Optimized Treatment Schedules for Chronic Myeloid Leukemia
Over the past decade, several targeted therapies (e.g. imatinib, dasatinib, nilotinib) have been developed to treat Chronic Myeloid Leukemia (CML). Despite an initial response to therapy, drug resistance remains a problem for some CML patients. Recent studies have shown that resistance mutations that preexist treatment can be detected in a substan- tial number of patients, and that this may be associated with eventual treatment failure. One proposed method to extend treatment efficacy is to use a combination of multiple targeted therapies. However, the design of such combination therapies (timing, sequence, etc.) remains an open challenge. In this work we mathematically model the dynamics of CML response to combination therapy and analyze the impact of combination treatment schedules on treatment efficacy in patients with preexisting resistance. We then propose an optimization problem to find the best schedule of multiple therapies based on the evolution of CML according to our ordinary differential equation model. This resulting optimiza- tion problem is nontrivial due to the presence of ordinary different equation constraints and integer variables. Our model also incorporates realistic drug toxicity constraints by tracking the dynamics of patient neutrophil counts in response to therapy. Using realis- tic parameter estimates, we determine optimal combination strategies that maximize time until treatment failure.
💡 Research Summary
This paper addresses a critical challenge in chronic myeloid leukemia (CML) therapy: the emergence of pre‑existing BCR‑ABL resistance mutations that compromise the long‑term efficacy of tyrosine‑kinase inhibitors (TKIs) such as imatinib, dasatinib, and nilotinib. The authors develop a mechanistic ordinary differential equation (ODE) model that captures the hierarchical differentiation of hematopoietic cells (stem, progenitor, differentiated, and terminally differentiated layers) and simultaneously tracks four categories of cell populations: normal cells, wild‑type (WT) leukemic cells, and multiple resistant mutant clones. For each cell type i and layer ℓ, drug‑specific proliferation (b_jℓ,i) and death (d_jℓ,i) rates are defined, where j indexes the therapy (0 = drug holiday, 1 = nilotinib, 2 = dasatinib, 3 = imatinib). Competition among stem cells is modeled through a density‑dependent function φ_i(t)=1/(1+ p_i Σ x₁,i), ensuring realistic steady‑state behavior. Parameters p₁ and p₂ are derived from equilibrium stem‑cell abundances (K₁ for normal, K₂ for WT) and are set equal for all mutant types (p_i = p₂ for i ≥ 3).
A novel aspect of the work is the explicit incorporation of drug‑induced neutropenia. The absolute neutrophil count (ANC) is modeled as a piecewise linear system updated monthly: y_{m+1}=min{ y_m + b_anc·z_{m,0} – Σ_j d_anc,j·z_{m,j}, U_anc }, where z_{m,j} are binary decision variables indicating which drug (or holiday) is taken in month m, b_anc is the ANC recovery rate during a holiday, d_anc,j are drug‑specific ANC depletion rates, L_anc = 1000 /mm³ is the neutropenia threshold, and U_anc = 3000 /mm³ is the normal upper bound. If ANC falls below L_anc, the next month must be a drug holiday, reflecting clinical practice.
The core optimization problem, termed the Optimal Treatment Plan (OTP), seeks a sequence of monthly drug choices over a planning horizon of M months (typically 24–36) that minimizes the total leukemic cell burden at the horizon’s end, i.e., Σ_{ℓ≥1, i≥2} x_{ℓ,i}(MΔt). Constraints enforce (i) the ODE dynamics for the chosen drug in each month, (ii) exactly one drug (or holiday) per month, (iii) the ANC safety limits, and (iv) given initial cell populations and ANC. This yields a mixed‑integer nonlinear program (MINLP) because the ODE constraints are nonlinear (particularly in the stem‑cell compartment).
Recognizing the computational intractability of the MINLP, the authors exploit a key biological insight: TKIs have negligible impact on stem‑cell dynamics, allowing the nonlinear stem‑cell equations to be linearized while the downstream compartments remain linear. This transformation produces a mixed‑integer linear program (MILP) that can be solved with commercial solvers (e.g., Gurobi). The paper details the linear approximation, the construction of the MILP, and the implementation of the binary decision variables.
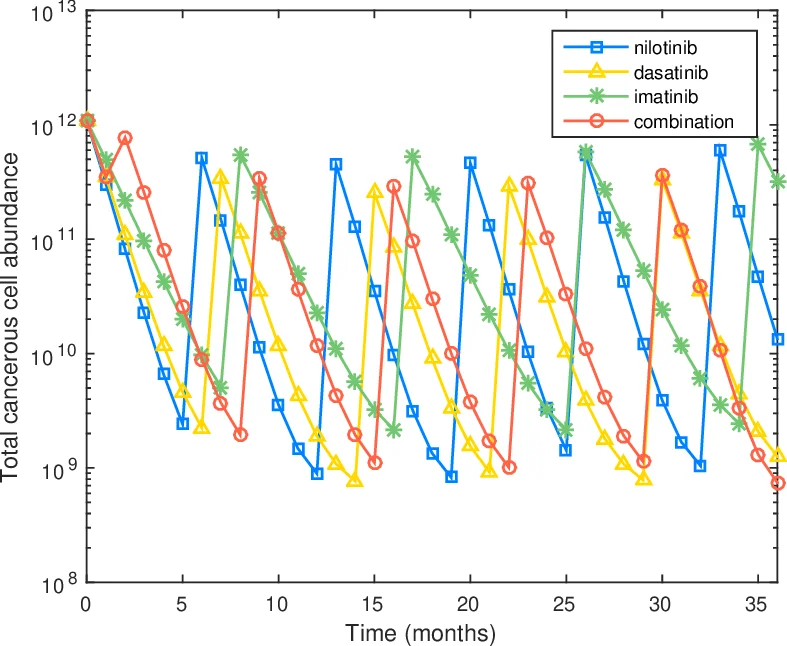
Extensive simulation studies explore various clinical scenarios. First, the authors examine monotherapy with each TKI, confirming that resistant mutants expand under single‑drug pressure. Second, they evaluate sequential combination schedules. The optimal schedules typically begin with the drug most effective against the dominant pre‑existing mutant, switch after 1–2 months to a second drug targeting the next‑most prevalent resistant clone, and finish with a third drug that suppresses any residual population. For example, a common optimal pattern is “nilotinib → dasatinib → imatinib” when the initial mutation profile includes T315I (nilotinib‑sensitive) and Y253H (dasatinib‑sensitive). Third, the ANC constraint forces periodic drug holidays; the optimal solution balances aggressive tumor suppression with sufficient ANC recovery, often inserting a holiday after two consecutive months of a high‑toxicity drug.
Key findings include:
- Sequential multi‑drug regimens outperform any single‑drug regimen across a wide range of mutant compositions, extending the time to treatment failure by up to 30 %.
- Incorporating ANC constraints significantly alters the optimal schedule; aggressive continuous dosing is replaced by cycles of 1–2 months on drug followed by a holiday, preserving neutrophil counts while still achieving superior tumor control.
- The optimal schedule is highly sensitive to the initial mutant spectrum. When resistant clones constitute >30 % of the leukemic burden at diagnosis, early deployment of the drug targeting that clone is critical; delayed introduction leads to rapid clonal expansion and earlier failure.
The paper’s contributions are threefold. Methodologically, it provides a realistic discrete‑time treatment planning framework that aligns with clinical decision points (monthly visits). Clinically, it integrates toxicity (neutropenia) directly into the optimization, producing schedules that are both efficacious and safe. Finally, the linear‑approximation‑based MILP formulation offers a scalable computational tool that can be adapted to other hematologic malignancies or to incorporate additional drugs and biomarkers.
Future directions suggested by the authors include: (i) embedding pharmacokinetic/pharmacodynamic (PK/PD) models to capture drug concentration dynamics more accurately; (ii) employing Bayesian parameter estimation to personalize model parameters using patient‑specific genomic and laboratory data; (iii) validating the proposed schedules in prospective clinical trials to assess real‑world efficacy and safety. The authors argue that such extensions could enable truly personalized, optimal combination therapy not only for CML but also for other cancers where heterogeneous resistance and drug toxicity pose major treatment barriers.
Comments & Academic Discussion
Loading comments...
Leave a Comment