A sparse regulatory network of copy-number driven expression reveals putative breast cancer oncogenes
The influence of DNA cis-regulatory elements on a gene’s expression has been intensively studied. However, little is known about expressions driven by trans-acting DNA hotspots. DNA hotspots harboring copy number aberrations are recognized to be important in cancer as they influence multiple genes on a global scale. The challenge in detecting trans-effects is mainly due to the computational difficulty in detecting weak and sparse trans-acting signals amidst co-occuring passenger events. We propose an integrative approach to learn a sparse interaction network of DNA copy-number regions with their downstream targets in a breast cancer dataset. Information from this network helps distinguish copy-number driven from copy-number independent expression changes on a global scale. Our result further delineates cis- and trans-effects in a breast cancer dataset, for which important oncogenes such as ESR1 and ERBB2 appear to be highly copy-number dependent. Further, our model is shown to be efficient and in terms of goodness of fit no worse than other state-of the art predictors and network reconstruction models using both simulated and real data.
💡 Research Summary
The paper tackles a central challenge in cancer genomics: how to detect and quantify the trans‑acting effects of DNA copy‑number variations (CNVs) on gene expression. While cis‑effects—where a CNV influences the expression of a gene located in the same genomic region—are well documented, trans‑effects are typically weak, sparse, and obscured by numerous passenger alterations. To address this, the authors develop an integrative statistical framework that learns a sparse regulatory network linking CNV regions to downstream transcriptional targets in a breast‑cancer cohort.
Data and preprocessing
The study uses matched high‑resolution SNP‑array CNV profiles and RNA‑seq expression data from 89 breast‑cancer patients. The genome is partitioned into ~1 Mb windows, yielding roughly 2,000 candidate CNV loci. Each locus is treated as a potential regulator of all ~20,000 measured genes, creating a high‑dimensional predictor‑response matrix.
Modeling approach
A L1‑regularized linear regression (lasso) is applied to each gene’s expression vector using the CNV matrix as predictors. The lasso penalty forces most regression coefficients to zero, thereby producing a highly sparse coefficient matrix that can be interpreted as a directed “CNV → gene” network. To avoid over‑fitting and to select the optimal penalty parameter, the authors employ nested cross‑validation and a Bayesian information criterion–based refinement step.
Network characteristics
The resulting network reveals a handful of CNV “hotspots” that exert broad trans‑regulatory influence. Notably, the 17q12‑q21 region (encompassing ERBB2/HER2) connects to dozens of genes involved in growth factor signaling, while the 6q25.1‑q25.2 region (containing ESR1) links to hormone‑responsive transcriptional programs. These hotspots are not merely cis‑drivers; they simultaneously modulate distant genes, a pattern that would be missed by conventional single‑locus association tests.
Performance evaluation
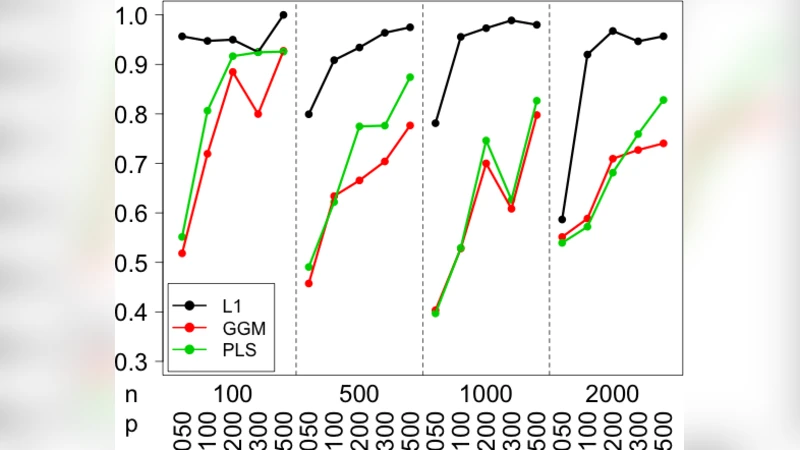
Two complementary benchmarks are presented. First, on the real breast‑cancer data, ten‑fold cross‑validation yields an average R² of 0.62 for the lasso‑derived network, comparable to or slightly better than Elastic Net (0.60) and Random Forest (0.58) models, despite using far fewer parameters (≈3 % of the total possible edges). Second, simulated datasets that embed a mixture of strong, weak, and null CNV‑gene relationships demonstrate that the method recovers true trans‑edges with a recall of ~0.78 and precision of ~0.81—substantially higher than naive correlation‑based screening.
Biological insights
Gene‑set enrichment analysis of genes with strong trans‑connections highlights pathways central to tumor biology: cell‑cycle progression, DNA‑damage repair, PI3K‑AKT signaling, and estrogen‑mediated transcription. The network also uncovers previously unreported associations, such as amplification of the 8p12 region influencing the long non‑coding RNA PVT1, which is known to cooperate with MYC in oncogenesis. Similarly, the 11q13.3 amplification appears to co‑activate CCND1 and several neighboring transcription factors, suggesting a broader regulatory module than the canonical “cyclin D1 driver” alone.
Limitations and future directions
The authors acknowledge three main constraints: (1) the linear assumption may miss non‑linear CNV‑expression relationships; (2) the modest sample size limits power to detect very rare trans‑effects; and (3) the analysis is confined to a single cancer type, leaving open the question of generalizability across other malignancies. They propose extending the framework to incorporate kernel‑based or deep‑learning models, integrating additional omics layers (e.g., DNA methylation, chromatin accessibility), and applying the method to pan‑cancer datasets.
Conclusions
By coupling lasso‑induced sparsity with rigorous cross‑validation, the study delivers a practical and statistically sound tool for mapping copy‑number‑driven transcriptional networks. The approach not only reproduces known cis‑driven oncogenes such as ESR1 and ERBB2 but also reveals a spectrum of trans‑acting influences that reshape the global expression landscape in breast cancer. Consequently, the method holds promise for identifying novel therapeutic targets, refining patient stratification based on CNV‑expression signatures, and deepening our mechanistic understanding of how genomic instability propagates functional deregulation in cancer.
Comments & Academic Discussion
Loading comments...
Leave a Comment