 2 , i ∈ { 0 , ..., K } , (4) where the discretized real space coordinates are discretized in steps of r d and reciprocal space coordinates are discretized in steps of 1 m i · r d . Mathematically , ptychography can be understood as a special case of the generalized phase r etrieval pr oblem : giv en a phase-less vector of measurements y ∈ R m + find a complex vector z ∈ C n such that y = | A z | 2 , (5) where A : C n → C m is an arbitrary linear operator . W e follo w here the notations in [ 46 ] to formulate ptychography as a generalized phase retriev al problem. First, we vectorize the transmission function as T V ∈ C N with N = n 1 · n 2 ∈ N . W e introduce the matrix R ( i ) ∈ R M × N , which extracts an m 1 × m 2 sized area centered at position ~ r i from T . W ith these notations in place, the relationship between the noise-free dif fraction measurements collected in a ptychography experiment and T V can be represented compactly as I = | FQT V | 2 = | PT V | 2 , (6) (7) where P is constructed by cropping K regions from T , multiplying by the incoming beam, and a 2D discrete Fourier transform F , i.e. P = FQ . I ∈ R K M z }| { I 1 . . . I K M = F ∈ C K M × K M z }| { F · · · 0 . . . . . . . . . 0 · · · F Q ∈ C K M × N z }| { diag ( ψ ) R ( 1 ) . . . diag ( ψ ) R ( K ) T V 2 (8) . (9) The matrix P ∈ C K M × N is sometimes called design matrix because its entries determine the measurement outcome and reflect the experimental design. In the last decades many algorithms to solve this problem have been de vised, of which we only revie w a small fraction with regards to low-dose reconstruction in the following section. For the subsequent analysis, we denote the KM row v ectors of P as p i . Bay esian optimization with truncated gradients The most prominent iterativ e algorithms to solve the ptychographic phase retrie val problems are the dif ference map (DM) [ 12 ] algorithm and the extended ptychographic iterati ve engine (ePIE) [ 18 ]. The dif ference map belongs to the family of algorithms which use projections onto non-con ve x sets to reach a fix-point, the solution lying at the intersection of the two sets. It can be shown that the standard algorithm of alternating projections is equiv alent to steepest descent optimization with a Gaussian likelihood and is not suited for low-dose reconstructions [ 46 ] 7/ 14 because the Poisson distribution arising from discretized count e v ents differs too strongly from a Gaussian at v ery low electron counts. While this equality does not hold for the more elaborate projection algorithms lik e DM and relaxed a veraged alternating reflections (RAAR) [ 52 ], they also fail in practice at lo w doses [ 23 , 53 ], and statistical reconstruction methods have to be used. Thibault and Guizar-Sicairos [ 22 ] have analyzed maximum likelihood methods in conjunction with a conjugate gradient update rule as a refinement step, after the DM algorithm has con ver ged. They demonstrate improved SNR compared to the DM algorithm alone. They note, howe ver , that starting directly with maximum likelihood optimization often poses con v ergence problems. The PIE algorithm can be formulated as maximum-a-posteriori (MAP) optimization with a Gaussian likelihood function and an independently weighted Gaussian prior of the object change [ 38 , 53 ], combined with a stochastic gradient-like update rule. The noise performance of the PIE algorithm has been in vestigated in [ 53 ] and found to be worse than the Poissonian lik elihood model at lo w counting statistics. While practically very robust, both algorithms can get stuck in local minima and until recently [ 46 ], no proof of con v ergence to a global minimum existed. [ 53 ] suggest to use a global gradient update at the start to avoid stagnation, and [ 13 ] use a restarted version with the stochastic gradient update rule, after remov al of phase v ortex artifacts. Due to the lack of algorithms with prov able con v erge guarantees, the mathematical community has recently picked up the problem and a host of new algorithms with prov able con ver gence has been dev eloped, which we do not elaborate on here but point the interested reader to the summary articles [ 54 , 55 ] and the article [ 56 ], which refers to the most recent dev elopments. Here we focus on dev elopments which specifically target lo w-dose applications. Notable in this area is the work by Katko vnik et al. [ 57 ], which in addition to the maximum likelihood estimate introduces a transform-domain sparsity constraint on the object and optimizes two objecti ve functions in an alternating fashion: one for the maximizing the likelihood and one for obtaining a sparse representation of the transmission function. Howe ver , instead of including the Poissonian likelihood directly , an observation filtering step is performed with a Gaussian likelihood. T o obtain a sparse representation of the object, the popular BM3D denoising filter is used. Another recent, similar approach uses dictionary learning to obtain a sparse representation of the transmission function [ 58 ], ho wev er only real-v alued signals are treated. During the writing of this paper , Y ang et al. suggested using the one-step in version technique for low-dose ptychography [ 59 ], howe ver no statistical treatment of the measurement process is included so far , which could be a promising av enue for future research. W e formulate ptychographic phase retriev al as a Bayesian inference problem by introducing the probability of the transmission function T V , giv en a set of measurements y = ( y 1 , y 2 , ... , y K M ) T ∈ R K M + with the Bayes’ rule: P ( T V | y ) = P ( y | T V ) P ( T V ) P ( y ) (10) Since the measurements y i follow the Poisson distrib ution y i ∼ Poisson ( I i ( T V )) , (11) the Likelihood function is gi ven by P ( y i | T V ) = I i ( T V ) y i y i ! e − I i ( T V ) . (12) The prior distribution is usually chosen such that it fav ors realistic solutions, so that noise is suppressed in the reconstructed image. Here we ev aluate two dif ferent models. A simple prior, suggested in [ 38 ], penalizes large gradients in the image with a Gaussian distrib ution on the gradient of the transmission function, which is also kno wn as T ikhonov re gularization. P T ik honov ( T ) = exp ( − µ 0 κ || ∇ T ( ~ r ) || 2 ) = exp ( − µ 0 κ N ∑ i = 1 ( D x T V ) 2 i + ( D y T V ) 2 i ) (13) with κ = 8 N 2 N m || I || 1 chosen as in [ 38 ] to scale the numerical value of the prior to be close to the likelihood. D x and D y are the discrete forward difference operators. The second prior we e valuate is based on the work by Katko vnik et al. [ 57 ] and uses sparse modeling to denoise the transmission function: P s parse ( T V ) = exp ( − µ || T V − T V s parse || 2 ) (14) Here, T V s parse is b uilt up by applying the BM3D collaborati v e filtering algorithm [ 60 ], which we describe here shortly . The BM3D algorithm computes the transform-domain sparse representation in 4 steps: 1 ) it finds the image patches similar to a gi ven image patch and groups them in a 3D block 2 ) 3D linear transform of the 3D block; 3 ) shrinking of the transform spectrum coefficients; 4 ) in verse 3D transformation. As input for the BM3D algorithm we transform T V into hue-saturation-value format, the phase representing hue and the 8/ 14 amplitude representing value, with full saturation. The prior P s parse ( T ) reduces the difference between the denoised version of the current transmission function and the transmission function itself. W e note that during the writing of this manuscript an e xtensiv e e v aluation of denoising phase retrie v al algorithms was published [ 61 ], which also ev aluates BM3D denoising using an algorithm similar to the one presented here and finds superior performance compared to other denoising strate gies such as total v ariation or nonlocal means. W e do not take into account the marginal like lihood P ( y ) due to the high dimensionality of the problem. Giv en the likelihood function and the prior distribution, one can write do wn the objecti ve function for the MAP estimate: T V M AP : = argmin T V L M AP ( T V ) (15) The gradient of the likelihood is gi ven as L ( T V ) = K M ∑ i = 1 | p i T V | 2 − y i log ( | p i T V | 2 ) . (16) and the MAP objectiv e functions are L M AP ( T V ) = − log P ( y | T V ) P ( T V ) P ( y ) = L ( T V ) + µ 0 κ || ∇ T ( ~ r ) || 2 (17) and L BM 3 D − M AP ( T V ) = L ( T V ) + µ 1 || T V − T V s parse || 2 (18) W e calculate the gradient of L M AP ( T ) ∇ L M AP ( T ) = K M ∑ i = 1 2 p i T V 1 − y i | p i T V | 2 p † i + 2 µ 0 κ N ∑ i = 1 ( D x T V ) i + ( D y T V ) i (19) ∇ L BM 3 D − M AP ( T ) = K M ∑ i = 1 2 p i T V 1 − y i | p i T V | 2 p † i + µ 1 T V − T V s parse (20) Since 17 and 18 are non-conv ex functions, there is no guarantee that standard gradient descent conv erges to a global minimum. Recently , a non-conv ex algorithm for the generalized phase retrieval problem with Poisson noise was presented [ 62 ] that prov ably con v er ges to a global minimum with suitable initialization. It introduces a iteration-dependent regularization on the gradients of the likelihood to remo ve terms which ha v e a negati v e effect on the search direction. Namely , it introduces a truncation criterion E i ( T V ) = ( y i − p i T V 2 ≤ α h K M k y − I k 1 p i T V k T V k 2 ) (21) that acts on the gradient of the likelihood and suppresses the influence of measurements that are too incompatible with the reconstruction. The regularized likelihood gradient is then ∇ L E i ( T V ) = K M ∑ i ∈ E i ( T V ) | p i T V | 2 − y i log ( | p i T V | 2 ) . (22) W e compute the next step using conjugate gradient descent [ 63 , 64 ], since this lead to much faster conv er gence compared to the update procedure described in [ 62 ]. Initialization T runcated spectral initialization for ptychography was first proposed by Marchesini et al. [ 46 ], based on the notion that the highest intensities in the diffraction pattern carry the strongest phase information. They compute the phase of the largest eigen vector of the following hermitian operator: 1 | y i | > ε FQ ( Q † Q ) − 1 Q † F † 1 | y i | > ε , (23) where 1 y > ε is an indicator vector of the same dimension as y and ε is chosen so that the largest 20 percent of the intensities are allowed to contribute. The largest eigen v alue of a sparse hermitian matrix can be efficiently computed either with po wer iterations [ 65 ], or with the Arnoldi method [ 66 ]. In [ 62 ], truncated spectral initialization with a truncation rule with 1 | y i | < α 2 0 λ 2 0 is used, with λ 0 = q ∑ K M i = 1 y i . W e found the initialization of Marchesini et al. 9/ 14 to produce better initializations for ptychographic datasets. This may be due to the nature of the ptychographic measurement vectors p i , which are different from the measurement v ectors used in [ 62 ]. W e leav e further analysis of this matter for future research. It is also important to note that the truncated spectral initialization only produces visually correct initial phase to a dose of roughly 100 e − / ˚ A 2 . Fig. 6 b) shows an example initialization for a dose of 100 e − / ˚ A 2 . For doses below this value, we initialized the transmission function with unity transmission and normal-distributed phase with mean 0 . 1 , 0 . 2 , and 0 . 3 for hemoglobin, 20S proteasome and ribosome respecti vely , and variance of 0 . 1 , which usually provides a lo wer starting error than the spectral initialization. W ith this sample-dependent random initialization we found no problem of con v ergence for all algorithms tested in this paper . a b c 0 2 5 5 0 7 5 1 0 0 1 2 5 1 5 0 i t e r a t i o n # 1 0 2 1 0 1 1 0 0 NRMSE BM 3 D -M AP MA P Ti k h o n o v - MA P a Figure 6. a) Con ver gence behavior of dif ferent gradient update rules b) Example for the transmission function initialization T 0 after 70 power iterations, for an electron dose of 100 e − / ˚ A 2 , intensities were truncated at the 80th percentile. c) T V s parse for human ribosome after 60 iterations of BM3D-MAP . Scale bar is 10 nm. Reconstruction parameters All ptychography reconstructions were performed with a probe area overlap of 75 % in real-space, where the probe area is defined by all pixels contributing more than 1 % of the maximum intensity . This corresponds to a step size of roughly 3 nm , depending on the probe used. For the reconstructions sho wn in Fig. 2 a total of 589 diffraction patterns were created using the random illumination. At a dose of 20 e − / ˚ A 2 , this corresponds to 5000 electrons per diffraction pattern. For the re gularization parameters we performed a grid search e v aluating the final NRMSE and found the values to be µ 1 = 4e − 2 , µ 2 = 1e − 1 . W e choose the biorthogonal spline wav elet transform as the linear transform for BM3D as it achiev es the best PSNR for high noise [ 67 ]. Implementation Details The algorithms presented in this paper were implemented with the torch scientific computing framework [ 68 ]. The gradient update routines were adapted from the optim package for torch [ 64 ]. For ef ficient computing on the graphics processing unit (GPU) with complex numbers, the zcutorch library for cuda was dev eloped [ 69 ]. hyperparameter optimization was done with the hypero [ 70 ] package for torch. For BM3D denoising we use the C++ implementation [ 71 ]. The code was run on an Intel i7-6700 processor with 32GB RAM and a NV idia T itan X GPU with 12GB RAM. The runtime for optimization with L M AP was 26 s , and for optimization with L BM 3 D − M AP 35 s , because the BM3D algorithm used here is not implemented on the GPU and the BM3D denoising is computationally more intensiv e. Conclusion In this paper we ha ve demonstrated via numerical e xperiments the possibility to retrie v e high resolution electron transmission phase information of biological macromolecules using ptychography and Bayesian optimization. Using the methods presented in this paper , it should be possible to achie ve resolutions belo w 1 nm for true single particle imaging of large molecular comple xes like human ribosome and a resolution around 4 ˚ A with simple av eraging of 15 datasets. W e have gi v en a detailed explanation of the optimization and initialization procedures used and hav e emphasized the importance of choosing an appropriate illumination function. W e note that, while the high data redundancy in a ptychographic dataset empirically makes it experimentally very robust, there is much room for improv ements in terms of measurement complexity . For the results presented here, the measurement dimension KM is larger than the problem dimension N by a factor of at least 30, while the theoretical limit for successful phase retriev al is KM = 4N [ 72 ]. By reducing the number of measurements the variance of each indi vidual measurement could be reduced, yielding an improved SNR in the reconstruction. Therefore the dev elopment of an optimized experimental scheme including design of the illumination function and scanning scheme is a promising direction of research and may enable significant improv ement to the results presented here. 10/ 14 W e would like to point out two obstacles that one may hav e to ov ercome in the experimental realization of our results. Firstly the best results are to be expected when recoding zero-loss dif fraction patterns with the use of an energy filter . The energy filter may introduce phase distortions into the dif fraction patterns which may need to be accounted for in the reconstruction algorithm. One could achie ve this by first reconstructing the incoming w av efront in characterization experiment without ener gy filter and then reconstructing the aberration induced by the ener gy filter with a fix ed, kno wn incoming wa vefront. Secondly , although beam induced movements are expected to be reduced by a lar ge amount due to spot-scanning, the remaining mo vement may cause problems in the reconstruction. Statistically stationary sample mov ements can be accounted for in the reconstruction algorithm [ 23 , 73 ], but beam induced motions are likely to be non-stationary , and dedicated algorithms may need to be dev eloped to account for it. Cryogenic ptychographic imaging of biological samples is also being developed in the X-ray sciences [ 74 ], and our results could equally be implemented there to improv e the dose-ef fecti veness. Finally , the methods presented here may find application in electron phase imaging of radiation-sensitive samples under non-cryogenic conditions and the incorporation of Bayesian methods into in-focus ptychographic reconstruction procedures [ 16 , 59 ] may provide similar gains in SNR as the ones discussed here while also keeping the analytical capabilities of traditional STEM imaging. Ackno wledgements This work was funded by the Max Planck Society through institutional support. P .M.P . ackno wledges support from the International Max Planck Research School for Ultrafast Imaging & Structural Dynamics. A uthor contrib utions statement P .M.P ., R.J.D.M. and R.B. conceived the idea. P .M.P and W .Q wrote the simulation code. P .M.P wrote the ptychographic reconstruction code. P .M.P wrote the paper with input and discussions from R.B, G.K, and R.J.D.M. Competing financial interests The authors declare no competing financial interests. Reference resolution e − / ˚ A 2 D’Alfonso et al. [ 28 ] ∼ 1 . 5 ˚ A 3 . 98 × 10 4 Y ang et al. [ 27 ] atomic 1 . 3 × 10 4 Putkunz et al.[ 26 ] ∼ 1 ˚ A 9 . 2 × 10 6 Humphry et al. [ 30 ] ∼ 2 . 3 ˚ A 3 . 33 × 10 3 T able 1. List of previously published electron ptychograph y experiments and used electron dose References 1. Bartesaghi, A., Matthies, D., Banerjee, S., Merk, A. & Subramaniam, S. Structure of β -galactosidase at 3.2- ˚ A resolution obtained by cryo-electron microscopy . PNAS 111 , 11709–11714 (2014). 00063. 2. Groll, M. et al. Structure of 20S proteasome from yeast at 2.4 ˚ A resolution. Nature 386 , 463–471 (1997). 02142. 3. Scheres, S. H. W . RELION: Implementation of a Bayesian approach to cryo-EM structure determination. Journal of Structur al Biology 180 , 519–530 (2012). 00587. 4. Bai, X.-c., Fernandez, I. S., McMullan, G. & Scheres, S. H. Ribosome structures to near-atomic resolution from thirty thousand cryo-EM particles. eLife 2 , e00461 (2013). 00255 PMID: 23427024. 5. Subramaniam, S., K ¨ uhlbrandt, W . & Henderson, R. CryoEM at IUCrJ: A ne w era. IUCrJ 3 , 3–7 (2016). 00013. 6. Bai, X.-c., McMullan, G. & Scheres, S. H. W . How cryo-EM is re v olutionizing structural biology . T r ends in Biochemical Sciences 40 , 49–57 (2015). 00152. 7. Scheres, S. H. Beam-induced motion correction for sub-megadalton cryo-EM particles. eLife 3 , e03665 (2014). 00149. 8. Glaeser, R. M. How good can cryo-EM become? Nat Meth 13 , 28–32 (2016). 00012. 11/ 14 9. Danev , R. & Baumeister , W . Cryo-EM single particle analysis with the V olta phase plate. eLife 5 , e13046 (2016). 00016 PMID: 26949259. 10. Danev , R., Buijsse, B., Khoshouei, M., Plitzko, J. M. & Baumeister , W . V olta potential phase plate for in-focus phase contrast transmission electron microscopy . PNAS 111 , 15635–15640 (2014). 00077 PMID: 25331897. 11. Zernike, F . How I Disco vered Phase Contrast. Science 121 , 345–349 (1955). 00490. 12. Thibault, P . et al. High-resolution scanning x-ray diffraction microscopy . Science 321 , 379–82 (2008). 00619. 13. Maiden, A. M., Sarahan, M. C., Stagg, M. D., Schramm, S. M. & Humphry , M. J. Quantitative electron phase imaging with high sensitivity and an unlimited field of vie w . Scientific Reports 5 , 14690 (2015). 00005. 14. Diaz, A. et al. Quantitative x-ray phase nanotomography . Phys. Rev . B 85 , 1–4 (2012). 15. Hoppe, W . Trace structure analysis, ptychograph y , phase tomography . Ultramicr oscopy 10 , 187–198 (1982). 00043. 16. Rodenbur g, J. M. The phase problem, microdiffraction and wa velength-limited resolution — a discussion. Ultramicr oscopy 27 , 413–422 (1989). 00018. 17. Thibault, P ., Dierolf, M., Bunk, O., Menzel, A. & Pfeiffer , F . Probe retriev al in ptychographic coherent diffracti v e imaging. Ultramicr oscopy 109 , 338–43 (2009). 18. Maiden, A. M. & Rodenburg, J. M. An improv ed ptychographical phase retriev al algorithm for diffracti ve imaging. Ultramicr oscopy 109 , 1256–62 (2009). 19. Guizar-Sicairos, M. & Fienup, J. R. Phase retriev al with transverse translation di versity: A nonlinear optimiza- tion approach. Opt. Expr ess, OE 16 , 7264–7278 (2008). 00219. 20. Zhang, F . et al. T ranslation position determination in ptychographic coherent dif fraction imaging. Opt. Expr ess, OE 21 , 13592–13606 (2013). 00085. 21. Maiden, A. M., Humphry , M. J., Sarahan, M. C., Kraus, B. & Rodenburg, J. M. An annealing algorithm to correct positioning errors in ptychography . Ultramicr oscopy 120 , 64–72 (2012). 00068. 22. Thibault, P . & Guizar-Sicairos, M. Maximum-likelihood refinement for coherent diffracti v e imaging. New J . Phys. 14 , 063004 (2012). 00081. 23. Pelz, P . M. et al. On-the-fly scans for X-ray ptychography . Appl. Phys. Lett. 105 , 251101 (2014). 00011. 24. Clark, J. N., Huang, X., Harder , R. J. & Robinson, I. K. Dynamic Imaging Using Ptychography. Phys. Rev . Lett. 112 , 113901 (2014). 00004. 25. Marchesini, S., Schirotzek, A., Y ang, C., W u, H.-t. & Maia, F . Augmented projections for ptychographic imaging. In verse Pr oblems 29 , 115009 (2013). 00027. 26. Putkunz, C. T . et al. Atom-Scale Ptychographic Electron Dif fractiv e Imaging of Boron Nitride Cones. Phys. Rev . Lett. 108 , 073901 (2012). 00027. 27. Y ang, H. et al. Simultaneous atomic-resolution electron ptychography and Z-contrast imaging of light and heavy elements in comple x nanostructures. Nature Communications 7 , 12532 (2016). 00001. 28. D’Alfonso, A. J., Allen, L. J., Saw ada, H. & Kirkland, A. I. Dose-dependent high-resolution electron ptychography . Journal of Applied Physics 119 , 054302 (2016). 00000. 29. Lubk, A. & R ¨ oder , F . Phase-space foundations of electron holography . Phys. Re v . A 92 , 033844 (2015). 00000. 30. Humphry , M., Kraus, B. & Hurst, A. Ptychographic electron microscop y using high-angle dark-field scattering for sub-nanometre resolution imaging. Nat. ldots 3 , 730–737 (2012). 00123. 31. Bullough, P . & Henderson, R. Use of spot-scan procedure for recording lo w-dose micrographs of beam-sensiti ve specimens. Ultramicr oscopy 21 , 223–230 (1987). 00088. 32. Brink, J., Chiu, W . & Dougherty , M. Computer-controlled spot-scan imaging of crotoxin complex crystals with 400 keV electrons at near -atomic resolution. Ultramicr oscopy 46 , 229–240 (1992). 00034. 33. Downing, K. H. Spot-scan imaging in transmission electron microscopy . Science 251 , 53–59 (1991). 00000 PMID: 1846047. 12/ 14 34. Fermi, G., Perutz, M. F ., Shaanan, B. & Fourme, R. The crystal structure of human deoxyhaemoglobin at 1.74 A resolution. J. Mol. Biol. 175 , 159–174 (1984). 00000 PMID: 6726807. 35. Anger , A. M. et al. Structures of the human and Drosophila 80S ribosome. Natur e 497 , 80–85 (2013). 00169. 36. V ulo vi ´ c, M. et al. Image formation modeling in cryo-electron microscopy . Journal of Structural Biology 183 , 19–32 (2013). 00027. 37. Kirkland, E. Advanced Computing in Electr on Micr oscopy (2010). URL http://books.google.com/ books?hl=en&lr=&id=YscLlyaiNvoC&oi=fnd&pg=PR5&dq=Advanced+Computing+in+ Electron+Microscopy&ots=KA3FApM1kW&sig=T6oRRFaEMUJuL3d3_GlC2dynwMs . 00000. 38. Thibault, P . & Menzel, A. Reconstructing state mixtures from dif fraction measurements. Natur e 494 , 68–71 (2013). 00135. 39. Enders, B. et al. Ptychography with broad-bandwidth radiation. Appl. Phys. Lett. 104 , 171104 (2014). 00019. 40. McMullan, G., Faruqi, A. R., Clare, D. & Henderson, R. Comparison of optimal performance at 300 keV of three direct electron detectors for use in low dose electron microscopy . Ultramicr oscopy 147 , 156–163 (2014). 00000 arXiv: 1406.1389. 41. Ryll, H. et al. A pnCCD-based, fast direct single electron imaging camera for TEM and STEM. J. Inst. 11 , P04006 (2016). 00003. 42. McMullan, G., Chen, S., Henderson, R. & Faruqi, A. R. Detectiv e quantum ef ficiency of electron area detectors in electron microscopy . Ultramicr oscopy 109 , 1126–1143 (2009). 00137. 43. Meyer , R. R. & Kirkland, A. I. Characterisation of the signal and noise transfer of CCD cameras for electron detection. Micr osc. Res. T ech. 49 , 269–280 (2000). 00112. 44. Stark, H., Zemlin, F . & Boettcher, C. Electron radiation damage to protein crystals of bacteriorhodopsin at different temperatures. Ultramicr oscopy 63 , 75–79 (1996). 00060. 45. Heel, M. V . & Schatz, M. Fourier shell correlation threshold criteria q. J. Struct. Biol. 151 , 250–262 (2005). 00000. 46. Marchesin, S., T u, Y . & W u, H.-t. Alternating Projection, Ptychographic Imaging and Phase Synchronization. arXiv Pr epr . arXiv1402.0550 1–29 (2014). URL . 00000, arXiv: 1402.0550v1 . 47. Li, P . et al. Multiple mode x-ray ptychography using a lens and a fixed dif fuser optic. J. Opt. 18 , 054008 (2016). 00001. 48. Maiden, A. M., Morrison, G. R., Kaulich, B., Gianoncelli, A. & Rodenb urg, J. M. Soft X-ray spectromicroscopy using ptychography with randomly phased illumination. Nature Communications 4 , 1669 (2013). 00042. 49. Guizar-Sicairos, M. et al. Role of the illumination spatial-frequency spectrum for ptychography . Phys. Rev . B 86 , 100103 (2012). 00030. 50. Li, P ., Edo, T . B. & Rodenb urg, J. M. Ptychographic inv ersion via W igner distribution decon volution: Noise suppression and probe design. Ultramicr oscopy 147 , 106–113 (2014). 00007. 51. Ophus, C. et al. Efficient linear phase contrast in scanning transmission electron microscopy with matched illumination and detector interferometry . Nat Commun 7 , 10719 (2016). 00000. 52. Luke, D. R. Relaxed av eraged alternating reflections for dif fraction imaging. Inver se Pr oblems 21 , 37 (2005). 00000. 53. Godard, P ., Allain, M., Chamard, V . & Rodenburg, J. Noise models for low counting rate coherent diffrac- tion imaging. Opt. Expr ess 20 , 25914–34 (2012). URL http://www.ncbi.nlm.nih.gov/pubmed/ 23187408 . 00024. 54. Jaganathan, K., Eldar , Y . C. & Hassibi, B. Phase Retriev al: An Overvie w of Recent Developments. Arxiv (2015). URL . 00024, 1510.07713 . 55. Shechtman, Y . et al. Phase Retrie val with Application to Optical Imaging: A contemporary ov erview . IEEE Signal Pr ocessing Magazine 32 , 87–109 (2015). 00016. 13/ 14 56. Sun, J., Qu, Q. & Wright, J. A geometric analysis of phase retrie v al. In 2016 IEEE International Symposium on Information Theory (ISIT) , 2379–2383 (2016). 00042. 57. Katkovnik, V . & Astola, J. Sparse ptychographical coherent dif fractiv e imaging from noisy measurements. J. Opt. Soc. Am. A. Opt. Image Sci. V is. 30 , 367–79 (2013). 00004. 58. T illmann, A. M., Eldar , Y . C. & Mairal, J. DOLPHIn #x2014;Dictionary Learning for Phase Retrie val. IEEE T ransactions on Signal Pr ocessing 64 , 6485–6500 (2016). 00000. 59. Y ang, H., Ercius, P ., Nellist, P . D. & Ophusa, C. Enhanced phase contrast transfer using ptychograph y combined with a pre-specimen phase plate in a scanning transmission electron microscope. Ultramicr oscopy 00001. 60. Danielyan, A., Katkovnik, V . & Egiazarian, K. BM3D Frames and V ariational Image Deblurring. IEEE T ransactions on Image Pr ocessing 21 , 1715–1728 (2012). 00195. 61. Chang, H. & Marchesini, S. A general framework for denoising phaseless dif fraction measurements (2016). URL https://128.84.21.199/abs/1611.01417 . 00001. 62. Chen, Y . & Candes, E. J. Solving Random Quadratic Systems of Equations Is Nearly as Easy as Solving Linear Systems. arXiv:1505.05114 [cs, math, stat] (2015). URL . 00087 arXiv: 1505.05114, 1505.05114 . 63. Shewchuk, J. R. An Introduction to the Conjugate Gradient Method W ithout the Agonizing Pain. T ech. Rep., Carnegie Mellon Uni versity , Pittsb urgh, P A, USA (1994). 01940. 64. T orch.optim. URL https://github.com/torch/optim/ . 00000. 65. Mises, R. V . & Pollaczek-Geiringer , H. Praktische V erfahren der Gleichungsaufl ¨ osung . Z. angew . Math. Mech. 9 , 152–164 (1929). 00185. 66. Lehoucq, R., Sorensen, D. & Y ang, C. ARP ACK User s’ Guide: Solution of Lar ge-Scale Eigen value Pr oblems with Implicitly Restarted Arnoldi Methods . SIAM e-books (Society for Industrial and Applied Mathematics (SIAM, 3600 Market Street, Floor 6, Philadelphia, P A 19104), 1998). URL https://books.google.co. in/books?id=45ZbeycoITcC . 02150. 67. Lebrun, M. An Analysis and Implementation of the BM3D Image Denoising Method. Image Pr ocessing On Line 2 , 175–213 (2012). 00065. 68. Collobert, R., Kavukcuoglu, K. & Farabet, C. T orch7: A Matlab-like En vironment for Machine Learning. In BigLearn, NIPS W orkshop (2011). 00445. 69. Z-cutorch - Complex number support for cutorch. URL https://github.com/PhilippPelz/ z- cutorch . 00000. 70. Leonard, N. Hypero - Hyperparameter optimization for torch. URL https://github.com/ Element- Research/hypero . 00000. 71. Lebrun, M. Bm3d - C++ implementation of BM3D denoising. URL https://github.com/gfacciol/ bm3d . 00000. 72. Balan, R., Casazza, P . & Edidin, D. On signal reconstruction without phase. Applied and Computational Harmonic Analysis 20 , 345–356 (2006). 00232. 73. Clark, J. N. et al. Dynamic sample imaging in coherent diffracti ve imaging. Opt. Lett. 36 , 1954–6 (2011). 00016. 74. Diaz, A. et al. Three-dimensional mass density mapping of cellular ultrastructure by ptychographic X-ray nanotomography . Journal of Structur al Biology 192 , 461–469 (2015). 00005. 14/ 14
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment