Capacity and Expressiveness of Genomic Tandem Duplication
The majority of the human genome consists of repeated sequences. An important type of repeated sequences common in the human genome are tandem repeats, where identical copies appear next to each other. For example, in the sequence $AGTC\underline{TGTG}C$, $TGTG$ is a tandem repeat, that may be generated from $AGTCTGC$ by a tandem duplication of length $2$. In this work, we investigate the possibility of generating a large number of sequences from a \textit{seed}, i.e.\ a small initial string, by tandem duplications of bounded length. We study the capacity of such a system, a notion that quantifies the system’s generating power. Our results include \textit{exact capacity} values for certain tandem duplication string systems. In addition, motivated by the role of DNA sequences in expressing proteins via RNA and the genetic code, we define the notion of the \textit{expressiveness} of a tandem duplication system as the capability of expressing arbitrary substrings. We then \textit{completely} characterize the expressiveness of tandem duplication systems for general alphabet sizes and duplication lengths. In particular, based on a celebrated result by Axel Thue from 1906, presenting a construction for ternary square-free sequences, we show that for alphabets of size 4 or larger, bounded tandem duplication systems, regardless of the seed and the bound on duplication length, are not fully expressive, i.e. they cannot generate all strings even as substrings of other strings. Note that the alphabet of size 4 is of particular interest as it pertains to the genomic alphabet. Building on this result, we also show that these systems do not have full capacity. In general, our results illustrate that duplication lengths play a more significant role than the seed in generating a large number of sequences for these systems.
💡 Research Summary
The paper investigates the generative power of bounded‑length tandem duplication systems, a model motivated by the prevalence of tandem repeats in the human genome. A tandem duplication system is defined as a triple (Σ, s, T) where Σ is a finite alphabet, s is a finite seed string, and T is a set of duplication rules. The specific rule set T_tan≤k consists of all tandem duplications that copy a substring of length at most k and insert it immediately after its original occurrence. Two quantitative notions are introduced: capacity and expressiveness. Capacity, cap(S) = lim sup_{n→∞} (log_{|Σ|}|S∩Σⁿ|)/n, measures the exponential growth rate of the number of length‑n strings that can be generated, normalized by the alphabet size; its maximum possible value is 1, which corresponds to a system that can essentially generate all strings. A system is fully expressive if every possible string over Σ appears as a substring of some string in the system; lack of full expressiveness necessarily forces capacity to be strictly less than 1.
The authors first illustrate the concepts with a binary example (seed “01”, duplication length ≤2) that generates all strings beginning with 0 and ending with 1, yielding capacity 1 and full expressiveness (any binary word appears as a substring after padding with 0 and 1). They then move to more complex settings where counting and characterization become non‑trivial.
Capacity results.
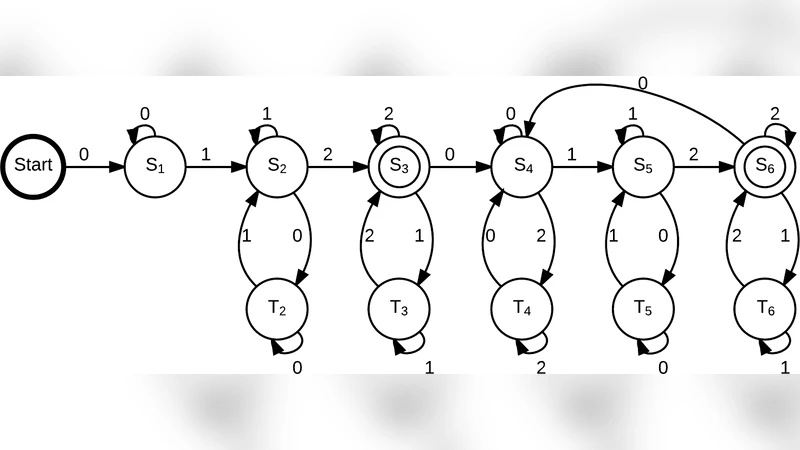
For the ternary system (Σ={0,1,2}, seed “012”, duplication length ≤3) they construct a finite automaton (Fig. 2) that exactly captures the language generated. By applying Perron–Frobenius theory to the automaton’s adjacency matrix they compute the dominant eigenvalue and obtain an exact capacity value:
\
Comments & Academic Discussion
Loading comments...
Leave a Comment