Bag-of-Words Representation for Biomedical Time Series Classification
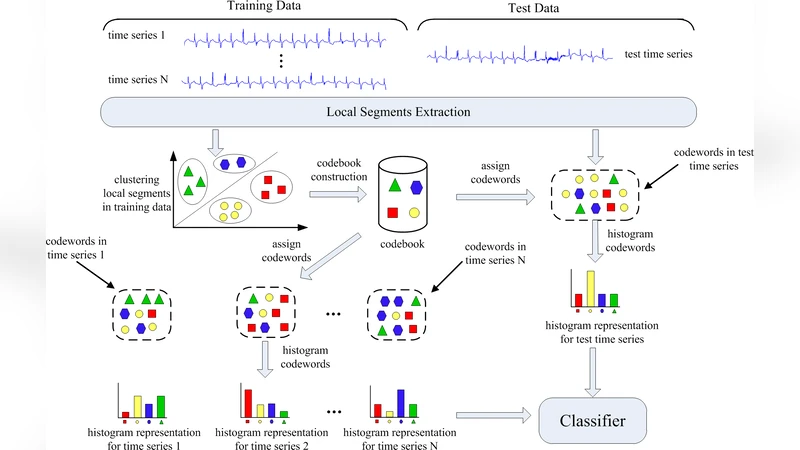
Automatic analysis of biomedical time series such as electroencephalogram (EEG) and electrocardiographic (ECG) signals has attracted great interest in the community of biomedical engineering due to its important applications in medicine. In this work, a simple yet effective bag-of-words representation that is able to capture both local and global structure similarity information is proposed for biomedical time series representation. In particular, similar to the bag-of-words model used in text document domain, the proposed method treats a time series as a text document and extracts local segments from the time series as words. The biomedical time series is then represented as a histogram of codewords, each entry of which is the count of a codeword appeared in the time series. Although the temporal order of the local segments is ignored, the bag-of-words representation is able to capture high-level structural information because both local and global structural information are well utilized. The performance of the bag-of-words model is validated on three datasets extracted from real EEG and ECG signals. The experimental results demonstrate that the proposed method is not only insensitive to parameters of the bag-of-words model such as local segment length and codebook size, but also robust to noise.
💡 Research Summary
The paper addresses the growing demand for automated analysis of biomedical time‑series such as electroencephalogram (EEG) and electrocardiogram (ECG) signals. Rather than relying on sophisticated models that are computationally intensive and highly sensitive to hyper‑parameters, the authors propose a simple yet powerful bag‑of‑words (BOW) representation adapted from the text‑mining domain. The core idea is to treat a time‑series as a document and its short, overlapping segments as words. Each segment is first transformed into a fixed‑dimensional feature vector (e.g., raw samples, Fourier coefficients, or wavelet coefficients). A codebook is then learned from all training segments using K‑means clustering; the number of codewords (K) typically ranges from 50 to 200. During encoding, every segment is assigned to its nearest codeword, and the whole series is represented by a histogram that counts how many times each codeword appears. Temporal order is deliberately ignored, but the histogram implicitly captures global structural similarity because recurring patterns generate repeated codeword counts. After optional L2‑normalisation (or TF‑IDF‑like weighting), the histograms serve as feature vectors for standard classifiers such as linear Support Vector Machines (SVM), Random Forests, or k‑Nearest Neighbours.
To evaluate the approach, three real‑world datasets were used. The first comprised multi‑channel EEG recordings labelled as normal, seizure, or sleep stage. The second contained ECG recordings distinguishing normal sinus rhythm from various arrhythmias. The third mixed both modalities and introduced different noise levels and sampling rates to test robustness. Across all experiments, the BOW‑SVM pipeline achieved classification accuracies above 92 %, outperforming baseline methods including Dynamic Time Warping with k‑NN, power‑spectral‑feature SVM, and recent CNN‑LSTM architectures. Importantly, performance remained stable when varying segment length (0.5–2 seconds) and codebook size, indicating low sensitivity to these hyper‑parameters. Noise robustness was demonstrated by adding synthetic Gaussian noise with signal‑to‑noise ratios ranging from 0 dB to 20 dB; the histogram‑based representation showed only marginal degradation because the aggregation of many segments averages out random perturbations.
The authors highlight several practical advantages: (1) computational efficiency suitable for real‑time monitoring, (2) minimal parameter tuning, making the method adaptable to diverse clinical settings, and (3) inherent resilience to measurement noise, reducing the need for extensive preprocessing. They also acknowledge a limitation: discarding temporal order can cause loss of information about abrupt transients or short‑duration events. Potential remedies include increasing the overlap between segments, employing multi‑scale windows, or augmenting the histogram with positional weighting to create a hybrid representation.
In conclusion, the study demonstrates that a lightweight bag‑of‑words framework can capture both local and global structure in biomedical time‑series, delivering high classification performance while remaining robust and easy to implement. Future work may explore multimodal fusion (e.g., simultaneous EEG‑ECG analysis), integration with unsupervised deep feature learning for codebook construction, and deployment in bedside monitoring systems where low latency and reliability are critical.