Combining predictors of natively unfolded proteins to detect a twilight zone between order and disorder in generic datasets
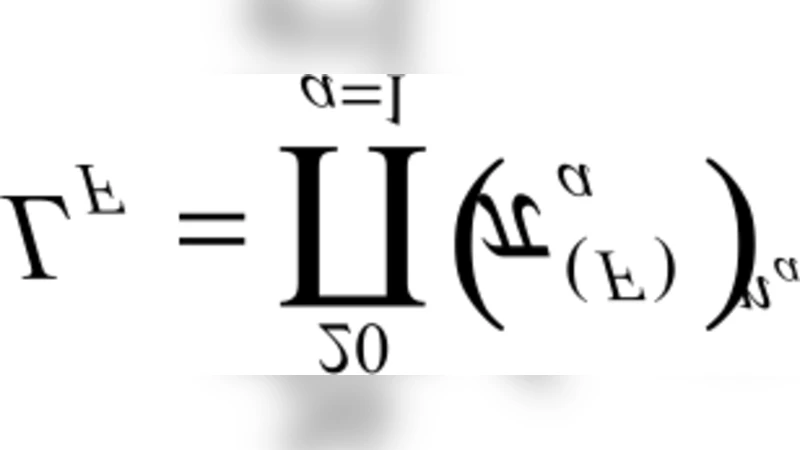
Natively unfolded proteins lack a well defined three dimensional structure but have important biological functions, suggesting a re-assignment of the structure-function paradigm. Many proteins have amino acidic compositions compatible both with the folded and unfolded status, and belong to a twilight zone between order and disorder. This makes difficult a dichotomic classification of protein sequences into folded and natively unfolded ones. In this methodological paper dichotomic folding indexes are considered: hydrophobicity-charge, mean packing, mean pairwise energy, Poodle-W and a new global index, that is called here gVSL2, based on the local disorder predictor VSL2. The performance of these indexes is evaluated on different datasets. Poodle-W, gVSL2 and mean pairwise energy have good performance and stability in all the datasets considered and are combined into a strictly unanimous combination score SSU, that leaves proteins unclassified when the consensus of all combined indexes is not reached. The unclassified proteins: i) belong to an overlap region in the vector space of amino acidic compositions occupied by both folded and unfolded proteins; ii) are composed by approximately the same number of order-promoting and disorder-promoting amino acids; iii) have a mean flexibility intermediate between that of folded and that of unfolded proteins. These proteins reasonably have physical properties intermediate between those of folded and those of natively unfolded proteins and their structural properties and evolutionary history are worth to be investigated.
💡 Research Summary
The paper addresses the challenge of classifying protein sequences into folded or natively unfolded categories when many sequences possess amino‑acid compositions compatible with both states. Such sequences occupy a “twilight zone” where a simple dichotomous label is ambiguous. The authors evaluate five global folding indexes: hydrophobicity‑charge, mean packing, mean pairwise energy, Poodle‑W, and a newly defined index called gVSL2, which aggregates the local disorder predictor VSL2 into a single sequence‑level score. Using several benchmark datasets that include clearly folded, clearly unfolded, and mixed‑type proteins, each index is assessed for accuracy, sensitivity, specificity, and area under the ROC curve. Hydrophobicity‑charge and mean packing show variable performance across datasets, whereas Poodle‑W, gVSL2, and mean pairwise energy consistently achieve high AUC values (>0.90) and low false‑positive rates.
To exploit the complementary strengths of the three robust indexes, the authors introduce a Strictly Unanimous Score (SSU). SSU assigns a protein to the folded or unfolded class only when all three indexes agree; otherwise the protein is left unclassified. This conservative strategy eliminates many borderline cases and results in roughly 10 % of proteins being marked as “unclassified.” The unclassified set is then examined in detail. In the multidimensional space of amino‑acid composition, these proteins lie in the overlap region between the folded and unfolded clusters. They contain nearly equal numbers of order‑promoting (e.g., Cys, Trp, Phe) and disorder‑promoting residues (e.g., Arg, Lys, Gln), and their average flexibility scores are intermediate between the two groups. Consequently, they exhibit physical properties that are midway between fully structured and intrinsically disordered proteins.
The paper argues that such proteins likely represent conditionally disordered or “semi‑folded” states that can shift toward order or disorder in response to environmental cues, post‑translational modifications, or binding events. The gVSL2 index is highlighted as a useful bridge between local disorder predictions (VSL2 provides per‑residue disorder probabilities) and global classification, by summarizing the per‑residue scores into a single metric. Mean pairwise energy, derived from statistical potentials of inter‑residue interactions, adds a physics‑based perspective to the otherwise statistical classifiers.
Overall, the study demonstrates that combining multiple, independently validated folding indexes under a strict consensus rule yields a reliable method for detecting proteins that reside in the twilight zone. The SSU framework not only improves classification confidence for clearly folded or unfolded proteins but also systematically isolates a biologically interesting subset that warrants further experimental investigation. Future work on structural determination, functional assays, and evolutionary analysis of these intermediate‑property proteins could illuminate new aspects of the relationship between disorder and function in the proteome.
Comments & Academic Discussion
Loading comments...
Leave a Comment