Multi-Organ Cancer Classification and Survival Analysis
Accurate and robust cell nuclei classification is the cornerstone for a wider range of tasks in digital and Computational Pathology. However, most machine learning systems require extensive labeling from expert pathologists for each individual proble…
Authors: Stefan Bauer, Nicolas Carion, Peter Sch"uffler
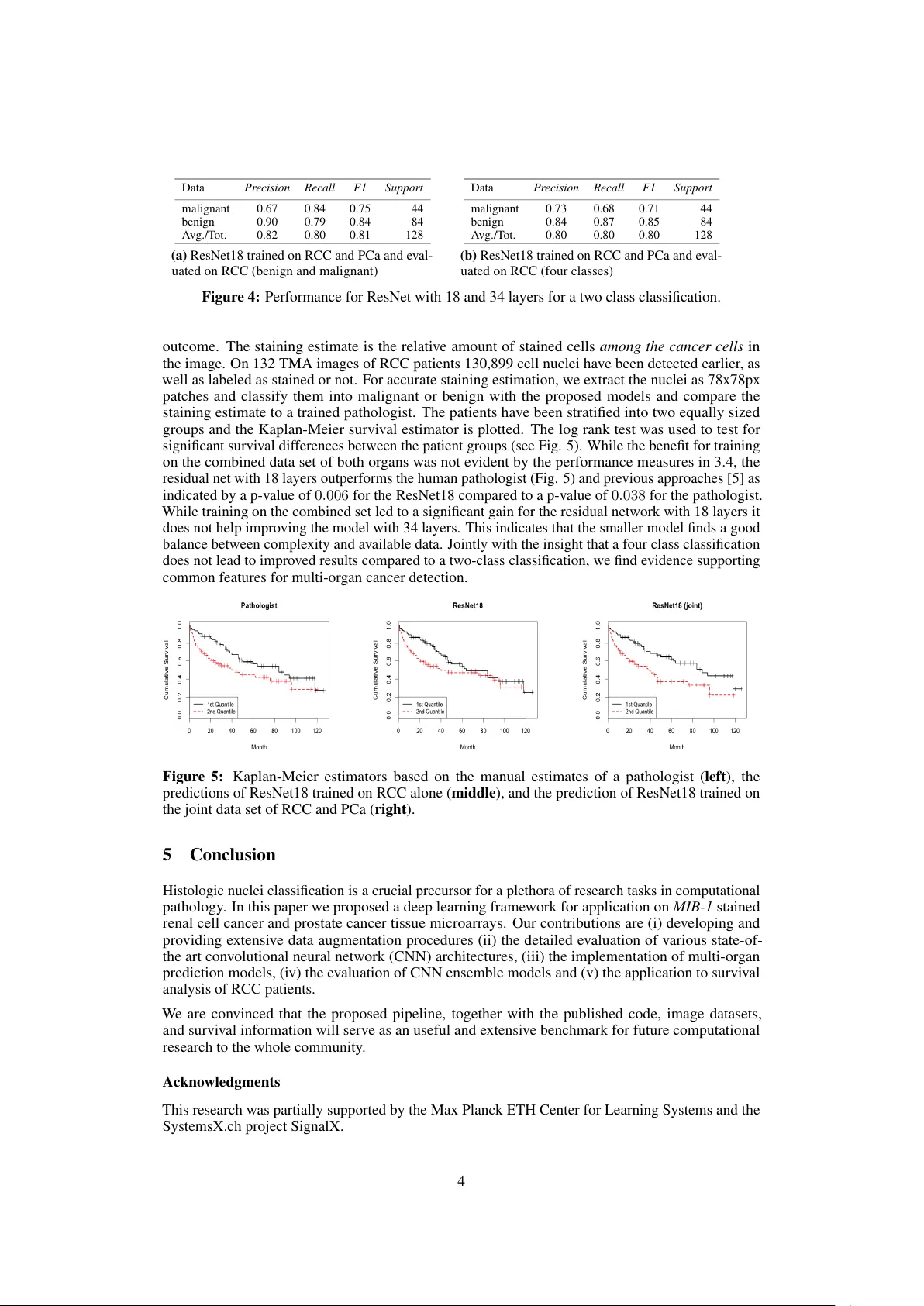
Multi-Organ Cancer Classification and Sur viv al Analysis Stefan Bauer , Nicolas Carion, Joachim M. Buhmann Department of Computer Science ETH Zurich, Switzerland {bauers, ncarion} @inf.ethz.ch jbuhmann @inf.ethz.ch Peter Schüffler , Thomas Fuchs Memorial Sloan Kettering Cancer Center New Y ork, USA {schueffp, fuchst}@mskcc.org Peter W ild Institute of Surgical P athology Univ ersity Hospital Zurich peter.widl@usz.ch Abstract Accurate and robust cell nuclei classification is the cornerstone for a wider range of tasks in digital and Computational Pathology . Ho wev er , most machine learning systems require extensi ve labeling from e xpert pathologists for each individual problem at hand, with no or limited abilities for knowledge transfer between datasets and organ sites. In this paper we implement and ev aluate a variety of deep neural network models and model ensembles for nuclei classification in renal cell cancer (RCC) and prostate cancer (PCa). W e propose a con volutional neural network system based on residual learning which significantly improv es ov er the state-of-the-art in cell nuclei classification. Finally , we sho w that the combination of tissue types during training increases not only classification accuracy b ut also ov erall surviv al analysis. 1 Introduction T o facilitate automated cancer diagnosis and prognosis, computational pathology is pro viding fully automated image analysis pipelines, e.g. [ 5 ], [ 4 ] or [ 13 ]. While these results already match or surpass the classification accuracy of expert pathologists [ 1 ], they require extensi ve feature engineering and extensi ve e xpert labels for specific cancer types. The ongoing success in machine learning and computer vision demonstrates the remarkable learning abilities of deep networks for image recognition [e.g. 12 ]. Deep learning algorithms hav e already been successfully applied in computational pathology , e.g. for the ICPR 2012 Contest on Mitosis Detection in Breast Cancer Histological Images [ 2 ] and similarly for the MICCAI 2013 Grand Challenge [17]. Our main moti vation is to in v estigate the performance of networks trained from scratch with fix ed parameters to see the transfer of learned concepts from one organ to the next. T o the best of our knowledge, there exists only very limited information on common features of cancer cells from different organs, thus potentially requiring the tailoring of current frame works to specific cancer scoring tasks. In 2 we describe both datasets and provide benchmark comparisons in 3. Given the typical small data sets in computational pathology , we focus on data augmentation in 3.1 and study the learning of neural networks of dif ferent depths and ensembles when trained from scratch in 3.2 and 3.3, as well as the transfer of trained networks from one organ to the ne xt (with fixed parameters) and on the joint data set with two, three or four classes in 3.4. W e additionally v alidate our results by sho wing that not 30th Conference on Neural Information Processing Systems (NIPS 2016), Barcelona, Spain. only the classification accuracy is impro ved b ut also o verall survi val analysis in 4. Our approaches significantly improv e ov er state-of-the-art in cell nuclei classification and suggest common patterns in renal cell carcinoma (RCC) and prostate cancer (PCa). In addition we provide our code and data for bench-marking and future research in computational pathology . 2 Data Renal Cell Carcinoma (RCC) belongs to the 10 most common cancers in western societies’ mor - tality [ 7 ]. Clear cell renal cell carcinoma (ccRCC) is a common subtype of RCC occurring on cells with clear cytoplasm. Since this cancer dev elops metastases in a very early stage (commonly before diagnosis), the prognosis for RCC patients is usually pessimistic [ 16 ]. T issue microarrays (TMA) serve as an important tool for molecular biomarker discovery , since they enable the screening of dozens or ev en hundreds of specimen simultaneously . Our data basis are eight ccRCC TMA images. Each image is fully labeled by tw o pathologists, indicating location and class of all malignant and benign cell nuclei. From 1633 found nuclei, the two pathologists agreed with the labels on 1272. These 1272 well-labeled nuclei (890 benign and 382 malignant) were extracted as patches of size 78x78 pixels centered at labeled nuclei and serv e as our original study data (see Fig 1). This dataset was first published and analysed in [5] and serves jointly with [13] as a comparison. Prostate cancer (PCa) is one of the most common cancer types in western male society . It is the second most frequently diagnosed cancer for human males worldwide, and the sixth leading cause of cancer related death [ 9 ]. Howe ver , research is ongoing for the de velopment of specific biomarkers for the early diagnosis and the deeper understanding of PCa [ 6 ]. W e incorporate six new TMA images of PCa patients, twice labeled by two pathologists. From 1195 detected nuclei, they agreed on the label of 826 (207 benign, 619 malignant). 3 Experiments (a) Renal cell (b) Prostate cell Figure 1: Examples of 78 × 78 patches. W e used the Caffe library [ 10 ] to train variants of small Cifar10 [ 11 ], AlexNet [ 12 ], ImageNet [ 3 ] and googlenet [ 15 ] like deep networks. Giv en our small data set we additionally implemented the newly dev eloped residual networks [ 8 ], which outperforms pre vious approaches in the ImageNet competition. A residual network with 18 layers is denoted by ResNet18 and one with 34 layers by ResNet34. All larger models e.g. ImageNet quickly led to ov erfitting and poor results due to the small sample size. Additionally , we tested the inclusion of custom weights in the cost function e.g. for AlexNet [ 12 ], in order to ov ercome class biases. Ho wev er , experiments sho wed that this compensation does not impro ve the error rates. While the code for the best performing ResNet’ s is already provided by Facebook https://github.com/facebook/fb.resnet.torch , we provide our code for the customized data augmentation and both data sets for bench-marking and future research. The best classifier using hand-crafted feature engineering, achie ves a classification accuracy of 83% for RCC [ 13 ], which is as good as the manual annotation: the inter -pathologist accuracy for classification of 1633 renal clear cell carcinoma nuclei is 80% . Replicating the approaches in [ 13 ] for PCa, these values ev en increase up to 90% . The automated staining estimation pipeline is implemented in the free Jav a program TMARKER [ 14 ]. The reported performance measures are r ecall , pr ecision , F1-scor e and support . 3.1 Data augmentation W e randomly split the data into 80 % , 10% and 10% for training, testing and v alidation. Due to the computational cost we only apply a one fold cross v alidation and split and average the data only once. For comparison for some nets the results of a double split experiment are reported. Giv en the low number of samples av ailable one focus of our work is data augmentation and we apply the following techniques to the training set while av eraging over all predictions for the v alidation set: firstly , the nucleus patch is scaled down to a randomly chosen size in [64 : 78] . After that, we select uniformly at random a cr opping of size 64 × 64 , and we mirr or it with probability 1 / 2 . Since only the shape, and not the orientation or the color of the nuclei is discriminative for classifying it as malignant or benign 2 [ 13 ], we also apply a r otation by a random angle between 0 ◦ and 360 ◦ , and in addition grayscaling . Each picture is randomly perturbed 50 times, giving alltogether 60T pictures in the RCC and 40T for the PCa dataset. 3.2 RCC Using a random partition with 80% for training and 10% for testing and validation, the performance is comparable or significantly better to the hand-crafted approach in [ 13 ] with a score of 83% (2b). While the ov erall performance of the Cifar10 net is comparable to the hand-crafted approach in [ 13 ], it apparently has difficulties with the prediction for the malignant cells as indicated by low precision and recall. Due to the limited number of samples in the v alidation set a multiple random partitions into training, testing and validation might already reduce the chance of an unfav orable validation set, as shown for the residual netw orks. Data Precision Recall F1 Support malignant 0.79 0.86 0.83 44 benign 0.93 0.88 0.90 84 A vg./T ot. 0.88 0.88 0.88 128 (a) ResNet18 on RCC Data Precision Recall F1 Support malignant 0.79 0.68 0.73 44 benign 0.84 0.90 0.87 84 A vg./T ot. 0.83 0.83 0.82 128 (b) ResNet34 on RCC Figure 2: Renal cell carcinoma (RCC) performance comparison of residual networks (ResNets) with different number of layers. 3.3 PCa In addition to the renal cell carcinoma data, we tested the different deep learning architectures on MIB-1 stained prostate cancer TMAs. The performance for both the residual networks with 18 and 34 layers is close to the intersection of two pathologists Fig. 3a and Fig. 3b. Howe ver both nets misclassify a different patch. An ensemble of both nets with equal weights only misclassifies one of the two patches, since the confidence of the residual net with 34 layers is high enough to o vercome the wrong label of the residual net with 18 layers. When trained on the combined data of RCC and PCa, all pictures (and thus the two pictures as well) are correctly classified by both ResNet18 and ResNet34 (see Section 3.4). Data Precision Recall F1 Support malignant 0.99 1.00 0.99 69 benign 1.00 0.93 0.96 14 A vg./T ot. 0.99 0.99 0.99 83 (a) ResNet18 on PCa Data Precision Recall F1 Support malignant 0.99 1.00 0.99 69 benign 1.00 0.93 0.96 14 A vg./T ot. 0.99 0.99 0.99 83 (b) ResNet34 on PCa Figure 3: Performance of networks with dif ferent layers for prostate cancer . While only one sample is misclassified for both networks, it is a dif ferent patch each time. 3.4 Multi-organ RCC and PCa data For the multi-or gan cancer classification we conducted two dif ferent kinds of experiments: first, we run a four class classification with two classes per org an (i.e. malignant and benign for prostate and malignant and benign for renal cells); second, we only used two classes (i.e. malignant and benign). While the ev aluation on the RCC set decreases the accuracy on the validation set to 80% (see 4b), the true performance might be higher . The training accuracy is around 86% and the surviv al analysis in Section 4 shows a significant impro vement for the RCC data. Like wise, the performance on the PCa data is improved since no w no sample is miss-classified and we exactly replicate the results from the intra-pathologist agreement. W e regard it as a positi ve feature that: No cell of one or gan was labeled as cell of a differ ent or gan. Similarly , the two-class residual networks with 18 and 34 layers trained on the combined data set ha ve 100% accuracy on the PCa data while the performance for the RCC drops to 80% . In addition to the residual networks, the Cifar10 model trained on the joint set and validated on the PCa data achie ves very good results. 4 Surviv al Analysis In addition to classification, we tested our approaches on follow-up surviv al data on 132 RCC patient. In RCC, the staining estimate of the proliferation protein MIB-1 is corelated with the overall survi val 3 Data Pr ecision Recall F1 Support malignant 0.67 0.84 0.75 44 benign 0.90 0.79 0.84 84 A vg./T ot. 0.82 0.80 0.81 128 (a) ResNet18 trained on RCC and PCa and e v al- uated on RCC (benign and malignant) Data Pr ecision Recall F1 Support malignant 0.73 0.68 0.71 44 benign 0.84 0.87 0.85 84 A vg./T ot. 0.80 0.80 0.80 128 (b) ResNet18 t rained on RCC and PCa and e v al- uated on RCC (four classes) Figur e 4: Performance for ResNet with 18 and 34 layers for a tw o class classification. outcome. The staining estimate is the relati v e amount of stained cells among the cancer cells in the image. On 132 TMA images of RCC patients 130,899 cell nuclei ha v e been detected earlier , as well as labeled as stained or not. F or accurate staining estimation, we e xtract the nuclei as 78x78px patches and classify them into malignant or benign with the proposed models and compare the staining estimate to a trained pathologist. The patients ha v e been stratified into tw o equally sized groups and the Kaplan-Meier survi v al estimator is plotted. The log rank test w as used to test for significant survi v al dif ferences between the patient groups (see Fig. 5). While the benefit for training on the combined data set of both or g ans w as not e vident by the performance measures in 3.4, the residual net with 18 layers outperforms the human pathologist (Fig. 5) and pre vious approaches [ 5 ] as indicated by a p-v alue of 0 . 006 for the ResNet18 compared to a p-v alue of 0 . 038 for the pathologist. While training on the combined set led to a significant g ain for the res idual netw ork with 18 layers it does not help impro ving the model with 34 layers. This indicates that the smaller model finds a good balance between comple xity and a v ailable data. Jointly with the insight that a four class classification does not lead to impro v ed results compared to a tw o-class classification, we find e vidence supporting common features for multi-or g an cancer detection. Figur e 5: Kaplan-Meier estimators based on the manual estimates of a pathologist ( left ), the predictions of ResNet18 t rained on RCC alone ( middle ), and the prediction of ResNet18 trained on the joint data set of RCC and PCa ( right ). 5 Conclusion Histologic nuclei classification is a crucial precursor for a plethora of research tasks in computational pathology . In this paper we proposed a deep learning frame w ork for applicati on o n MIB-1 stained renal cell cancer and prostate cancer tissue microarrays. Our contrib utions are (i) de v eloping and pro viding e xtensi v e data augmentation procedures (ii) the detailed e v aluation of v arious state-of- the art con v olutional neural netw ork (CNN) architectures, (iii) the implementation of multi-or g an prediction models, (i v) the e v aluation of CNN ensemble models and (v) the application to survi v al analysis of RCC patients. W e are con vinced that the proposed pipeline, together with the published code, i mage datasets, and survi v al information will serv e as an useful and e xtensi v e benchmark for fut ure computational research to the whole community . Ackno wledgments This research w as partially supported by the Max Planck ETH Center for Learning Systems and the SystemsX.ch project SignalX. 4 References [1] Manuele Bicego, A ydın Ula¸ s, Peter J. Schüf fler, Umberto Castellani, V ittorio Murino, André Martins, Pedro Aguiar , and Mario Figueiredo. Renal Cancer Cell Classification Using Genera- tiv e Embeddings and Information Theoretic Kernels. In P attern Recognition in Bioinformatics , volume 7036, pages 75–86. 2011. [2] Dan Ciresan, Alessandro Giusti, Luca Gambardella, and Jür gen Schmidhuber . Mitosis detection in breast cancer histology images with deep neural netw orks. Medical Image Computing and Computer-Assisted Intervention MICCAI , 2013. [3] Jia Deng, W ei Dong, Richard Socher , Li-Jia Li, Kai Li, and Li Fei-Fei. ImageNet: A large-scale hierarchical image database. In CVPR , pages 248–255. IEEE, 2009. [4] Thomas Fuchs and Joachim M. Buhmann. Computational Pathology: Challenges and Promises for T issue Analysis. Computerized Medical Imaging and Graphics , 35(7–8):515–530, 2011. [5] Thomas Fuchs, Peter W ild, Holger Moch, and Joachim M. Buhmann. Computational Pathology Analysis of T issue Microarrays Predicts Survi v al of Renal Clear Cell Carcinoma Patients. 5242:1–8, 2008. [6] S Gillessen, I Cima, R Schiess, P W ild, M Kalin, P Schueffler , JM Buhmann, H Moch, R Aebersold, and W Krek. Cancer genetics-guided discovery of serum biomarker signatures for prostate cancer . In ASCO Annual Meeting Pr oceedings , volume 28, page 4564, 2010. [7] David J Grignon and Mingxin Che. Clear cell renal cell carcinoma. Clinics in laboratory medicine , 25(2):305–316, 2005. [8] Kaiming He, Xiangyu Zhang, Shaoqing Ren, and Jian Sun. Deep residual learning for image recognition. arXiv preprint , 2015. [9] Ahmedin Jemal, Freddie Bray , Melissa M Center , Jacques Ferlay , Elizabeth W ard, and Da vid Forman. Global cancer statistics. CA: a cancer journal for clinicians , 61(2):69–90, 2011. [10] Y angqing Jia, Ev an Shelhamer , Jeff Donahue, Ser gey Karaye v , Jonathan Long, Ross Girshick, Sergio Guadarrama, and T rev or Darrell. Caffe: Con volutional architecture for fast feature embedding. arXiv preprint , 2014. [11] Alex Krizhe vsky and Geoffre y Hinton. Learning multiple layers of features from tiny images, 2009. [12] Alex Krizhe vsky , Ilya Sutskever , and Geoffrey E. Hinton. Imagenet classification with deep con volutional neural networks. In Advances in Neural Information Pr ocessing Systems 25 , pages 1106–1114. 2012. [13] Peter Schueffler , Thomas Fuchs, Cheng Soon Ong, V olker Roth, and Joachim M. Buhmann. Computational TMA Analysis and Cell Nucleus Classification of Renal Cell Carcinoma. In Pr oc. of 32nd D A GM confer ence on P attern reco gnition , pages 202–211, 2010. [14] Peter J. Schüffler , Thomas J. Fuchs, Cheng S. Ong, Peter W ild, and Joachim M. Buhmann. TMARKER: A Free Software T oolkit for Histopathological Cell Counting and Staining Estima- tion. J ournal of P athology Informatics , 4(2):2, 2013. [15] Christian Szegedy , W ei Liu, Y angqing Jia, Pierre Sermanet, Scott Reed, Dragomir Anguelo v , Dumitru Erhan, V incent V anhoucke, and Andre w Rabinovich. Going deeper with con volutions. In CVPR , 2015. [16] Andrea T annapfel, Helmut A Hahn, Alexander Katalinic, Rainer J Fietkau, Reinhard Kuehn, and Christian W W ittekind. Prognostic value of ploidy and proliferation markers in renal cell carcinoma. Cancer , 77(1):164–171, 1996. [17] Mitko V eta, Paul J V an Diest, Stefan M W illems, Haibo W ang, Anant Madabhushi, Angel Cruz-Roa, Fabio Gonzalez, Anders BL Larsen, Jacob S V estergaard, Anders B Dahl, et al. Assessment of algorithms for mitosis detection in breast cancer histopathology images. Medical image analysis , 20(1):237–248, 2015. 5
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment