Statistical power and prediction accuracy in multisite resting-state fMRI connectivity
Connectivity studies using resting-state functional magnetic resonance imaging are increasingly pooling data acquired at multiple sites. While this may allow investigators to speed up recruitment or increase sample size, multisite studies also potentially introduce systematic biases in connectivity measures across sites. In this work, we measure the inter-site effect in connectivity and its impact on our ability to detect individual and group differences. Our study was based on real, as opposed to simulated, multisite fMRI datasets collected in N=345 young, healthy subjects across 8 scanning sites with 3T scanners and heterogeneous scanning protocols, drawn from the 1000 functional connectome project. We first empirically show that typical functional networks were reliably found at the group level in all sites, and that the amplitude of the inter-site effects was small to moderate, with a Cohen’s effect size below 0.5 on average across brain connections. We then implemented a series of Monte-Carlo simulations, based on real data, to evaluate the impact of the multisite effects on detection power in statistical tests comparing two groups (with and without the effect) using a general linear model, as well as on the prediction of group labels with a support-vector machine. As a reference, we also implemented the same simulations with fMRI data collected at a single site using an identical sample size. Simulations revealed that using data from heterogeneous sites only slightly decreased our ability to detect changes compared to a monosite study with the GLM, and had a greater impact on prediction accuracy. Taken together, our results support the feasibility of multisite studies in rs-fMRI provided the sample size is large enough.
💡 Research Summary
This paper investigates how systematic “site effects” inherent to multisite resting‑state functional MRI (rs‑fMRI) studies influence both univariate statistical power and multivariate prediction accuracy. Using real data rather than purely simulated datasets, the authors assembled a cohort of 345 healthy young adults (age 18‑46) scanned at eight different 3 T sites (Germany, United Kingdom, Australia, United States) drawn from the 1000 Functional Connectome Project. After rigorous preprocessing—including slice‑time correction, motion scrubbing (frames with FD > 0.5 mm removed), regression of white‑matter, ventricular, motion parameters, and 6 mm spatial smoothing—functional connectivity matrices were constructed for a 100‑region brain parcellation. Connectivity for each region pair was quantified as the Fisher‑z transformed Pearson correlation of the regional time series.
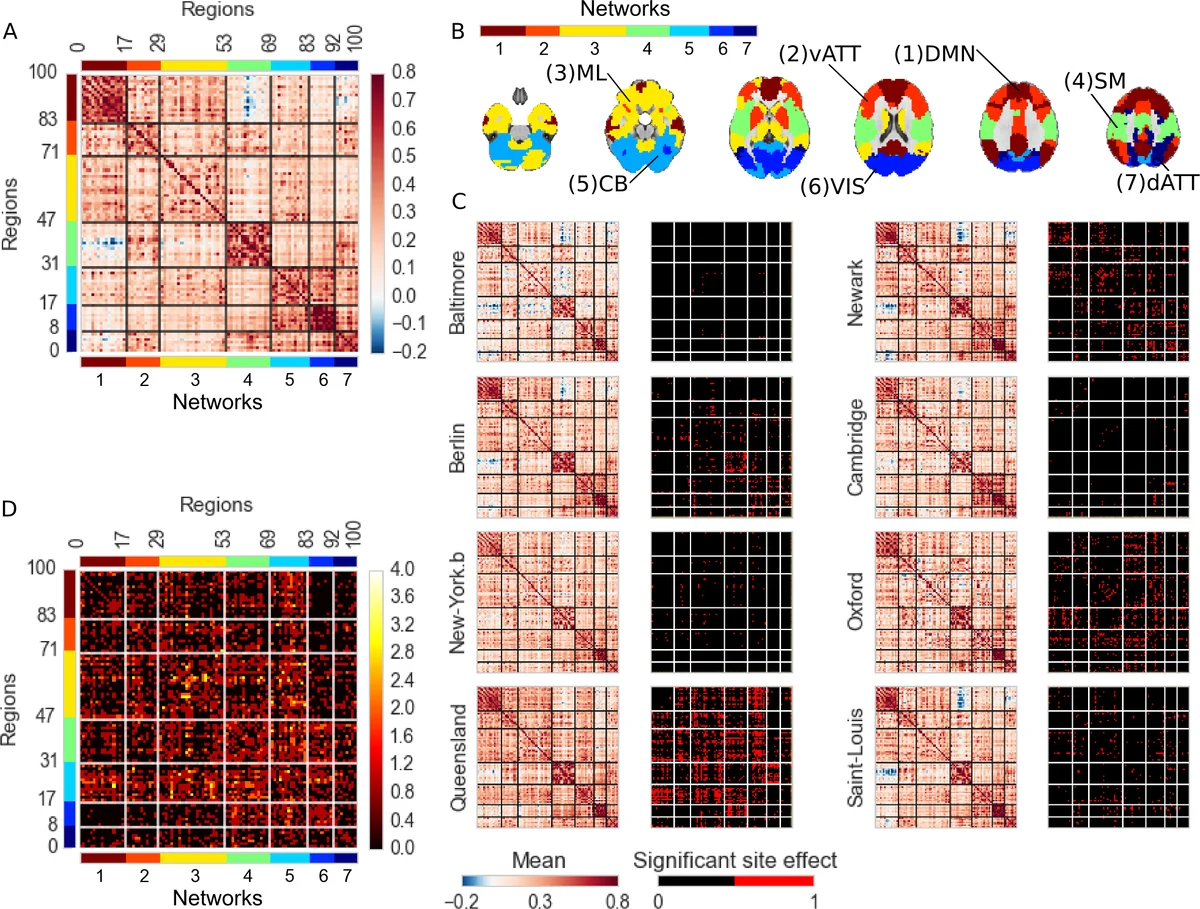
To quantify inter‑site bias, the authors fitted a general linear model (GLM) for every connection, incorporating age, sex, residual motion, and dummy variables for each site. The contrast of each site versus the grand mean yielded β‑coefficients and associated p‑values, corrected for multiple comparisons using a Benjamini‑Hochberg false‑discovery rate (q = 0.05). Cohen’s d effect sizes across all connections averaged 0.34 (maximum ≈ 0.48), indicating small‑to‑moderate systematic differences between sites.
The core of the study involved Monte‑Carlo simulations that added synthetic “pathological” effects to the real data. By varying the effect size (δ = 0.2–0.8), total sample size (N = 40, 80, 120, 200), and the balance of groups across sites, the authors created realistic multisite and monosite scenarios with identical overall N. Two analytical pipelines were evaluated: (1) univariate GLM testing each connection separately, and (2) a linear support‑vector machine (SVM) that ingests the full connectivity matrix to predict group membership. Each simulation was repeated 1,000 times to estimate statistical power (proportion of significant GLM tests) and prediction accuracy (cross‑validated SVM classification rate).
Results showed that multisite designs incurred only a modest loss of GLM power—typically a 5 % reduction relative to a monosite design—and this loss became negligible (≤ 1 %) when the total sample exceeded 100 participants. In contrast, SVM accuracy suffered a larger decrement (6–9 % lower) under multisite conditions, reflecting the greater sensitivity of high‑dimensional classifiers to systematic variance. Nevertheless, when the sample size reached 120 or more, the accuracy gap narrowed to ≤ 2 %. Importantly, a preprocessing step that regressed out site dummy variables before SVM training (“site‑regression”) did not improve classification performance, suggesting that simple linear removal of site effects is insufficient for multivariate models.
The authors discuss several implications. First, for conventional GLM‑based group comparisons, multisite rs‑fMRI studies are viable provided the sample is sufficiently large and covariates (age, sex, motion) are modeled; site effects are relatively small compared with within‑subject physiological variability. Second, multivariate predictive modeling is more vulnerable to site heterogeneity, indicating a need for more sophisticated harmonization techniques (e.g., ComBat, domain adaptation) beyond basic regression. Third, balancing group sizes across sites mitigates power loss, emphasizing careful study design in consortium‑level projects. Limitations include the relatively short scan duration (6–7.5 min), a homogeneous demographic (young, predominantly Caucasian), and reliance on a 100‑region parcellation; future work should explore longer acquisitions, diverse populations, and alternative parcellations or graph‑theoretic features.
In conclusion, this work provides an empirical quantification of inter‑site variability in rs‑fMRI connectivity and demonstrates that, with adequate sample sizes (≈ 120 subjects) and proper statistical control, multisite studies can achieve statistical power and predictive accuracy comparable to single‑site investigations. The findings support the growing trend toward large‑scale, collaborative rs‑fMRI consortia while highlighting the necessity of methodological safeguards for multivariate analyses.
Comments & Academic Discussion
Loading comments...
Leave a Comment