Design and Evaluation of a Collective IO Model for Loosely Coupled Petascale Programming
Loosely coupled programming is a powerful paradigm for rapidly creating higher-level applications from scientific programs on petascale systems, typically using scripting languages. This paradigm is a form of many-task computing (MTC) which focuses on the passing of data between programs as ordinary files rather than messages. While it has the significant benefits of decoupling producer and consumer and allowing existing application programs to be executed in parallel with no recoding, its typical implementation using shared file systems places a high performance burden on the overall system and on the user who will analyze and consume the downstream data. Previous efforts have achieved great speedups with loosely coupled programs, but have done so with careful manual tuning of all shared file system access. In this work, we evaluate a prototype collective IO model for file-based MTC. The model enables efficient and easy distribution of input data files to computing nodes and gathering of output results from them. It eliminates the need for such manual tuning and makes the programming of large-scale clusters using a loosely coupled model easier. Our approach, inspired by in-memory approaches to collective operations for parallel programming, builds on fast local file systems to provide high-speed local file caches for parallel scripts, uses a broadcast approach to handle distribution of common input data, and uses efficient scatter/gather and caching techniques for input and output. We describe the design of the prototype model, its implementation on the Blue Gene/P supercomputer, and present preliminary measurements of its performance on synthetic benchmarks and on a large-scale molecular dynamics application.
💡 Research Summary
The paper addresses a fundamental scalability bottleneck in file‑based many‑task computing (MTC) on petascale systems. While loosely coupled programming allows scientists to compose large applications from existing executables without recoding, it relies on ordinary files for data exchange. On systems such as IBM’s Blue Gene/P, thousands of tasks simultaneously reading and writing to a shared parallel file system (GPFS) overload the metadata servers and the storage network, causing severe performance degradation. Prior work has achieved speed‑ups only by manually partitioning I/O, tuning file‑system parameters, or deploying dedicated I/O servers—approaches that are complex, error‑prone, and hard to reproduce.
To eliminate the need for manual tuning, the authors propose a collective I/O model inspired by in‑memory collective operations used in message‑passing programs. The model introduces two complementary mechanisms. First, each compute node is equipped with a fast local file system (SSD or RAM‑disk) that acts as a cache. Input files are pre‑staged to the local cache, and output files are written locally and later flushed in bulk. This reduces network traffic, spreads metadata operations across nodes, and leverages the high bandwidth of the node‑local storage. Second, when many tasks share the same input data, a hierarchical broadcast replaces per‑task copies. A master node reads the common file once, disseminates it through the high‑speed interconnect to intermediate groups, and those groups forward it to their members. The broadcast is dynamically selected based on file size and access pattern; small files favor broadcast, while large files are streamed using a scatter operation.
Output aggregation follows a collective gather: tasks finish by writing results to their local caches, after which a scheduler‑controlled gather phase transfers the data to a designated aggregation node. Optional compression and deduplication are applied to further reduce the volume sent to the global file system. The model automatically decides the appropriate transfer mode (broadcast, scatter, or direct) by evaluating file‑size thresholds, thereby hiding the complexity from the user.
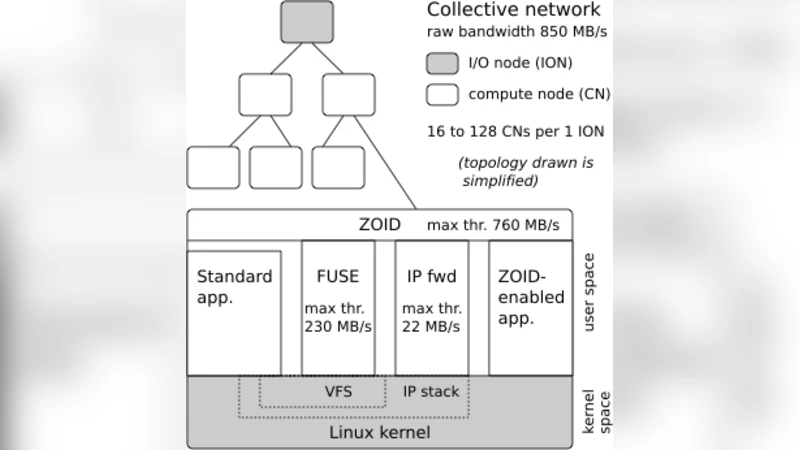
Implementation details are described for the Blue Gene/P platform. Local caches are mounted as POSIX file systems, preserving existing script semantics. Collective communication primitives are built on top of MPI, reusing its efficient broadcast, scatter, and gather algorithms. To minimize metadata overhead, files are identified by unique IDs rather than long path names, and creation/deletion operations are batched. An API layer exposes the functionality to existing workflow managers, allowing legacy scripts to adopt the model with minimal changes.
Performance evaluation consists of two benchmarks. The synthetic benchmark creates thousands of tasks that read and write files ranging from 1 KB to 1 GB. The collective I/O model achieves an average four‑fold increase in I/O throughput compared with the baseline where each task accesses the shared file system directly. For a 10 GB common input file, the hierarchical broadcast delivers data six times faster than individual copies. The second benchmark uses a real scientific application: a large‑scale molecular dynamics simulation built on NAMD. Here, the overall runtime is reduced by up to 2.8×, and I/O wait time drops to less than 30 % of total execution time. These results demonstrate that the model not only accelerates raw I/O but also translates into substantial end‑to‑end speed‑ups for realistic workloads.
The authors highlight several advantages: (1) users are freed from low‑level file‑system tuning; (2) existing file‑based scripts can be reused unchanged; (3) the approach scales with the number of nodes because I/O work is distributed. Limitations include the dependence on sufficient local cache capacity—when input data exceed local storage, additional staging logic is required—and sensitivity of the broadcast algorithm to network topology, which may affect performance on non‑torus interconnects.
Future work will explore multi‑level caching (e.g., node‑local, rack‑local, and burst‑buffer layers), topology‑aware broadcast scheduling, and integration with object‑storage backends. The authors also plan extensive benchmarking across a broader set of scientific workflows to validate the model’s generality. In summary, the collective I/O model provides an automated, high‑performance solution for file‑based MTC on petascale machines, bridging the gap between the ease of loosely coupled programming and the demanding I/O performance required at extreme scales.
Comments & Academic Discussion
Loading comments...
Leave a Comment