Scalable Algorithms for Generating and Analyzing Structural Brain Networks with a Varying Number of Nodes
Diffusion Magnetic Resonance Imaging (MRI) exploits the anisotropic diffusion of water molecules in the brain to enable the estimation of the brain’s anatomical fiber tracts at a relatively high resolution. In particular, tractographic methods can be used to generate whole-brain anatomical connectivity matrix where each element provides an estimate of the connectivity strength between the corresponding voxels. Structural brain networks are built using the connectivity information and a predefined brain parcellation, where the nodes of the network represent the brain regions and the edge weights capture the connectivity strengths between the corresponding brain regions. This paper introduces a number of novel scalable methods to generate and analyze structural brain networks with a varying number of nodes. In particular, we introduce a new parallel algorithm to quickly generate large scale connectivity-based parcellations for which voxels in a region possess highly similar connectivity patterns to the rest of the regions. We show that the corresponding regional structural consistency is always superior to randomly generated parcellations over a wide range of parcellation sizes. Corresponding brain networks with a varying number of nodes are analyzed using standard graph-theorectic measures, as well as, new measures derived from spectral graph theory. Our results indicate increasingly more statistical power of brain networks with larger numbers of nodes and the relatively unique shape of the spectral profile of large brain networks relative to other well-known networks.
💡 Research Summary
The paper presents a scalable, GPU‑accelerated pipeline for generating connectivity‑based brain parcellations and for analyzing the resulting structural brain networks across a wide range of node counts. Starting from high‑resolution diffusion MRI data (NKI/Rockland dataset), the authors reconstruct diffusion tensors for each voxel and apply probabilistic tractography (BEDPOSTX and probtrackx) to obtain a massive weighted connectivity matrix of size 155,794 × 155,794, where each entry reflects the number of streamlines linking a pair of voxels.
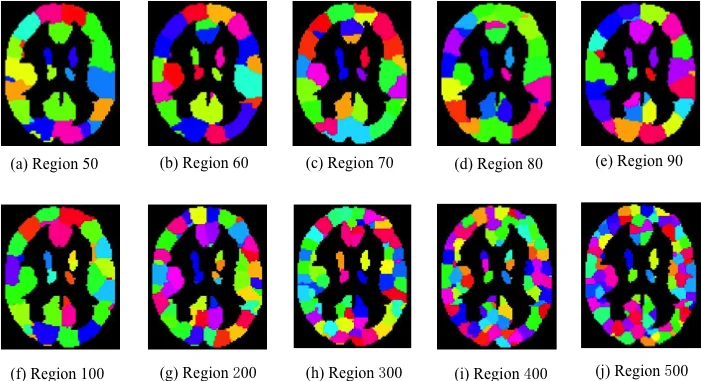
The core methodological contribution is a fast parallel spectral clustering algorithm that creates whole‑brain parcellations with up to 500 regions. Each voxel is represented by a “connectivity profile” – a vector of connection strengths to all other voxels – and spatial adjacency is enforced by linking voxels within a radius r = 2 voxels. The resulting spatial‑similarity graph is processed on a heterogeneous CPU‑GPU platform (Intel Xeon E5‑2690 + NVIDIA Tesla K20c) using CUDA, cuBLAS, cuSPARSE, Thrust, and ARPACK‑CUDA. By leveraging these libraries, the algorithm solves large sparse eigenvalue problems (implicitly restarted Arnoldi) in seconds, achieving an order‑of‑magnitude speed‑up over CPU‑only implementations.
The generated parcellations are evaluated against randomly generated ones. Across all tested numbers of parcels (k = 2 … 500), the connectivity‑based parcellations exhibit significantly higher intra‑parcel structural consistency, confirming that voxels grouped together share highly similar connectivity patterns.
Using these parcellations, the authors construct structural brain networks where nodes correspond to parcels and edge weights are the summed streamline counts between parcels. They then compute a suite of traditional graph‑theoretic metrics (characteristic path length, clustering coefficient, global/local efficiency, betweenness centrality, modularity) and a set of spectral‑graph metrics (Laplacian eigenvalue distribution, spectral gap, Fiedler vector, shape of the spectrum).
Key findings include:
-
Node‑count effect – As the number of nodes increases, networks display stronger small‑world characteristics, higher modularity, and greater statistical power for detecting group differences (e.g., age or disease cohorts).
-
Spectral uniqueness – Large‑scale brain networks possess a distinctive eigenvalue spectrum that differs markedly from other well‑studied complex networks (social, technological). The high‑frequency tail of the spectrum captures subtle structural nuances that are invisible to binary‑thresholded graph measures.
-
Robustness to thresholding – Spectral analyses operate directly on the original weighted graph, avoiding the information loss inherent in thresholding and binarization. Consequently, spectral metrics are less sensitive to arbitrary sparsity choices and provide more stable biomarkers.
-
Computational efficiency – The heterogeneous implementation reduces the time required for parcellation generation and eigenvalue computation from hours to minutes, enabling routine processing of large cohorts such as the Human Connectome Project.
The authors argue that the combination of connectivity‑driven parcellation and spectral network analysis offers a powerful framework for future neuroimaging studies, particularly for identifying biomarkers of neurological and psychiatric disorders and for probing developmental trajectories. The work also sets the stage for standardized, high‑throughput brain network pipelines that can be deployed across research sites worldwide.
Comments & Academic Discussion
Loading comments...
Leave a Comment