Stochastic switching between multistable oscillation patterns of the Min-system
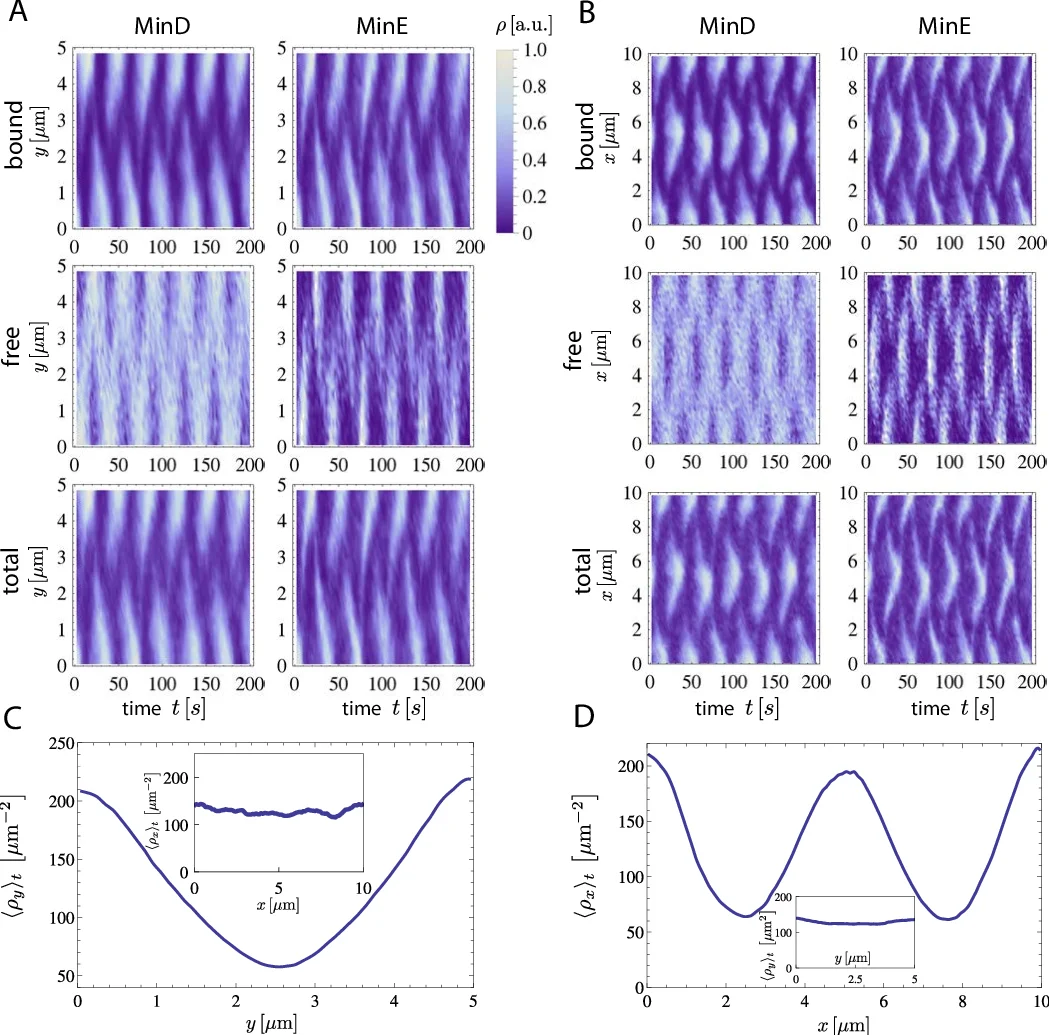
The spatiotemporal oscillation patterns of the proteins MinD and MinE are used by the bacterium E. coli to sense its own geometry. Strikingly, both computer simulations and experiments have recently shown that for the same geometry of the reaction volume, different oscillation patterns can be stable, with stochastic switching between them. Here we use particle-based Brownian dynamics simulations to predict the relative frequency of different oscillation patterns over a large range of three-dimensional compartment geometries, in excellent agreement with experimental results. Fourier analyses as well as pattern recognition algorithms are used to automatically identify the different oscillation patterns and the switching rates between them. We also identify novel oscillation patterns in three-dimensional compartments with membrane-covered walls and identify a linear relation between the bound Min-protein densities and the volume-to-surface ratio. In general, our work shows how geometry sensing is limited by multistability and stochastic fluctuations.
💡 Research Summary
The paper investigates how the Min protein system of Escherichia coli—a well‑studied reaction‑diffusion oscillator that determines the cell‑division site—behaves in three‑dimensional micro‑compartments of various shapes. While previous experiments showed that the same geometry can support several distinct, stable oscillation modes (pole‑to‑pole, transverse pole‑to‑pole, strip‑like, etc.) and occasional stochastic switching between them, quantitative data on mode frequencies and switching kinetics were lacking.
To address this gap, the authors employ particle‑based Brownian dynamics simulations that implement the “minimal” Min model originally proposed by Huang et al. The model includes five elementary reactions: (1a) membrane attachment of cytosolic MinD‑ATP, (1b) cooperative recruitment of additional MinD‑ATP to already bound MinD, (1c) recruitment of cytosolic MinE to membrane‑bound MinD, (1d) ATP‑hydrolysis‑driven release of a MinDE complex, and (1e) nucleotide exchange converting MinD‑ADP back to MinD‑ATP. Three parameter sets (A, B, C) derived from earlier literature are tested to assess robustness.
Simulation details: a fixed time step Δt = 10⁻⁴ s is used; free particles diffuse in three dimensions with D≈2.5 µm² s⁻¹, while membrane‑bound particles diffuse laterally with D_bound≈0.01 µm² s⁻¹. First‑order reactions are treated as Poisson processes, membrane‑attachment reactions use a proximity‑based rate (particles within 0.02 µm of the membrane attempt attachment with probability p = 1 − exp(−σ d Δt)), and second‑order reactions follow the Smoluchowski framework as implemented in the Smoldyn package. Reflective boundary conditions are applied at compartment walls.
Three geometric configurations are examined: Geometry A (only the bottom surface coated with a lipid membrane, mimicking flat patterned substrates), Geometry B (bottom plus side walls coated, reproducing open‑top micro‑chambers), and Geometry C (all six faces coated, representing fully sculpted cells).
Key findings:
-
Multistability of Oscillation Patterns – All three geometries support at least three distinct, long‑lived modes. Geometry A predominantly shows classic pole‑to‑pole oscillations, but occasional transverse pole‑to‑pole and strip‑like modes appear. Geometry B exhibits a mixture of longitudinal and transverse strip‑like waves, while Geometry C reveals a novel “edge‑localized” pattern where MinD concentrates along the compartment edges.
-
Geometry‑Dependent Mode Frequencies – The relative occurrence of each mode correlates strongly with the volume‑to‑surface ratio (V/S) and the fraction of membrane coverage. Small V/S favors pole‑to‑pole, whereas larger V/S promotes strip‑like and edge‑localized modes. Quantitatively, in Geometry C the edge‑localized mode occupies >30 % of the simulation time when V/S > 0.8 µm⁻¹.
-
Stochastic Switching Kinetics – By performing Fourier transforms on the time‑series of membrane‑bound protein densities and applying a custom pattern‑recognition pipeline (feature extraction + K‑means clustering), the authors automatically detect switching events. The mean residence time in a given mode is ~120 s (standard deviation ≈ 45 s), yielding switching rates that match experimental measurements from micro‑fabricated chambers. Parameter set B (higher attachment rates) accelerates switching by ~30 %, confirming the sensitivity of multistability to kinetic parameters.
-
Linear Relation Between Bound Protein Density and V/S – The average surface density of membrane‑bound MinD and MinE (ρ_bound) scales linearly with the volume‑to‑surface ratio across all geometries and parameter sets: ρ_bound ≈ α·(V/S) + β, with α≈0.45 µM·µm and β≈0.12 µM. This suggests a simple physical mechanism by which a cell could infer its own geometry from the concentration of bound Min proteins.
The study validates the particle‑based approach by reproducing experimental mode frequencies and, for the first time, providing quantitative switching times. It also uncovers previously unobserved patterns in fully membrane‑coated compartments and demonstrates that the minimal reaction set already contains all ingredients necessary for geometry sensing. The authors make their simulation code and analysis scripts publicly available, enabling future work on synthetic cell design, microfluidic pattern control, and stochastic bifurcations in other reaction‑diffusion systems.
In conclusion, the paper shows that multistability and stochastic fluctuations are intrinsic to the Min system’s ability to sense cellular geometry, and that particle‑level stochastic modeling is essential for capturing these phenomena. This insight bridges the gap between deterministic PDE descriptions and the noisy reality of intracellular protein dynamics, opening avenues for engineering robust spatial patterning in synthetic biology.
Comments & Academic Discussion
Loading comments...
Leave a Comment