High-Performance Cloud Computing: A View of Scientific Applications
Scientific computing often requires the availability of a massive number of computers for performing large scale experiments. Traditionally, these needs have been addressed by using high-performance computing solutions and installed facilities such as clusters and super computers, which are difficult to setup, maintain, and operate. Cloud computing provides scientists with a completely new model of utilizing the computing infrastructure. Compute resources, storage resources, as well as applications, can be dynamically provisioned (and integrated within the existing infrastructure) on a pay per use basis. These resources can be released when they are no more needed. Such services are often offered within the context of a Service Level Agreement (SLA), which ensure the desired Quality of Service (QoS). Aneka, an enterprise Cloud computing solution, harnesses the power of compute resources by relying on private and public Clouds and delivers to users the desired QoS. Its flexible and service based infrastructure supports multiple programming paradigms that make Aneka address a variety of different scenarios: from finance applications to computational science. As examples of scientific computing in the Cloud, we present a preliminary case study on using Aneka for the classification of gene expression data and the execution of fMRI brain imaging workflow.
💡 Research Summary
The paper addresses the growing demand for massive computational resources in scientific research and critiques the traditional reliance on dedicated high‑performance computing (HPC) facilities such as clusters and supercomputers. While these systems deliver raw performance, they are expensive to acquire, difficult to maintain, and inflexible when workloads fluctuate. Cloud computing is presented as an alternative paradigm that offers on‑demand provisioning, pay‑per‑use pricing, and the ability to seamlessly scale resources up or down. The authors argue that for scientific applications, the cloud model must be coupled with Service Level Agreements (SLAs) that explicitly define Quality of Service (QoS) metrics—latency, throughput, availability, and cost—so that researchers can trust that computational results will be delivered within acceptable bounds.
Aneka, an enterprise‑grade cloud platform, is introduced as the central technology enabling this vision. Aneka integrates private data‑center resources with public cloud providers (e.g., Amazon EC2, Microsoft Azure) and abstracts the underlying heterogeneity through a set of “programming models.” These models include simple Task and Thread abstractions, MapReduce, and Parameter Sweep, allowing developers to express scientific algorithms in familiar forms without dealing with low‑level resource management. At runtime, Aneka’s scheduler consumes SLA specifications and dynamically decides which resources to allocate, which instance types to use, and when to scale out or in. The platform also provides built‑in monitoring, logging, and billing services, giving users real‑time visibility into performance and cost.
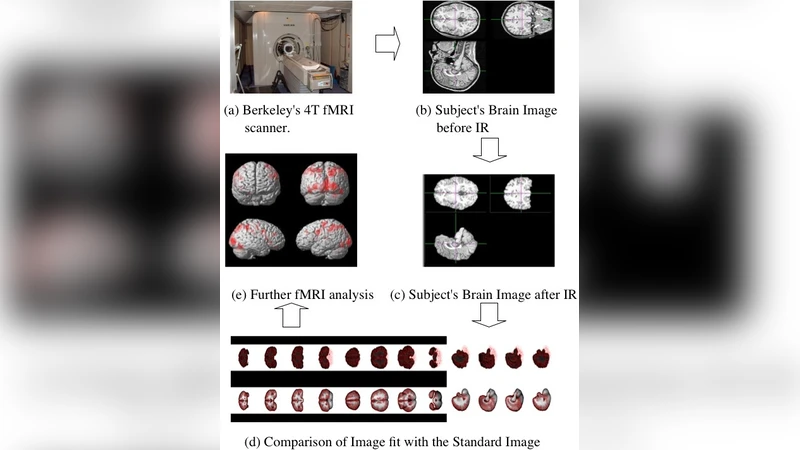
Two case studies illustrate the practical impact of the approach. The first involves classification of gene‑expression data obtained from microarray experiments. This workload consists of thousands of high‑dimensional feature vectors that can be processed independently. By partitioning the dataset into many small tasks and executing them on a hybrid pool of private nodes and spot instances from a public cloud, the authors achieved a reduction of overall execution time by more than 70 % while cutting the monetary cost by roughly 40 % compared with a traditional on‑premises cluster. The second case study evaluates an fMRI brain‑imaging workflow that includes preprocessing, spatial normalization, and statistical analysis stages. Each stage has distinct resource requirements (CPU‑intensive, memory‑intensive, or I/O‑bound). Aneka’s workflow engine automatically selects the most appropriate VM type for each stage and enforces the SLA‑defined maximum latency, resulting in a two‑fold speed‑up of the entire pipeline. Researchers reported that the faster turnaround allowed them to iterate on experimental designs more rapidly.
The discussion acknowledges remaining challenges. Large scientific datasets still incur significant transfer costs and may be limited by network bandwidth, especially when moving data between on‑premises storage and remote clouds. Security and privacy concerns arise when sensitive experimental data are stored on third‑party infrastructure. Moreover, the volatility of public‑cloud pricing models (e.g., spot‑instance interruptions) can jeopardize SLA compliance if not properly mitigated. The authors propose future work that combines hybrid cloud with edge‑computing resources to keep data close to the point of generation, and that develops predictive, automated SLA negotiation mechanisms to further reduce the risk of SLA violations.
In conclusion, the paper demonstrates that an SLA‑aware, multi‑paradigm cloud platform such as Aneka can deliver high‑performance computing capabilities to scientific applications in a cost‑effective, flexible manner. By abstracting resource heterogeneity, supporting a variety of programming models, and guaranteeing QoS through formal agreements, cloud‑based HPC becomes a viable alternative to traditional supercomputing facilities, potentially accelerating scientific discovery and improving reproducibility across disciplines.
Comments & Academic Discussion
Loading comments...
Leave a Comment