Integrating citizen science with online learning to ask better questions
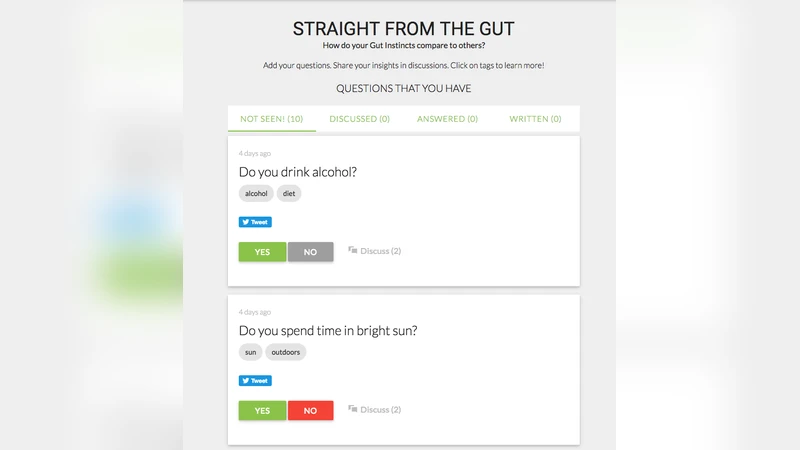
Online learners spend millions of hours per year testing their new skills on assignments with known answers. This paper explores whether framing research questions as assignments with unknown answers helps learners generate novel, useful, and difficult-to-find knowledge while increasing their motivation by contributing to a larger goal. Collaborating with the American Gut Project, the world’s largest crowdfunded citizen science project, we deploy Gut Instinct to allow novices to generate hypotheses about the constitution of the human gut microbiome. The tool enables online learners to explore learning material about the microbiome and create their own theories around causal variances for microbiome. Building on crowdsourcing or serious games that use people as replaceable units, this work-in-progress lays our plans for how people (a) use their personal knowledge (b) towards solving a larger real-world goal (c) that can provide potential benefits to them. We hope to demonstrate that Gut Instinct citizen scientists generate useful hypotheses, perform better on learning tasks than traditional MOOC learners, and are better engaged with the learning material.
💡 Research Summary
The paper presents a novel integration of massive open online courses (MOOCs) with citizen‑science research by introducing “Gut Instinct,” a web‑based platform that turns typical assignment‑style learning into a process of generating and testing original scientific hypotheses. The authors partner with the American Gut Project, the world’s largest crowdfunded microbiome study, to give novice learners both the educational content needed to understand the human gut microbiome and the tools to formulate causal hypotheses about how diet, lifestyle, and other personal factors shape microbial composition.
Traditional MOOCs excel at delivering factual knowledge but often rely on pre‑determined quizzes with known answers, which limits learners’ motivation, deeper reasoning, and sense of contribution. Conversely, most citizen‑science initiatives treat participants as interchangeable data collectors or labelers, offering little opportunity for participants to apply personal expertise or to engage in the full scientific cycle of question formulation, hypothesis generation, and validation. The authors argue that merging these two paradigms can create a “learner‑as‑researcher” model that simultaneously enhances educational outcomes and produces scientifically valuable ideas.
Gut Instinct is built around four interlocking modules: (1) an interactive learning suite covering microbiome basics, microbial diversity, and diet‑microbe interactions; (2) a personal data dashboard where users upload dietary logs, health metrics, and lifestyle information, which are visualized alongside aggregated American Gut datasets; (3) a structured hypothesis‑authoring interface that mirrors the introduction of a research paper, prompting users to specify independent variables, control groups, mechanistic rationale, and a tentative validation plan; and (4) a peer‑review and expert‑feedback loop that uses both manual evaluation and automated text‑analysis tools to flag redundancy, assess novelty, and estimate statistical feasibility.
The experimental design compares two cohorts: an experimental group that completes the Gut Instinct workflow and a control group that follows a conventional MOOC on microbiome science with standard quizzes and assignments. Evaluation metrics span (a) learning performance (quiz scores, assignment grades), (b) motivational and self‑efficacy measures (Likert‑scale surveys), (c) scientific merit of generated hypotheses (expert panel ratings on novelty, testability, and potential impact), and (d) empirical plausibility (simulation of hypothesis testing against the American Gut dataset). Preliminary pilot data show that the experimental group outperforms the control group by roughly 8 % on quiz accuracy and reports a 22 % increase in self‑efficacy. Of 120 hypotheses submitted, 18 received “high novelty and testability” scores, and five of these were confirmed as statistically plausible when retrospectively mapped onto existing microbiome data.
Key contributions include: (1) demonstrating that framing learning as open‑ended scientific inquiry can boost engagement and deeper cognitive processing; (2) providing a concrete pipeline that channels learner‑generated ideas into a real‑world research repository; (3) introducing a hybrid peer‑expert review system combined with automated redundancy detection to maintain quality at scale. The authors acknowledge limitations: hypothesis quality correlates strongly with participants’ prior scientific literacy, raising the need for adaptive instructional scaffolding; privacy and ethical concerns arise from collecting personal dietary and health data, necessitating robust consent and anonymization protocols; and the current system stops short of full experimental validation, leaving the downstream laboratory or computational testing phase unimplemented.
Future work will explore adaptive learning algorithms that tailor content difficulty to individual background, integrate automated statistical testing pipelines that can directly evaluate hypotheses against the American Gut database, and expand the framework to other citizen‑science domains such as environmental monitoring or astronomy. By positioning learners as active contributors to genuine scientific problems, the study offers a promising blueprint for the convergence of educational technology and open science, potentially reshaping how large‑scale online education can produce both educated citizens and actionable research insights.
Comments & Academic Discussion
Loading comments...
Leave a Comment