Non-Negative Matrix Factorizations for Multiplex Network Analysis
Networks have been a general tool for representing, analyzing, and modeling relational data arising in several domains. One of the most important aspect of network analysis is community detection or network clustering. Until recently, the major focus…
Authors: Vladimir Gligorijevic, Yannis Panagakis, Stefanos Zafeiriou
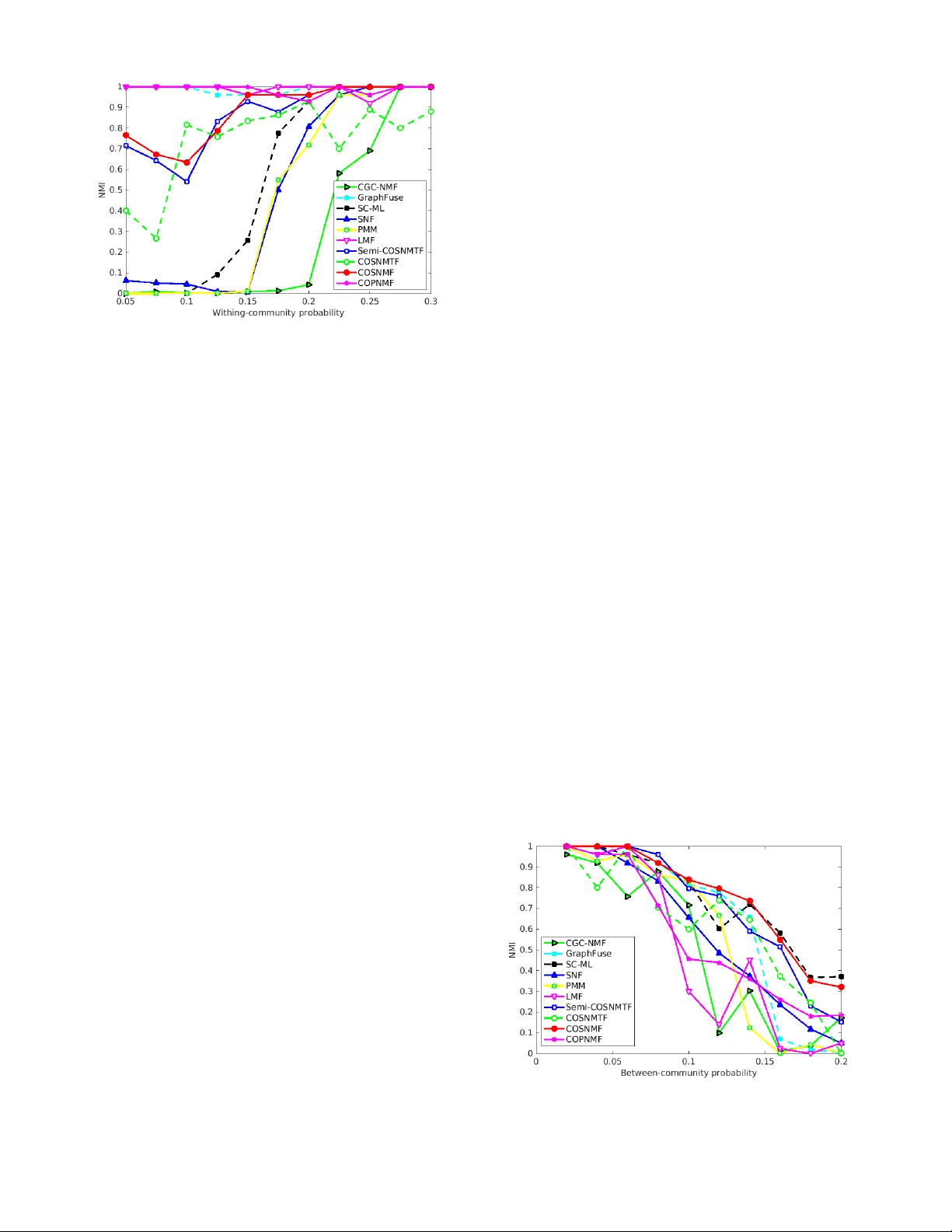
JOURNAL OF L A T E X CLASS FILES, V OL. 14, NO . 8, AUGUST 2015 1 Non-Negativ e Matrix F actor izations f or Multiple x Network Analysis Vladimir Gligorijevi ´ c, Y annis P anagakis, Member , IEEE and Stef anos Zafeiriou, Member , IEEE Abstract —Networks hav e been a general tool for representing, analyzing, and modeling relational data arising in se veral domains . One of the most important aspect of network analysis is community detection or netw ork cluster ing. Until recently , the major f ocus hav e been on discov ering community str ucture in single (i.e., monople x) networks. How ev er , with the advent of relational data with multiple modalities, multiple x networks, i.e., netw orks composed of multiple layers representing diff erent aspects of relations, have emerged. Consequently , community detection in multiplex netw or k, i.e., detecting clusters of nodes shared b y all lay ers, has become a new challenge. In this paper , we propose N etwork F usion for C omposite C omm unity E xtraction ( NF-CCE ), a new class of algorithms, based on four diff erent non–negative matrix factorization models, capable of e xtracting composite communities in multiplex netw orks. Each algorithm works in two steps: first, it finds a non–negative, lo w–dimensional feature representation of each netw or k la yer; then, it fuses the feature representation of la yers into a common non–negative, lo w–dimensional feature representation via collectiv e factorization. The composite clusters are extr acted from the common feature representation. W e demonstrate the superior performance of our algorithms over the state–of–the–art methods on various types of multiple x networks, including biological, social, economic, citation, phone communication, and brain multiple x networ ks. Index T erms —Multiplex networks, non-negativ e matrix factorization, community detection, network integration F 1 I N T R O D U C T I O N N E T W O R K S (or graphs 1 ) along with their theoretical foundations ar e powerful mathematical tools for rep- resenting, modeling, and analyzing complex systems arising in several scientific disciplines including sociology , biol- ogy , physics, and engineering among others [1]. Concretely , social networks, economic networks, biological networks, telecommunications networks, etc. are just a few examples of graphs in which a large set of entities (or agents) corre- spond to nodes (or vertices ) and relationships or interactions between entities correspond to edges (or links ). Structural analysis of these networks have yielded important findings in the corresponding fields [2], [3]. Community detection (also known as graph clustering or module detection ) is one of the foremost problems in network analysis. It aims to find groups of nodes (i.e., clusters, modules or communities ) that are more densely connected to each other than they are to the rest of the network [4]. Even thought the volume of resear ch on community detection is large, e.g., [5], [6], [7], [8], [9], [10], [11], [12], [13], the majority of these methods focus on networks with only one type of relations between nodes (i.e., networks of single type interaction). However , many real-world systems are naturally repre- sented with multiple types of r elationships, or with relation- ships that chance in time. Such systems include subsystems or layers of connectivity representing differ ent modes of complexity . For instance, in social systems, users in social networks engage in differ ent types of interactions (e.g., • V . Gligorijevi ´ c, Y . Panagakis and S. Zafeiriou are with the Department of Computing, Imperial College London, UK. Corresponding author: V . Gligorijevi ´ c, email: v .gligorijevic@imperial.ac.uk 1. we use terms graphs and network interchangeably throughout this paper personal, professional, social, etc.). In biology , different ex- periments or measurements can provide different types of interactions between genes. Reducing these networks to a single type interactions by disregarding their multiple modalities is often a very crude approximation that fails to capture a rich and holistic complexity of the system. In order to encompass a multimodal nature of these relations, a multiplex network representation has been proposed [14]. Multiplex networks (also known as multidimensional , mul- tiview or multilayer networks) have recently attracted a lot of attention in network science community . They can be repr esented as a set of graph layers that share a common set of nodes, but differ ent set of edges in each layer (cf. Fig. 1). W ith the emergence of this network representation, finding composite communities across different layers of multiplex network has become a new challenge [14], [15]. Here, distinct from the previous approaches (reviewed in Section 2), we focus on multiplex community detection . Concretely , a novel and general model, namely the N etwork F usion for C omposite C ommunity E xtraction (NF-CCE) along with its algorithmic framework is developed in Sec- tion 3. The heart of the NF-CCE is the Collective Non- negative Matrix Factorization (CNMF), which is employed in order to collectively factorize adjacency matrices repre- senting dif ferent layers in the network. The collective factor - ization facilitate us to obtain a consensus low-dimensional latent representation, shared across the decomposition, and hence to reveal the communities shared between the net- work layers. The contributions of the paper are as follows: 1) Inspir ed by recent advances in non-negative matrix factorization (NMF) techniques for graph clustering [16], [17] and by using tools for subspace analysis on Grassmann manifold [18], [19], [20], we propose JOURNAL OF L A T E X CLASS FILES, V OL. 14, NO . 8, AUGUST 2015 2 a general framework for extracting composite com- munities from multiplex networks. In particular , the framework NF-CCE, utilizes four dif ferent NMF techniques, each of which is generalized for collec- tive factorization of adjacency matrices r epresenting network layers, and u for computing a consensus low-dimensional latent feature matrix shared acr oss the decomposition that is used for extracting latent communities common to all network layers. T o this end, a general model involving factorization of net- works’ adjacency matrices and fusion of their low- dimensional subspace representation on Grassmann manifold is proposed in Section 3. 2) Unlike a few matrix factorization-based methods for multiplex community extraction that have been pr o- posed so far , e.g., [20], [21], [22], [23], that directly decompose matrices repr esenting network layers into a low-dimensional representation common to all network layers, NF-CCE is conceptually differ- ent. Namely , it works in two steps: first, it denoises each network layer by computing its non-negative low-dimensional repr esentation. Then it merges the low-dimensional representations into a consensus low-dimensional repr esentation common to all net- work layers. This makes our method more robust to noise, and consequently , it yields much stable clustering results. 3) Four efficient algorithms based on four dif ferent NMF techniques are developed for NF-CCE using the concept of natural gradient [19], [20] and pre- sented in the form of multiplicative update rules [24] in Section 3. The advantages of the NF-CCE over the state-of-the-art in community detection are demonstrated by conducting ex- tensive experiments on a wide range of real-world multiplex networks, including biological, social, economic, citation, phone communication, and brain multiplex networks. In particular , we compared the clustering performance of our four methods with 6 state-of-the-art methods and 5 baseline methods (i.e., single-layer methods modified for multiplex networks), on 9 differ ent real-world multiplex networks including 3 large-scale multiplex biological networks of 3 differ ent species. Experimental results, in Section 5, indicate that the proposed methods yield much stable clustering results than the state-of-the-art and baseline methods, by robustly handling incomplete and noisy network layers. The experiments conducted on multiplex biological networks of 3 differ ent species indicate NF-CCE as a superior method for finding composite communities (in biological networks also known as functional modules ). Moreover , the results also indicate that NF-CCE can extract unique and mor e function- ally consistent modules by considering all network layers together than by considering each network layer separately . Notations : throughout the paper , matrices are de- noted by uppercase boldface letters, e.g., X . Subscript in- dices denote matrix elements, e.g., X ij , whereas superscript indices in brackets denote network layer , e.g., X ( i ) . The set of real numbers is denoted by R . | · | denotes the cardinality of a set, e.g., | S | . A binary matrix of size n × m is represented by { 0 , 1 } n × m . G 1 (V,E 1 ) G 2 (V,E 2 ) G 3 (V,E 3 ) c 1 c 1 c 1 c 2 c 2 c 2 Fig. 1. An example of a multiple x networ k with 11 nodes present in three complementary layers denoted in diff erent colors. T wo different communities across all three la yers can be identified. 2 B AC K G R O U N D A N D R E L AT E D W O R K 2.1 Single-la yer (monoplex) netw orks In graph theory , a monoplex network (graph) can be repr e- sented as an ordered pair , G = ( V , E ) , where V is a set of n = | V | vertices or nodes, and E is a set of m = | E | edges or links between the vertices [25]. An adjacency matrix repr esenting a graph G is denoted by A ∈ { 0 , 1 } n × n , where A ij = 1 if there is an edge between vertices i and j , and A ij = 0 otherwise. Most of the real-world networks that we consider throughout the paper are represented as edge- weighted graphs , G = ( V , E , w ) , where w : E → R assigns real values to edges. In this case, the adjacency matrix instead of being a binary matrix, is a r eal one i.e., A ∈ R n × n , with entries characterizing the strength of association or interaction between the network nodes. Although, there is no universally accepted mathematical definition of the community notion in graphs, the probably most commonly accepted definition is the following: a com- munity is a set of nodes in a network that are connected more densely among each other than they are to the rest of the network [14]. Hence, the problem of community detection is as follows: given an adjacency matrix A of one network with n nodes and k communities, find the commu- nity assignment of all nodes, denoted by H ∈ { 0 , 1 } n × k , where H ij = 1 if nodes i belongs to community j , and H ij = 0 otherwise. W e consider the case of non-overlapping communities, where a node can belong to only one commu- nity , i.e., P k j =1 H ij = 1 . T o address the community detection problem in mono- plex networks, several methods have been proposed. Com- prehensive surveys of these methods are [4] and [13]. T o make the paper self-contained, her e, we briefly r eview some of the most repr esentative approaches, including graph par- titioning, spectral clustering, hierarchical clustering, modularity maximization, statistical inference and structure-based methods , as well as method that rely on non-negative matrix factoriza- tions : • Graph partitioning aims to group nodes into partitions such that the cut size , i.e., the total number of edges between any two partitions, is minimal. T wo widely used graph partitioning algorithms that also take into account the size of partitions are Ratio Cut and Normalized Cut [26]. Graph partitioning algorithms can be alternatively defined as spectral algorithms JOURNAL OF L A T E X CLASS FILES, V OL. 14, NO . 8, AUGUST 2015 3 in which the objective is to partition the nodes into communities based on their eigenvectors obtained from eigendecomposition of graph Laplacian matrix [27]. • In hierar chical clustering the goal is to reveal network communities and their hierarchical structure based on a similarity (usually topological) measure com- puted between pairs of nodes [28]. • Modularity-based algorithms ar e among the most pop- ular ones. Modularity was designed to measure the strength of partition of a network into communities. It is defined as a fraction of the edges that fall within the community minus the expected fraction when these edges are randomly distributed [5], [29]. V arious algorithms have been pr oposed for mod- ularity optimization, including gr eedy techniques, simulated annealing, spectral optimization, etc. [4]. • Statistical inference methods aims at fitting the gen- erative model to the network data based on some hypothesis. Most commonly used statistical infer ence method for community detection is the stochastic block model , that aims to approximate a given adja- cency matrix by a block structure [30]. Each block in the model represents a community . • Structur e-based methods aim to find subgraphs repre- senting meta definitions of communities. Their objec- tive is to find maximal cliques, i.e., the cliques which are not the subgraph of any other clique. The union of these cliques form a subgraph, whose components are interpreted as communities [31]. • Mor e recently , graph clustering methods that rely on the Non-Negative Matrix Factorization (NMF) [24] have been proposed e.g., [16], [17]. Their goal is to approximate a symmetric adjacency matrix of a given network by a product of two non-negative, low-rank matrices, such that they have clustering interpretation, i.e., they can be used for assigning nodes to communities. The proposed methods here, follow this line of research, but as opposed to the ex- isting methods [16], [17], the NF-CCE can effectively handle multiplex networks. 2.2 Multiplex netw orks A multiplex network is a set of N monoplex networks (or layers), G i ( V , E i ) , for i = 1 , . . . , N . The number of nodes in each layer is the same, n = | V | , while the connectivity pattern and the distribution of links in each layer differs, m i = | E i | (see Fig. 1). Similarly to monoplex networks, we consider the case where each layer repr esents a weighted, undirected graph, i.e., G i ( V , E i , w i ) . A multiplex network can be represented as a set of adjacency matrices encoding connectivity patterns of individual layers, A ( i ) ∈ R n × n , for i = 1 , . . . , N . The goal of community detection in multiplex networks is to infer shared, latent community assignment that best fits all given layers. Given that each layer contains incomplete and complementary information, this process of finding shared communities by integrating information from all layers is also known in the literature as network integration (fusion) [22], [32]. Unlike the case of monoplex networks, resear ch on com- munity detection in multiplex networks is scarce. Existing methods extract communities from multiplex networks first by aggregating the links of all layers into a single layer , and then applying a monoplex method to that single layer [22], [33], [34]. However , this approach does not account for shared information between layers and treats the noise present in each layer uniformly . Clearly , this is not the case in real-world multiplex networks where each level is contaminated by different noise in terms of magnitude and, possibly , distribution. Thus, by aggregating links from differ ent layers the noise in the aggregated layer signifi- cantly increases, resulting in a poor community detection performance. Current state-of-the-art methods are built on monoplex approaches and further generalized to multiplex networks. They can be divided into the following categories: • Modularity-based approaches that generalize the no- tion of modularity from single-layer to multi-layer networks [35]. Namely , to alleviate the above men- tioned limitations, the Principal Modularity Maxi- mization (PMM) [22] has been proposed. First, for each layer , PMM extracts structural features by opti- mizing its modularity , and thus significantly denois- ing each layer; then, it applies PCA on concatenated matrix of structural feature matrices, to find the principal vectors, followed by K-means to perform clustering assignment. The main drawback of this approach is that it treats structural feature matrices of all layers on equal basis (i.e., it is not capable of distinguishing between more and less informa- tive network layers, or complementary layers). Even though the noise is pr operly handled by this method, the complementarity aspect cannot be captured well by the integration step. • Spectral clustering approaches that generalize the eigendecomposition from single to multiple Lapla- cian matrices repr esenting network layers. One of the state-of-the-art spectral clustering methods for multi- plex graphs is the Spectral Clustering on Multi-Layer (SC-ML) [18]. First, for each network layer , SC-ML computes a subspace spanned by the principal eigen- vectors of its Laplacian matrix. Then, by interpreting each subspace as a point on Grassmann manifold, SC-ML merges subspaces into a consensus subspace from which the composite clusters are extracted. The biggest drawback of this methods is the underlying spectral clustering, that always finds tight and small- scale and, in some cases, almost trivial communities. For example, SC-ML cannot adequately handle net- work layers with missing or weak connections, or layers that have disconnected parts. • Information diffusion-based approaches that utilize the concept of diffusion on networks to integrate net- work layers. One of such methods is Similarity Net- work Fusion (SNF) proposed by W ang et al. [36]. SNF captures both shared and complementary in- formation in network layers. It computes a fused matrix from the similarity matrices derived from all layers through parallel interchanging dif fusion process on network layers. Then, by applying a spectral clustering method on the fused matrix they JOURNAL OF L A T E X CLASS FILES, V OL. 14, NO . 8, AUGUST 2015 4 extract communities. However , for sparse networks, the diffusion process, i.e., information propagation, is not very efficient and it may results in poor clustering performance. • Matrix and tensor factorization-based approaches that utilize collective factorization of adjacency matri- ces repr esenting network layers. A few matrix and tensor decomposition-based approaches have been proposed so far [21], [23], [37], [38]. T ang et al. [37] introduced the Linked Matrix Factorization (LMF) which fuses information from multiple network lay- ers by factorizing each adjacency matrix into a layer- specific factor and a factor that is common to all network layers. Dong et al. [23], introduced the Spec- tral Clustering with Generalized Eigendecomposi- tion (SC-GED) which factorizes Laplacian matrices instead of adjacency matrices. Papalexakis et al. [38] proposed GraphFuse, a method for clustering multi- layer networks based on sparse P ARAllel F ACtor (P ARAF AC) decomposition [39] with non-negativity constraints. Cheng et al. introduced Co-regularized Graph Clustering based on NMF (CGC-NMF). They factorize each adjacency matrix using symmetric NMF while keeping the Euclidean distance between their non-negative low-dimensional representations small. As already pointed out in Section 1, one of the major limitations of all of these factorization methods is that they treat each network layer on an equal basis and, unlike PMM or SC-ML, for example, they cannot filter out irrelevant information or noise. T o alleviate the drawbacks of the aforementioned meth- ods, the NF-CCE framework is detailed in the following section. It consists of 4 models, where each layer is first de- noised by computing its non-negative low-dimensional sub- space representation. Then, the low-dimensional subspaces are merged into a consensus subspace whose non-negative property enables clustering interpretation. The models are conceptually similar to the SC-ML method, since they use the same merging technique to find the common subspace of all layers. 3 P R O P O S E D F R A M E W O R K Here, we present four novel metods that are built upon 4 non-negative matrix factorization models, SNMF [16], PNMF [40], SNMTF [41] and Semi-NMTF [42], and extended for fusion and clustering of multiplex networks. Since the derivation of each method is similar , we present them in a unified framework, namely NF-CCE. NF-CCE extracts com- posite communities from a multiplex network consisting of N layers. In particular , given N -layered multiplex network repr esented by adjacency matrices, { A (1) , . . . , A ( N ) } , NF- CCE consists of two steps: Step 1 : For each network layer , i , we obtain its non-negative, low-dimensional repr esentation, H ( i ) , under column orthonormality constraints i.e., H ( i ) T H ( i ) = I , by using any of the non-negative factorization methods men- tioned above. Step 2 : W e fuse the low-dimensional representations into a common, consensus representation, H , by proposing a collective matrix factorization model. That is, we collectively decompose all adjacency matrices, A ( i ) into a common ma- trix, H , whilst enforcing the non-negative low-dimensional repr esentation of network layers, H ( i ) (computed in the previous step), to be close enough to the consensus low- dimensional repr esentation, H . The general objective func- tion capturing these two properties is written as follows: min H ≥ 0 J = N X i =1 J ( i ) ( H ; A ( i ) ) + α N X i =1 J ( i ) c ( H ; H ( i ) ) (1) where, J ( i ) is an objective function for clustering ith layer and J ( i ) c is the loss function quantifying the inconsistency between each low-dimensional representation H ( i ) , com- puted in Step 1 , and the consensus repr esentation H . Below we provide the details of the second step for each individual factorization technique. 3.1 Collective SNMF (CSNMF) W e factorize each individual adjacency matrix using Sym- metric NMF in the following way: A ( i ) ≈ HH T under the following constraints: H ≥ 0 and H T H = I ; where, i = 1 , . . . , N . The first part of our general objective function in Step 2 (Eq. 1) has the following form: J ( i ) ( H ; A ( i ) ) = k A ( i ) − HH T k 2 F (2) where H is the consensus, non-negative low-dimensional matrix, and F denotes Frobenius norm. 3.2 Collective PNMF (CPNMF) W e factorize each individual adjacency matrix using Projec- tive NMF in the following way: A ( i ) ≈ HH T A ( i ) under the following constraints: H ≥ 0 and H T H = I ; where, i = 1 , . . . , N . The first part of our general objective function in Step 2 (Eq. 1) has the following form: J ( i ) ( H ; A ( i ) ) = k A ( i ) − HH T A ( i ) k 2 F (3) where H is the consensus, non-negative low-dimensional matrix. 3.3 Collective SNMTF (CSNMTF) W e tri-factorize each individual adjacency matrix using Symmetric NMTF in the following way: A ( i ) ≈ HS ( i ) H T under the following constraints: H ≥ 0 and H T H = I ; where i = 1 , . . . , N . The first part of our general objective function in Step 2 (Eq. 1) has the following form: J ( i ) ( H ; A ( i ) , S ( i ) ) = k A ( i ) − HS ( i ) H T k 2 F (4) JOURNAL OF L A T E X CLASS FILES, V OL. 14, NO . 8, AUGUST 2015 5 where H is the consensus low-dimensional matrix. In the derivation of our algorithm, we distinguish be- tween two cases. In the first case, we consider S matrix to be non-negative, i.e., S ( i ) ≥ 0 . W e call that case CSNMTF . In the second case, we consider S ( i ) matrix to have both positive and negative entries. W e call this case collective symmetric semi-NMTF , or CSsemi-NMTF (or CSsNMTF). 3.4 Merging low-dimensional representation of graph lay ers on Grassmann Manifolds For the second term in our general objective function (Eq. 1) in Step 2 , we utilize the orthonormal property of non- negative, low-dimensional matrices, H ( i ) , and propose a distance measure based on this property . Namely , Dong et al. [18] proposed to use the tools from subspace analysis on Grassmann manifold. A Grassmann manifold G ( k , n ) is a set of k -dimensional linear subspaces in R n [18]. Given that, each orthonormal cluster indicator matrix, H i ∈ R n × k , spanning the corr esponding k -dimensional non-negative subspace, span ( H i ) in R n , is mapped to a unique point on the Grassmann manifold G ( k , n ) . The geodesic distance between two subspaces, can be computed by projection distance. For example, the square distance between two subspaces, H i and H j , can be computed as follows: d 2 proj ( H i , H j ) = k X i =1 sin 2 θ i = k − k X i =1 cos 2 θ i = k − tr ( H i H T i H j H T j ) T o find a consensus subspace, H , we factorize all the ad- jacency matrices, A ( i ) , and minimize the distance between their subspaces and the consensus subspace on Grassmann manifold. Following this approach, we can write the second part of our general objective function in the following way: J ( i ) c ( H , H ( i ) ) = k − tr ( HH T H ( i ) H ( i ) T ) (5) = k HH T − H ( i ) H ( i ) T k 2 F 3.5 Deriv ation of the general multiplicative update rule In Step 1 , we use well-known non-negative factorization techniques, namely SNMF , PNMF , SNMTF and Ssemi- NMTF , for which the update rules for computing low- dimensional non-negative matrices, H ( i ) , have been pro- vided in the corresponding papers [16], [40], [41], [42], respectively . They are summarized in T able 1. As for the Step 2 , we derive the update rules for each of the collective factorization techniques presented in Sections 3.1, 3.2 and 3.3. The details of the derivation are given in Section 2 of the online supplementary material. Here we provide a general update rule for Equation 1. W e minimize the general objective function shown in Equation 1, under the following constraints: H ≥ 0 and H T H = I . Namely , we adopt the idea from Ding et al. [41] to impose orthonormality constraint on H matrix, i.e., H T H = I ; that has been shown to lead to a more rigorous clustering interpretation [41]. Moreover , assignments of net- work nodes to composite communities can readily be done by examining the entries in rows of H matrix. Namely , we can interpret matrix H n × k as the cluster indicator matrix , where the entries in i -th row (after row normalization) can be interpreted as a posterior probability that a node i belongs to each of the k composite communities. In all our experiments, we apply hard clustering procedur e, where a node is assign to the cluster that has the largest probability value. W e derive the update rule, for matrix H , for minimizing the objective function (Eq. 1) following the procedure from the constrained optimization theory [43]. Specifically , we follow the strategy employed in the derivation of NMF [24] to obtain a multiplicative update rule for H matrix that can be used for finding a local minimum of the optimization problem (Eq. 1). The derivative of the objective function (Eq. 1) with respect to H is as follows: ∇ H J = N X i =1 ∇ H J ( i ) ( H ; A ( i ) ) − α N X i =1 H ( i ) H ( i ) T H (6) where the first term under summation can be decomposed into two non-negative terms, namely: ∇ H J ( i ) ( H ; A ( i ) ) = [ ∇ H J ( i ) ( H ; A ( i ) )] + − [ ∇ H J ( i ) ( H ; A ( i ) )] − where, [ ∇ H J ( i ) ( H ; A ( i ) )] + ≥ 0 , [ ∇ H J ( i ) ( H ; A ( i ) )] − ≥ 0 are non-negative terms. Depending on the type of collective factorization technique repr esented in Section 3.1, 3.2 or 3.3, the first term repr esents the derivative of the corresponding objective function, i.e., Equation 2, 3 or 4, respectively . The second term repr esents a derivative of Equation 5 with respect to H . T o incorporate the orthonormality constraint into the update rule, we introduce the concept of natural gradient by following the work of Panagakis et al. [20]. Namely , we shown in Section 3.4 that columns of H matrix span a vector subspace known as Grassmann manifold G ( k , n ) , i.e., span ( H ) ∈ G ( k, n ) [20]. Using that, Amari in [19] has showed that when an optimization problem is defined over a Grassmann manifold, the ordinary gradient of the optimization function (Equation 6) does not repr esent its steepest direction, but natural gradient does [19]. Therefor e, we define a natural gradient to optimize our objective function (1) under the orthornormality constraint. Following Panagakis et al. [20], the natural gradient of J on Grassmann manifold at H can be written in terms of the ordinary gradient as follows: e ∇ H J = ∇ H J − HH T ∇ H J (7) where, ∇ H J is the ordinary gradient given in Equation 6. Following the Karush-Kuhn-T ucker (KKT) complemen- tarity condition [43] and preserving the non-negativity of H , the general update rule for H matrix using the natural gradient is as follows: H j k ← H j k ◦ [ e ∇ H J ] − j k [ e ∇ H J ] + j k (8) where, “ ◦ ” denotes Hadamard product. JOURNAL OF L A T E X CLASS FILES, V OL. 14, NO . 8, AUGUST 2015 6 [ e ∇ H J ] − = HH T N X i =1 [ ∇ H J ( i ) ( H ; A ( i ) )] + + N X i =1 [ ∇ H J ( i ) ( H ; A ( i ) )] − + α N X i =1 H ( i ) H ( i ) T H [ e ∇ H J ] + = HH T N X i =1 [ ∇ H J ( i ) ( H ; A ( i ) )] − + N X i =1 [ ∇ H J ( i ) ( H ; A ( i ) )] + + α HH T N X i =1 H ( i ) H ( i ) T H The concrete update rule for each collective factorization method is summarized in T able 1 and united within NF-CCE algorithm (see Algorithm 1). Algorithm 1 NF-CCE Input : Adjacency matrices A ( i ) for each network layer i = 1 , . . . , N ; number of clusters k ; parameter α ; factorization technique: F ACTORIZA TION ∈ { SNMF , PNMF , SNMTF } Output : Consensus cluster indicator matrix H switch (F ACTORIZA TION) case ’SNMF’ : for i ∈ [1 , N ] do H ( i ) ← F ACTORIZA TION ( A ( i ) , k ) end for A avg ← N P i =1 A ( i ) + α 2 H ( i ) H ( i ) T H ← F ACTORIZA TION ( A avg , k ) case ’PNMF’ : for i ∈ [1 , N ] do H ( i ) ← F ACTORIZA TION ( A ( i ) , k ) end for A avg ← N P i =1 A ( i ) A ( i ) T + α H ( i ) H ( i ) T H ← F ACTORIZA TION ( A avg , k ) case ’SNMTF’ : for i ∈ [1 , N ] do ( H ( i ) , S ( i ) ) ← F ACTORIZA TION ( A ( i ) , k ) end for H ← CSNMTF ( { A ( i ) } N i =1 , { H ( i ) } N i =1 , { S ( i ) } N i =1 , k ) end switch 4 E X P E R I M E N T S W e test our methods on synthetic, as well as on real-world data. W e designed synthetic multiplex networks with clear ground truth information and different properties in terms of noise and complementary information of network layers. The goal is to address the robustness of our methods against noise and their ability to handle complementary information contained in layers. The real-world multiplex networks are taken from diverse experimental studies to demonstrate the applicability of our methods in a broad spectrum of disci- plines. Namely , we consider social and biological networks, networks of mobile phone communications, brain networks and networks constructed from bibliographic data. The bi- ological networks are treated as a special case because of their lack of ground truth clusters. W e pr ovide detailed anal- ysis of such networks based on the functional annotations of their nodes (genes). W e present results of comparative analysis of our proposed methods against state-of-the-art methods described in Section 2.2. Specifically , we compare our methods against PMM, SC-ML, SNF , LMF , GraphFuse and CGC-NMF . Moreover , we adopt the following single- layer methods, SNMF , SNMTF , PNMF and MM (modularity maximization) to be our baseline methods. 4.1 Synthetic m ultiplex networks W e generate two sets of synthetic multiplex networks. First type, that we denote SYNTH-C , is designed to demonstrate complementary information in layers; whereas the second type, that we denote SYNTH-N , is designed to demonstrate differ ent levels of noise between communities contained in layers. Our synthetic networks are generated by using planted partition model [44]. The procedure is as follows: we choose the total number of nodes n partitioned into N communities of equal or different sizes. For each layer , we split the corresponding adjacency matrix into blocks defined by the partition. Entries in each diagonal block, are filled with ones randomly , with probability p ii , repr esenting the within-community pr obability or also referr ed as com- munity edge density . W e also add random noise between each pair of blocks, ij , with probability p ij , repr esenting between-community probability . The larger the values of p ij are the harder the clustering is. Similarly , the smaller the values of p ii are the harder the clustering is. W e vary these probabilities across the layers to simulate complementary information and noise in the following way: SYNTH-C . W e generate two-layer multiplex networks with n = 200 nodes and N = 2 communities with equal number of nodes each. W e generate 11 differ ent multiplex networks with differ ent amounts of information between two layers. Namely , we vary the within-community prob- ability p 11 = { 0 . 05 , 0 . 075 , . . . , 0 . 3 } of the first community of the first layer across different multiplex networks, while fixing the within-community probability of the second com- munity , p 22 = 0 . 2 . In the second layer , we represent the com- plementary information by fixing the within-community probability of the first community to p 11 = 0 . 2 and varying within-cluster probability of the second community p 22 = { 0 . 05 , 0 . 075 , . . . , 0 . 3 } across the multiplex networks. For all multiplex networks, we set the same amount of nosy links, by fixing between-community probability to p 12 = 0 . 05 . SYNTH-N . Similar to SYNTH-C we generate two-layer multiplex networks with two communities ( n and N are the same as in SYNTH-C ). W e fix the within-community prob- ability of both communities and both layers to p 11 = 0 . 3 and p 22 = 0 . 3 across all multiplex networks. W e vary the between-community probability p 12 = { 0 . 02 , 0 . 04 , . . . , 0 . 2 } of the first layer , while keeping the between-community probability of the second layer fixed, p 12 = 0 . 02 , across all multiplex networks. The spy plots of adjacency matrices repr esenting layers of SYNTH-C and SYNTH-N are given in the Section 1 of the online supplementary material. 4.2 Real-w orld multiplex netw orks Below we provide a brief description of real-world multi- plex networks used in our comparative study: Bibliographic data, CiteSeer : the data ar e adopted fr om [45]. The network consist of N = 3 , 312 papers belonging to JOURNAL OF L A T E X CLASS FILES, V OL. 14, NO . 8, AUGUST 2015 7 T ABLE 1 Multiplicative update rules (MUR) f or single-lay er and multiplex netw or k analysis. Method Single-layer MUR Multiplex MUR SNMF H ( i ) j k ← H ( i ) j k ◦ A ( i ) H ( i ) j k H ( i ) H ( i ) T A ( i ) H ( i ) j k H j k ← H j k ◦ N P i =1 A ( i ) + α 2 H ( i ) H ( i ) T H j k HH T N P i =1 A ( i ) + α 2 H ( i ) H ( i ) T H j k PNMF H ( i ) j k ← H ( i ) j k ◦ A ( i ) A ( i ) T H ( i ) j k H ( i ) H ( i ) T A ( i ) A ( i ) T H ( i ) j k H j k ← H j k ◦ N P i =1 A ( i ) A ( i ) T + α H ( i ) H ( i ) T H j k HH T N P i =1 A ( i ) A ( i ) T + α H ( i ) H ( i ) T H j k SNMTF H ( i ) j k ← H ( i ) j k ◦ A ( i ) H ( i ) S ( i ) j k H ( i ) H ( i ) T A ( i ) H ( i ) S ( i ) j k H j k ← H j k ◦ N P i =1 A ( i ) HS ( i ) + α 2 N P i =1 H ( i ) H ( i ) T H j k HH T N P i =1 A ( i ) HS ( i ) + α 2 N P i =1 H ( i ) H ( i ) T H j k S ( i ) j k ← S ( i ) j k ◦ H ( i ) T A ( i ) H ( i ) j k H ( i ) T H ( i ) S ( i ) H ( i ) T H ( i ) j k SsNMTF H ( i ) j k ← H ( i ) j k ◦ h A ( i ) H ( i ) S ( i ) + + H ( i ) H ( i ) T [ A ( i ) H ( i ) S ( i ) − i j k h A ( i ) H ( i ) S ( i ) − + H ( i ) H ( i ) T [ A ( i ) H ( i ) S ( i ) + i j k H j k ← H j k h N P i =1 A ( i ) HS ( i ) + + HH T A ( i ) HS ( i ) − + α 2 H ( i ) H ( i ) T H i j k h N P i =1 A ( i ) HS ( i ) − + HH T A ( i ) HS ( i ) + + α 2 H ( i ) H ( i ) T H i j k S ( i ) ← H ( i ) T H ( i ) − 1 H ( i ) T A ( i ) H ( i ) ( H ( i ) T H ( i ) − 1 6 different resear ch categories, that we grouped into k = 3 pairs categories. W e consider these categories as the ground truth classes. W e construct two layers: citation layer , repre- senting the citation relations between papers extracted from the paper citation recor ds; and the paper similarity layer , constructed by extracting a vector of 3 , 703 most frequent and unique words for each paper , and then computing the cosine similarity between each pair of papers. W e construct the k-nearest neighbor graph from the similarity matrix by connecting each paper with its 10 most similar papers. Bibliographic data, CoRA : the data are adopted from [45]. The network consists of 1 , 662 machine learning papers grouped into k = 3 differ ent resear ch categories. Namely , Genetic Algorithms, Neural Networks and Probabilistic Methods. W e use the same approach as for CiteSeer dataset to construct the citation and similarity layers. Mobile phone data (MPD) : the data are adopted from [23]. The network consists of N = 3 layers repr esenting differ ent mobile phone communications between n = 87 phone users on the MIT campus; namely , the layers represent physical location, bluetooth scans and phone calls. The ground truth clusters are known and manually annotated. Social network data (SND) : the data are adopted fr om [46]. The dataset repr esents the multiplex social network of a corporate law partnership, consisting of N = 3 layers having three types of edges, namely , co-work, friendship and advice. Each layer has n = 71 nodes repr esenting employees in a law firm. Nodes have many attributes. W e use the location of employees’ offices as well as their status in the firm as the ground truth for clusters and perform two differ ent experiments, namely SND(o) and SND(s) respec- tively . Worm Brain Networks (WBN) : the data are retrieved from W ormAtlas 2 , i.e., from the original study of White et al. [47]. The network consist of n = 279 nodes repr esenting neur ons, connected via N = 5 different types of links (i.e., layers), repr esenting 5 different types of synapse. W e use neuron types as the ground truth clusters. Word T rade Networks (WTN) : the data r epresents differ ent types of trade relationships (import/export) among n = 183 2. http://www .wormatlas.org/ countries in the world [48]. The network consist of 339 layers r epresenting different products (goods). Since, for some products layers are very sparse, we retain the layers having more than n − 1 links, which resulted in N = layers. W e use geographic r egions (continents) of countries and eco- nomic trade categories for defining ground truth clusters 3 . Thus, we perform experiments with this two ground truth clusters, namely WTN (reg) , denoting geographic regions and WTN (cat) , denoting economic categories. In T able 2 we summarize the important statistics and information of real-world multiplex networks used in our experiments. 4.2.1 Multiple x biological networks W e obtained multiplex biological networks of 3 differ ent species, i.e., human biological network (HBN), yeast biolog- ical network (YBN) and mouse biological network (MBN) 4 , from the study of Mostafavi and Morris [49]. The network layers are constructed from the data obtained from differ - ent experimental studies and from the publicly available databases. The network layers represent differ ent types of interactions between genes 5 , including protein interactions, genetic interaction, gene co-expressions, protein localiza- tion, disease associations, etc. The number of nodes and layers in each network is summarized in T able 2. For each network and its genes, the corresponding GO annotations has also been provided by Mostafavi and Morris [49]. For details about network statistics and layer repr esentation we refer a reader to Mostafavi and Morris [49]. 4.3 Setup f or state-of-the-ar t methods Each of the state-of-the-art method takes as an input param- eter the number of clusters k that needs to be known in advance. Also, some of the methods take as input other types of parameters that needs to be determined. T o make 3. data about countries are downloaded from http://unctadstat. unctad.org 4. The network can be retrieved from: http://morrislab.med. utoronto.ca/ ∼ sara/SW/ 5. genes and their coded proteins are considered as the same type of nodes in networks layers JOURNAL OF L A T E X CLASS FILES, V OL. 14, NO . 8, AUGUST 2015 8 T ABLE 2 Real-world multiple x networks used for our comparativ e study . Net name n N ground truth Ref. CiteSeer 3,312 2 known ( k = 3 ) [45] CoRA 1,662 2 known ( k = 3 ) [45] MPD 87 3 known ( k = 6 ) [23] SND 71 3 known ( k = 3 ) [46] WBN 279 10 known ( k = 10 ) [47] WTN 183 14 known ( k = 5 ) [48] HBN 13,251 8 unknown ( k = 100 ) [49] YBN 3,904 44 unknown ( k = 100 ) [49] MBN 21,603 10 unknown ( k = 100 ) [49] the comparison fair , below we briefly explain each of the comparing method and provide the implementation and parameter fitting details that we use in all our experiments (for detailed procedure on parameter fitting, please refer to Section 3 in the online supplementary material): Baseline, single-layer methods (MM, SNMF , PNMF , SN- MTF and SsNMTF) . In order to apply them on multiplex network we first merge all the network layers into a sin- gle network described by the following adjacency matrix: A = 1 N P N i =1 A ( i ) . PMM [22] has a single parameter , ` , which represents the number of structural features to be extracted from each network layer . In all our runs, we compare the clustering performance by varying this parameter , but we also noted that the clustering performance does not change signifi- cantly when l k . SNF [36] the method is parameter-free. However , the method prefers data in the kernel matrix. Thus, we use diffusion kernel matrix representation of binary interaction networks as an input to this method. SC-ML [18] has a single regularization parameter , α , that balances the trade-off between two terms in the SC- ML objective function. In all our experiments we choose the value of α that leads to the best clustering performance. LMF [37] has a regularization parameter , α , that balances the influence of r egularization term added to objective func- tion to improve numerical stability and avoid over fitting. W e vary α in all our runs, and choose the value of α that leads to the best clustering performance. GraphFuse [38] has a single parameter , sparsity penalty factor λ , that is chosen by exhaustive grid search and the value of λ that leads to the best clustering performance is chosen. CGC-NMF [21] has a set of parameters γ ij ≥ 0 that bal- ance between single-domain and cross-domain clustering objective for each pair of layers ij . Given that in all our experiments the relationship between node labels for any pair of layers is one-to-one , we set γ ij = 1 (as in [21]) for all pairs of layers and throughout all our experiments. 4.4 Clustering ev aluation measures Here we discuss the evaluation measures used in our ex- periments to evaluate and compare the performance of our proposed methods with the above described state-of-the-art methods. Given that we test our methods on multiplex network with known and unknown ground truth cluster , we distinguish between two sets of measures: Known ground truth . For multiplex network with known ground truth clustering assignment, we use the following three widely used clustering accuracy measures: Purity [50], Normalized Mutual Information (NMI) [51] and Adjusted Rand Index (ARI) [51]. All three measures provide a quantitative way to compare the computed clusters Ω = { ω 1 , . . . , ω k } with respect to the ground truth classes: C = { c 1 , . . . , c k } . Purity repr esents percentage of the total number of nodes classified correctly , and it is defined as [50]: P ur ity (Ω , C ) = 1 n X k max j | ω k ∩ c j | where n is the total number of nodes, and | ω k ∩ c j | represents the number of nodes in the intersection of ω k and c j . T o trade-off the quality of the clustering against the number of clusters we use NMI . NMI is defined as [51]: N M I (Ω , C ) = I (Ω; C ) | H (Ω) + H ( C ) | / 2 where I is the mutual information between node clusters Ω and classes C , while H (Ω) and H ( C ) r epresent the entropy of clusters and classes respectively . Finally , Rand Index rep- resents percentage of true positive ( T P ) and true negative ( T N ) decisions assigns that are correct (i.e., accuracy). It is defined as: RI (Ω , C ) = T P + T N T P + F P + F N + T N where, F P and F N repr esent false positive and false neg- ative decisions respectively . ARI is defined to be scaled in range [0 , 1] [51]. All three measures are in the range [0 , 1] , and the higher their value, the better clustering quality is. Unknown ground truth . For biological networks, the ground truth clusters are unknown and evaluating the clustering results becomes more challenging. In order to evaluate the functional modules identified by our methods, we use Gene Ontology (GO) [52], a commonly used gene an- notation database. GO repr esents a systematic classification of all known protein functions organized as well-defined terms (also known as GO terms) divided into three main categories, namely Molecular Function (MF), Biological Pr o- cess (BP) and Cellular Component (CC) [52]. GO terms (i.e., annotations), representing gene functions, are hierar chically structured where low-level (general) terms annotate more proteins than high-level (specific) terms. Thus, in our analy- sis we aim to evaluate our clusters with high-level (specific) GO terms annotating not mor e than 100 genes. Additionally , we r emove GO terms annotating 2 or less proteins. Thus, for each gene in a network, we create a list of its corresponding GO term annotations. W e then analyze the consistency of each cluster , i , obtain by our method, by computing the redundancy [6], R i as follows: R i = 1 − − N GO P l =1 p l log 2 p l log 2 N GO where, N GO repr esents the total number of GO terms considered and p l repr esents the relative frequency of GO term l in cluster i . Redundancy is based on normalized Shannon’s entropy and its values range between 0 and JOURNAL OF L A T E X CLASS FILES, V OL. 14, NO . 8, AUGUST 2015 9 Fig. 2. The cluster ing performance of our proposed and other methods on 11 different SYNTH-C multiplex networ ks measured by NMI. On x - axis we present within-community probability , representing the density of connections of communities in the two complementary lay ers. 1. For clusters in which the majority of genes share the same GO terms (annotations) redundancy is close to 1, whereas for clusters in which the majority of genes have disparate GO terms the redundancy is close to 0. When comparing clustering results obtained by differ ent methods, we use the value of redundancy averaged over all clusters. Furthermore, the redundancy is a very suitable measure for clustering performance comparisons because its value does not depend on the number of clusters and unlike others evaluation measures for biological network clustering [53], it is parameter-free. 5 R E S U L T S A N D D I S C U S S I O N 5.1 Clustering evaluation on artificial m ultiplex net- works The ability of our pr oposed methods to extract clusters fr om complementary layers, repr esented by SYNTH-C networks, is shown in Figure 2. The performance of our methods is compared with other methods and it is measured by NMI. By decreasing the within-community probability of comple- mentary clusters in both layer , i.e., by decr easing the density of connections withing communities and thus making com- munities harder to detect, we see a drastic decrease of per- formance in many methods, including SC-ML, PMM, SNF and CGC-NMF (Fig. 2). Furthermore, below some value of within-community probability , i.e., < 0 . 1 , the performance of these methods is equal or close to zero. Unlike them, our proposed methods, particularly CSNMF , CSsNMTF and CPNMF show significantly better performance. Specifically , CPNMF demonstrates constant performance for all values of withing-community probability . The similar results can also be observed for GraphFuse and LMF . Given that, we can observe that CPNMF method is mostly successful in utilizing complementary information contained in all layers and achieving the highest clustering results. In terms of noise, incorporated into SYNTH-N networks, the ranking between the methods in terms of cluster- ing performance is dif ferent. By increasing the between- community probability of the first layer , and thus intro- ducing more noise between communities, the clustering performance of all methods decreases (Fig. 3). Our proposed methods, CSNMF , CSNMTF and CSsNMTF , along with SC-ML demonstrate the best performance across different values of within-community probability , which makes them more robust to noise than other methods. On the other hand, other methods methods are characterized with sig- nificantly lower clustering performance. Surprisingly , CP- NMF method, which is giving the best performance for complementary layers, perform significantly worse on nosy networks than other methods. 5.2 Clustering evaluation on real-world multiplex net- works T able 3 presents the Purity , NMI and ARI of our four proposed collective factorization methods, along with five differ ent baseline methods and six differ ent widely used state-of-the-art methods on six different real-world net- works. The first important observation is that all four pro- posed collective NMF methods (CSNMF , CPNMF , CSNMTF and CSsemi-NMTF) perform better than their correspond- ing baseline methods (SNMF , PNMF , SNMTF and Ssemi- NMTF) on all real-world multiplex networks. Thus, the strategy of merging layers into a single layer always leads to underperformance. Moreover , single-layer modularity maximization (MM) algorithm is outperformed by baseline, single-layer NMF methods in almost all real-world net- works, except for WTN networks where MM significantly outperforms baseline NMF methods, and SND(o) wher e MM performs better than SNMF , SNMTF and Ssemi-NMTF , but not better than PNMF . In comparison to the state-of- the-art methods (PMM, SNF , SC-ML, LMF , GraphFuse and CGC-NMF), at least one of our proposed methods out- performs them all (in terms of either Purity , NMI or ARI or all three measures) in all real-world multiplex network. Moreover , for example, on MPD network, both CSNMF and CSsemi-NMTF perform better than all other methods, with CSemi-NMTF being the best in terms of Purity and NMI; on SND(s) network, CSNMF , CSNMTF and CSsemi-NMTF perform better than all other methods, with CSNMTF per- forming the best in terms of all three measures; on WBN Fig. 3. The cluster ing performance of our proposed and other methods on 10 different SYNTH-N multiplex networ ks measured by NMI. On x - axis we present between-community probability , representing the noise lev el between communities in the first la yer . JOURNAL OF L A T E X CLASS FILES, V OL. 14, NO . 8, AUGUST 2015 10 T ABLE 3 Clustering accuracy measures for methods (from left to right): MM, SNMF , PNMF , SNMTF , SsNMTF , PMM, SNF , SC-ML, LMF , GraphFuse, CGC-NMF , CSNMF , CPNMF , CSNMTF , CSsNMTF applied on real-world multi-lay er networks (from top to bottom): CiteSeer , CoRA , MPD , SND , WBN , WTN , HBN , YBN and MBN . MM SNMF PNMF SNMTF SsNMTF PMM SNF SC-ML LMF GF CGC CSNMF CPNMF CSNMTF CSsNMTF CiteSeer Purity 0.500 0.407 0.405 0.377 0.371 0.302 0.214 0.419 0.235 0.512 0.212 0.501 0.519 0.416 0.404 NMI 0.187 0.222 0.221 0.170 0.164 0.145 0.023 0.191 0.013 0.211 0.013 0.237 0.216 0.172 0.195 ARI 0.152 0.059 0.057 0.042 0.038 0.008 0.001 0.169 0.005 0.201 0.001 0.207 0.185 0.093 0.089 CoRA Purity 0.706 0.669 0.660 0.669 0.669 0.496 0.733 0.787 0.492 0.642 0.678 0.802 0.790 0.683 0.684 NMI 0.340 0.385 0.353 0.382 0.382 0.085 0.449 0.480 0.002 0.201 0.389 0.514 0.480 0.346 0.390 ARI 0.257 0.280 0.247 0.277 0.277 0.030 0.470 0.485 0.001 0.209 0.296 0.491 0.470 0.279 0.288 MPD Purity 0.563 0.678 0.666 0.620 0.678 0.689 0.620 0.701 0.471 0.689 0.678 0.701 0.655 0.655 0.724 NMI 0.313 0.471 0.466 0.384 0.473 0.533 0.395 0.495 0.191 0.565 0.457 0.504 0.451 0.458 0.521 ARI 0.147 0.268 0.259 0.228 0.272 0.383 0.280 0.379 0.029 0.411 0.357 0.394 0.368 0.346 0.422 SND(o) Purity 0.929 0.943 0.943 0.676 0.943 0.943 0.943 0.943 0.788 0.943 0.943 0.943 0.943 0.943 0.943 NMI 0.618 0.681 0.681 0.133 0.681 0.675 0.689 0.681 0.303 0.675 0.673 0.681 0.685 0.773 0.678 ARI 0.460 0.493 0.493 0.021 0.493 0.477 0.515 0.493 0.239 0.477 0.472 0.493 0.503 0.811 0.484 SND (s) Purity 0.619 0.577 0.633 0.634 0.619 0.591 0.633 0.591 0.633 0.619 0.662 0.633 0.633 0.747 0.605 NMI 0.038 0.025 0.052 0.055 0.041 0.037 0.057 0.030 0.053 0.045 0.0781 0.053 0.053 0.276 0.034 ARI 0.024 0.012 0.058 0.058 0.043 0.022 0.058 0.021 0.059 0.044 0.092 0.059 0.059 0.234 0.031 WBN Purity 0.473 0.512 0.501 0.476 0.523 0.473 0.534 0.272 0.283 0.509 0.516 0.577 0.537 0.548 0.530 NMI 0.333 0.382 0.400 0.327 0.363 0.373 0.425 0.079 0.098 0.426 0.370 0.463 0.432 0.404 0.424 ARI 0.199 0.226 0.213 0.112 0.180 0.130 0.211 0.001 0.009 0.216 0.211 0.291 0.233 0.237 0.225 WTN Purity 0.506 0.475 0.464 0.388 0.284 0.388 0.289 0.497 0.453 0.415 0.278 0.579 0.420 0.371 0.420 NMI 0.231 0.269 0.242 0.176 0.077 0.205 0.073 0.226 0.191 0.176 0.072 0.322 0.172 0.154 0.155 ARI 0.080 0.114 0.114 0.073 0.001 0.039 0.005 0.133 0.094 0.107 0.002 0.160 0.094 0.035 0.088 HBN 0.180 0.325 0.350 0.322 0.326 0.351 0.141 0.266 0.045 0.203 0.320 0.341 0.364 0.339 0.342 YBN R avg 0.027 0.374 0.336 0.372 0.358 0.343 0.163 0.257 0.100 0.114 0.311 0.383 0.342 0.383 0.381 MBN 0.015 0.416 0.387 0.462 0.441 0.355 0.211 0.320 0.180 0.298 0.328 0.422 0.401 0.416 0.433 network, both CSNMF and CSNMTF perform better than all other methods, with CSNMTF being the best in terms of Purity and ARI, and CSNMF being the best in terms of NMI; on WTN network, CSNMF , CSNMTF and CSsemi-NMTF perform better than other methods, with CSNMF being the best in terms of all three measures. 5.2.1 Clustering ev aluation on multiple x biological networks In table 3, we also present the average redundancy ( R av g ) obtained by clustering multiplex biological networks with our four methods, as well as with state-of-the-art and baseline methods. The results, again, indicate the superior perfor- mance of our methods over the state-of-the-art and baseline methods, except in the case of MBN, where SNMTF , applied on merged network layers, yields the highest redundancy . Furthermore, we compare the functional consistency of clusters obtain by collective integration of all network layers with the functional consistency of clusters obtained from each individual network layer . The results for all three biological networks, obtained by applying SNMF on each individual network layer and CSNMF on all network layers together , are depicted in Fig. 4. W e observe that each in- dividual network layer has biologically consistent clusters. However , the highest level of biological consistency , mea- sured by the average redundancy , is achieved when all the layers are fused together (red bars in Fig.4). 6 C O N C L U S I O N In this paper , we address the problem of composite commu- nity detection in multiplex networks by proposing NF-CCE, a general model consisting of four novel methods, CSNMF , CPNMF , CSNMTF and CSsemi-NMTF , based on four non- negative matrix factorization techniques. Each of the pro- posed method works in a similar way: in the first step, it decomposes adjacency matrices representing network layers Fig. 4. Av erage redundancy of each individual network lay er (in blue), computed by SNMF , and of their fused representation (in red), com- puted by CSNMF , f or networks: (A) HBN, (B) YBN and (C) MBN. Method parameters are: k = 300 and α = 0 . 01 . into low-dimensional, non-negative feature matrices; then, in the second step, it fuses the featur e matrices of layers into a consensus non-negative, low-dimensional feature matrix common to all network layers, from which the composite clusters are extracted. The second step is done by collective matrix factorization that maximizes the shared information JOURNAL OF L A T E X CLASS FILES, V OL. 14, NO . 8, AUGUST 2015 11 between network layers by optimizing the distance between each of the non-negative feature matrices representing lay- ers and the consensus feature matrix. The ability of our methods to integrate complementary as well as noisy network layers more efficiently than the state-of-the-art methods has been demonstrated on artifi- cially generated multiplex networks. In terms of cluster- ing accuracy , we demonstrate the superior performance of our proposed methods over the baseline and state-of-the-art methods on nine real-world networks. W e show that sim- ple averaging of adjacency matrices repr esenting network layers (i.e., merging network layers into a single network repr esentation), the strategy that is usually practiced in the literature, leads to the worst clustering performance. More- over , our experiments indicate that widely-used modularity maximization methods are significantly outperformed by NMF-based methods. NF-CCE can be applied on multiplex networks from differ ent domains, ranging from social, phone communica- tion and bibliographic networks to biological, economic and brain networks, demonstrating the diverse applicability of our methods. R E F E R E N C E S [1] M. Newman, Networks: An Introduction . New Y ork, NY , USA: Oxford University Press, Inc., 2010. [2] S. Boccaletti, V . Latora, Y . Moreno, M. Chavez, and D.-U. Hwang, “Complex networks: Structure and dynamics,” Physics Reports , vol. 424, no. 45, pp. 175 – 308, 2006. [3] S. H. Strogatz, “Exploring complex networks,” Nature , vol. 410, pp. 268–276, 2001. [4] S. Fortunato, “Community detection in graphs,” Physics Reports , vol. 486, no. 3–5, pp. 75–174, 2010. [5] M. Girvan and M. E. J. Newman, “Community structure in social and biological networks,” Proceedings of the National Academy of Sciences , vol. 99, no. 12, pp. 7821–7826, 2002. [6] J. B. Pereira-Leal, A. J. Enright, and C. A. Ouzounis, “Detection of functional modules from protein interaction networks,” Proteins: Structure, Function, and Bioinformatics , vol. 54, no. 1, pp. 49–57, 2004. [7] J. Leskovec, K. J. Lang, and M. Mahoney , “Empirical comparison of algorithms for network community detection,” in Proceedings of the 19th International Conference on World Wide Web , ser . WWW ’10. New Y ork, NY , USA: ACM, 2010, pp. 631–640. [8] J. Chen and B. Y uan, “Detecting functional modules in the yeast proteinpr otein interaction network,” Bioinformatics , vol. 22, no. 18, pp. 2283–2290, 2006. [9] J. Duch and A. Arenas, “Community detection in complex net- works using extremal optimization,” Phys. Rev. E , vol. 72, p. 027104, Aug 2005. [10] V . D. Blondel, J.-L. Guillaume, R. Lambiotte, and E. Lefebvr e, “Fast unfolding of communities in large networks,” Journal of Statistical Mechanics: Theory and Experiment , vol. 2008, no. 10, p. P10008, 2008. [11] M. Mitrovi ´ c and B. T adi ´ c, “Spectral and dynamical properties in classes of sparse networks with mesoscopic inhomogeneities,” Phys. Rev . E , vol. 80, p. 026123, Aug 2009. [12] M. A. Porter , J.-P . Onnela, and P . J. Mucha, “Communities in networks,” Notices Amer . Math. Soc. , vol. 56, no. 9, pp. 1082–1097, 2009. [13] S. E. Schaeffer , “Survey: Graph clustering,” Comput. Sci. Rev . , vol. 1, no. 1, pp. 27–64, Aug. 2007. [14] M. Kivel ¨ a, A. Arenas, M. Barthelemy , J. P . Gleeson, Y . Moreno, and M. A. Porter , “Multilayer networks,” Journal of Complex Networks , 2014. [15] J. Kim and J.-G. Lee, “Community detection in multi-layer graphs: A survey ,” SIGMOD Rec. , vol. 44, no. 3, pp. 37–48, dec 2015. [16] D. Kuang, H. Park, and C. H. Ding, “Symmetric nonnegative matrix factorization for graph clustering.” in SDM , vol. 12. SIAM, 2012, pp. 106–117. [17] F . W ang, T . Li, X. W ang, S. Zhu, and C. Ding, “Community dis- covery using nonnegative matrix factorization,” Data Min. Knowl. Discov . , vol. 22, no. 3, pp. 493–521, May 2011. [18] X. Dong, P . Frossard, P . V andergheynst, and N. Nefedov , “Clus- tering on multi-layer graphs via subspace analysis on grassmann manifolds,” T rans. Sig. Pr oc. , vol. 62, no. 4, pp. 905–918, feb 2014. [19] S.-I. Amari, “Natural gradient works efficiently in learning,” Neu- ral Comput. , vol. 10, no. 2, pp. 251–276, feb 1998. [20] Y . Panagakis, C. Kotropoulos, and G. R. Arce, “Non-negative multilinear principal component analysis of auditory temporal modulations for music genre classification,” IEEE T ransactions on Audio, Speech, and Language Processing , vol. 18, no. 3, pp. 576–588, March 2010. [21] W . Cheng, X. Zhang, Z. Guo, Y . W u, P . F . Sullivan, and W . W ang, “Flexible and robust co-regularized multi-domain graph cluster- ing,” in Proceedings of the 19th ACM SIGKDD International Confer- ence on Knowledge Discovery and Data Mining , ser . KDD ’13. New Y ork, NY , USA: ACM, 2013, pp. 320–328. [22] L. T ang, X. W ang, and H. Liu, “Uncovering groups via hetero- geneous interaction analysis,” in Proceedings – IEEE International Conference on Data Mining, ICDM , 2009, pp. 503–512. [23] X. Dong, P . Frossard, P . V andergheynst, and N. Nefedov , “Clus- tering with multi-layer graphs: A spectral perspective,” IEEE T ransactions on Signal Processing , vol. 60, no. 11, pp. 5820–5831, Nov 2012. [24] D. D. Lee and H. S. Seung, “Algorithms for non-negative matrix factorization,” in In NIPS . MIT Press, 2000, pp. 556–562. [25] D. B. W est et al. , Introduction to graph theory . Pr entice hall Upper Saddle River , 2001, vol. 2. [26] J. Shi and J. Malik, “Normalized cuts and image segmentation,” IEEE T rans. Pattern Anal. Mach. Intell. , vol. 22, no. 8, pp. 888–905, Aug 2000. [27] U. von Luxburg, “A tutorial on spectral clustering,” Statistics and Computing , vol. 17, no. 4, pp. 395–416, 2007. [28] T . Hastie, R. T ibshirani, and J. Friedman, The Elements of Statistical Learning . Springer , 2001. [29] M. E. J. Newman, “Modularity and community structure in net- works,” Proceedings of the National Academy of Sciences , vol. 103, no. 23, pp. 8577–8582, 2006. [30] P . W . Holland, K. B. Laskey , and S. Leinhardt, “Stochastic block- models: First steps,” Social Networks , vol. 5, no. 2, pp. 109 – 137, 1983. [31] M. G. Everett and S. P . Borgatti, “Analyzing clique overlap,” Connections , vol. 21, no. 1, pp. 49–61, 1998. [32] S. Zhang, H. Zhao, and M. K. Ng, “Functional module analy- sis for gene coexpression networks with network integration,” IEEE/ACM T rans. Comput. Biol. Bioinformatics , vol. 12, no. 5, pp. 1146–1160, Sep. 2015. [33] M. Berlingerio, M. Coscia, and F . Giannotti, “Finding and charac- terizing communities in multidimensional networks,” in Advances in Social Networks Analysis and Mining (ASONAM), 2011 Interna- tional Conference on , July 2011, pp. 490–494. [34] M. A. Rodriguez and J. Shinavier , “Exposing multi-relational net- works to single-relational network analysis algorithms,” Journal of Informetrics , vol. 4, no. 1, pp. 29 – 41, 2010. [35] P . J. Mucha, T . Richardson, K. Macon, M. A. Porter , and J.-P . Onnela, “Community structure in time-dependent, multiscale, and multiplex networks,” Science , vol. 328, no. 5980, pp. 876–878, 2010. [36] B. W ang, A. M. Mezlini, F . Demir , M. Fiume, Z. T u, M. Brudno, B. Haibe-Kains, and A. Goldenberg, “Similarity network fusion for aggregating data types on a genomic scale,” Nature Methods , vol. 11, no. 3, pp. 333–337, 2014. [37] W . T ang, Z. Lu, and I. S. Dhillon, “Clustering with multiple graphs,” in Proceedings of the 2009 Ninth IEEE International Con- ference on Data Mining , ser . ICDM ’09. W ashington, DC, USA: IEEE Computer Society , 2009, pp. 1016–1021. [38] E. E. Papalexakis, L. Akoglu, and D. Ienco, “Do more views of a graph help? community detection and clustering in multi-graphs,” in Proceedings of the 16th International Conference on Information Fusion, FUSION 2013, Istanbul, T urkey, July 9-12, 2013 , 2013, pp. 899–905. [39] R. A. Harshman, “Foundations of the P ARAF AC procedure: Mod- els and conditions for an” explanatory” multi-modal factor analy- sis,” UCLA Working Papers in Phonetics , vol. 16, no. 1, p. 84, 1970. [40] Z. Y ang and E. Oja, “Linear and nonlinear projective nonnegative matrix factorization,” IEEE T ransactions on Neural Networks , vol. 21, no. 5, pp. 734–749, May 2010. JOURNAL OF L A T E X CLASS FILES, V OL. 14, NO . 8, AUGUST 2015 12 [41] C. Ding, T . Li, W . Peng, and H. Park, “Orthogonal nonnegative matrix T-factorizations for clustering,” in Proceedings of the 12th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining , ser . KDD ’06. New Y ork, NY , USA: ACM, 2006, pp. 126–135. [42] C. H. Q. Ding, T . Li, and M. I. Jordan, “Convex and semi- nonnegative matrix factorizations,” IEEE T ransactions on Pattern Analysis and Machine Intelligence , vol. 32, no. 1, pp. 45–55, Jan 2010. [43] S. Boyd and L. V andenberghe, Convex Optimization . New Y ork, NY , USA: Cambridge University Press, 2004. [44] A. Condon and R. M. Karp, “Algorithms for graph partitioning on the planted partition model,” Random Struct. Algorithms , vol. 18, no. 2, pp. 116–140, Mar . 2001. [45] P . Sen, G. M. Namata, M. Bilgic, L. Getoor , B. Gallagher , and T . Eliassi-Rad, “Collective classification in network data,” AI Mag- azine , vol. 29, no. 3, pp. 93–106, 2008. [46] T . A. B. Snijders, P . E. Pattison, G. L. Robins, and M. S. Hand- cock, “New specifications for exponential random graph models,” Sociological Methodology , vol. 36, no. 1, pp. 99–153, 2006. [47] J. G. White, E. Southgate, J. N. Thomson, and S. Brenner , “The structure of the nervous system of the nematode caenorhabditis elegans,” Philosophical T ransactions of the Royal Society of London B: Biological Sciences , vol. 314, no. 1165, pp. 1–340, 1986. [48] M. De Domenico, V . Nicosia, A. Arenas, and V . Latora, “Structural reducibility of multilayer networks,” Natur e communications , vol. 6, p. 6864, 2015. [49] S. Mostafavi and Q. Morris, “Fast integration of heterogeneous data sources for predicting gene function with limited annota- tion,” Bioinformatics , vol. 26, no. 14, pp. 1759–1765, 2010. [50] Y . Zhao and G. Karypis, “Empirical and theoretical comparisons of selected criterion functions for document clustering,” Mach. Learn. , vol. 55, no. 3, pp. 311–331, jun 2004. [51] C. D. Manning, P . Raghavan, and H. Sch ¨ utze, Introduction to Information Retrieval . New Y ork, NY , USA: Cambridge University Press, 2008. [52] M. Ashburner , C. A. Ball, J. A. Blake, D. Botstein, H. Butler , J. M. Cherry , A. P . Davis, K. Dolinski, S. S. Dwight, J. T . Eppig, M. A. Harris, D. P . Hill, L. Issel-T arver , A. Kasarskis, S. Lewis, J. C. Matese, J. E. Richardson, M. Ringwald, G. M. Rubin, and G. Sherlock, “Gene Ontology: tool for the unification of biology ,” Nature Genetics , vol. 25, no. 1, pp. 25–29, May 2000. [53] Y .-K. Shih and S. Parthasarathy , “Identifying functional modules in interaction networks through overlapping markov clustering,” Bioinformatics , vol. 28, no. 18, pp. i473–i479, 2012.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment