Noise enhanced persistence in a biochemical regulatory network with feedback control
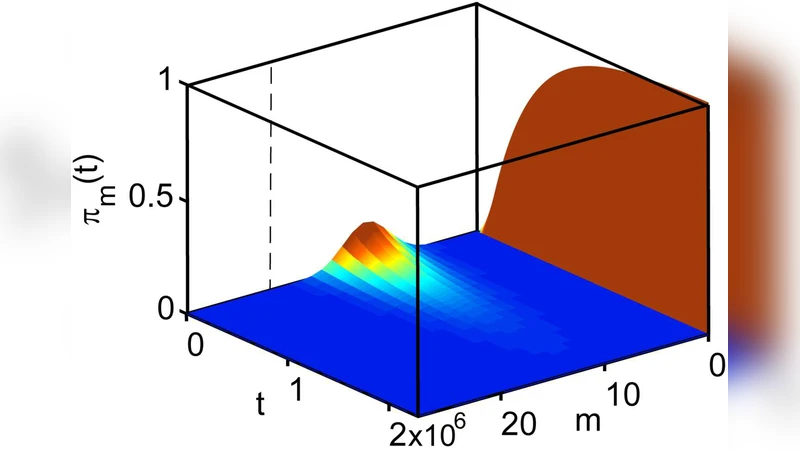
We find that discrete noise of inhibiting (signal) molecules can greatly delay the extinction of plasmids in a plasmid replication system: a prototypical biochemical regulatory network. We calculate the probability distribution of the metastable state of the plasmids and show on this example that the reaction rate equations may fail in predicting the average number of regulated molecules even when this number is large, and the time is much shorter than the mean extinction time.
💡 Research Summary
The paper investigates how discrete stochastic fluctuations of inhibitory signal molecules can dramatically prolong the survival of plasmids in a prototypical plasmid replication system, a classic example of a biochemical regulatory network with negative feedback control. The authors begin by formulating a master equation that treats both plasmid copies and inhibitor molecules as integer‑valued stochastic variables. Reaction rates are defined according to the underlying biochemical steps: plasmid replication is up‑regulated when inhibitor concentration is low and down‑regulated when inhibitors bind to the replication machinery. The deterministic counterpart, the reaction‑rate equations (RRE), predicts a steady‑state plasmid number that depends only on average concentrations. However, when the number of inhibitor molecules is small, the system experiences pronounced shot noise, and the deterministic description can become misleading.
To capture the effect of this noise, the authors employ a large‑deviation (WKB) approximation to the master equation, deriving an effective Hamiltonian and a corresponding “potential” landscape. The minima of this potential correspond to metastable (quasi‑steady) states where plasmids persist for long periods before an eventual stochastic extinction event. By solving the Hamilton‑Jacobi equations, they obtain an explicit expression for the probability distribution of the metastable plasmid number and an analytical estimate of the mean extinction time. Their calculations reveal two striking phenomena. First, the discrete noise of the inhibitors creates intermittent windows in which the inhibitory pressure is temporarily weakened, allowing plasmid replication to surge and thereby delaying extinction far beyond the deterministic prediction. Second, even when the average plasmid count is large (hundreds of copies) and the observation time is much shorter than the mean extinction time, the RRE underestimates the true average plasmid number because the probability distribution is skewed by noise. In other words, the “law of large numbers” does not guarantee convergence to the deterministic value in this regime.
A systematic parameter sweep shows how the feedback strength, the synthesis and degradation rates of the inhibitor, and the maximal replication rate shape the metastable distribution. Stronger feedback pushes the metastable peak to lower plasmid numbers but also amplifies the sensitivity to stochastic fluctuations, while faster inhibitor turnover reduces noise and brings the stochastic average closer to the deterministic one, yet a residual discrepancy remains. The authors also discuss the biological implications: in synthetic biology, intentional modulation of inhibitor noise could be used to engineer circuits with desired persistence properties; in natural contexts, the same mechanism may help explain why antibiotic‑resistance plasmids can persist in bacterial populations far longer than would be expected from simple deterministic models.
Overall, the study provides a rigorous quantitative framework that demonstrates the failure of reaction‑rate equations to predict average molecule numbers and extinction dynamics in regimes where discrete noise is significant. By combining master‑equation analysis, large‑deviation theory, and explicit calculation of metastable distributions, the work highlights the crucial role of stochasticity in biochemical networks with feedback control and offers practical insights for both theoretical modeling and experimental design.
Comments & Academic Discussion
Loading comments...
Leave a Comment