Bayesian Optimization with Dimension Scheduling: Application to Biological Systems
Bayesian Optimization (BO) is a data-efficient method for global black-box optimization of an expensive-to-evaluate fitness function. BO typically assumes that computation cost of BO is cheap, but experiments are time consuming or costly. In practice…
Authors: Doniyor Ulmasov, Caroline Baroukh, Benoit Chachuat
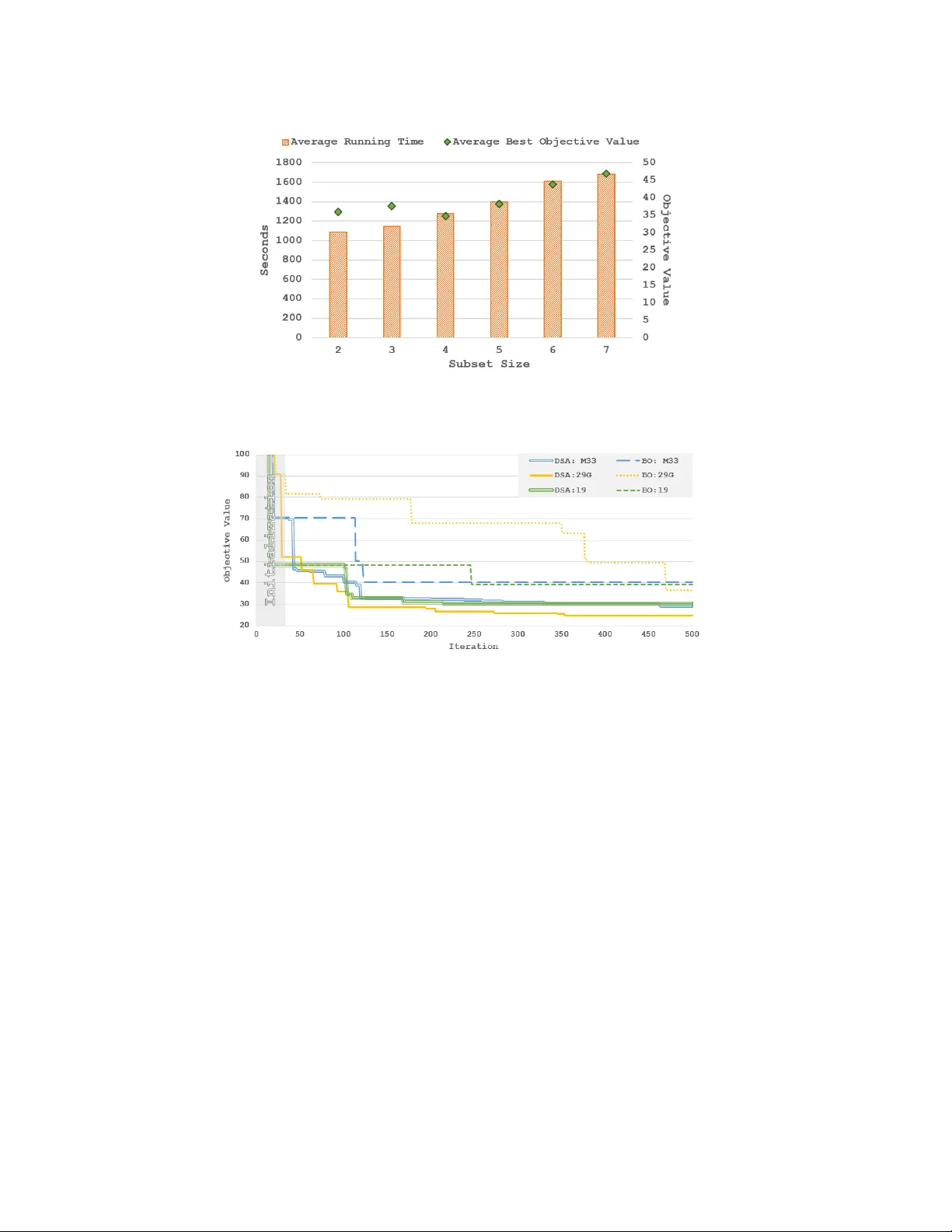
Bayesian Optimization with Dimension Scheduling: A pplication to Biological Systems Doniyor Ulmasov a , Caroline Baroukh b , Benoit Chachuat c , Marc Peter Deisenroth a,* and Ruth Misener a,* a Department of Computing, Imperial Colle ge London, UK b INRA, Laboratory of En vir onmental Biotec hnology , F rance c Department of Chemical Engineering, Imperial Colle ge London, UK *m.deisenr oth, r .misener@imperial.ac.uk; +44 (0) 20759 48315 Nov ember 18, 2015 Abstract Bayesian Optimization (BO) is a data-efficient method for global black-box optimization of an expensi ve-to-e valuate fitness function. BO typically assumes that computation cost of BO is cheap, but experiments are time consuming or costly . In practice, this allo ws us to optimize ten or fewer critical parameters in up to 1,000 e xperiments. But experiments may be less e xpensiv e than BO methods assume: In some simulation models, we may be able to conduct multiple thousands of experiments in a few hours, and the computational burden of BO is no longer negligible compared to experimentation time. T o address this challenge we introduce a new Dimension Scheduling Algorithm (DSA), which reduces the computational burden of BO for many experiments. The key idea is that DSA optimizes the fitness function only along a small set of dimensions at each iteration. This DSA strategy (1) reduces the necessary computation time, (2) finds good solutions faster than the traditional BO method, and (3) can be parallelized straightforwardly . W e ev aluate the DSA in the context of optimizing parameters of dynamic models of microalgae metabolism and show f aster con vergence than traditional BO. Keyw ords: Bayesian Optimization, Black-Box Optimization, Parameter Estimation, Microalgae Metabolism 1. Intr oduction Bayesian Optimization (BO) is a data-efficient method for global black-box optimization of an expensi ve-to-e valuate fitness function. The working hypothesis of BO is that experiments are very expensi ve, e.g., in terms of time or money , while BO computations are relatively cheap. BO has been applied to a wide variety of problems, including tuning critical parameters of Deep Neural Networks (Dahl et al., 2013), learning controller parameters of walking robots (Calandra et al., 2015) or automatic algorithm configuration (Hutter et al., 2011; Snoek et al., 2012). T o balance exploration and exploitation during optimization BO uses Gaussian Processes (GPs) to model a posterior distrib ution over fitness functions from av ailable experiments (Jones et al., 1998). Similar to experimental design, an acquisition function is applied to the GP posterior ov er fitness functions to suggest the next (optimal) e xperiment. In all applications, there is a common theme: BO perfor- mance de grades in high dimensions and/or with large data amount of observ ation points due to the scaling of the Gaussian Processes (GP), the surrogate functions. In practice, BO is limited to opti- mizing about ten parameters and about 1,000 experiments since GP predictions/ev aluations scale linearly in the number of dimensions but cubically in the number data points when we integrate 2 Ulmasov , Bar oukh, Chachuat, Deisenr oth, Misener out the hyperparameters. Dynamic models of biological processes allow us to test biological hypotheses while running fewer costly , real-world experiments (Floudas and Pardalos, 2000). This paper considers estimating biological parameters (e.g., reaction rate kinetics) by minimizing the squared error between model and experimental data points (Rodrigues-Fernandez et al., 2006). W e propose BO for efficient parameter estimation of a dynamic microalgae metabolism model (Baroukh et al., 2014). The forcing function is based on light e xposure and nitrate input; experimental data has been collected for measurable outputs including lipids, carbohydrates, carbon organic biomass, nitrogen organic biomass and chlorophyll. But our method is general and may be applied to an y process model. There are several timescales for collecting microalgae metabolism data: an experiment of Lacour et al. (2012) may tak e 10 days while each model simulation of Baroukh et al. (2014) runs in a frac- tion of a second. BO is traditionally applied to functions with an expensiv e ev aluation costs, e.g., running a 10 day experiment, but the objecti ve of this paper is testing biological hypotheses; we are specifically interested in running the simulation model many times for parameter estimation. Parameter estimation for dynamic systems has been alternativ ely proposed using deterministic global optimization (Michalik et al., 2009), heuristics (Rodrigues-Fernandez and Banga, 2006), and using gradient-based methods. Problems of this size are, practically speaking, out of the range of current deterministic global optimization technology , so we are left to consider heuristics and gradient-based approaches. W e prefer Bayesian optimization to heuristics such as genetic algo- rithms because the GPs allow us to immediately study sensitivity in the model response surface. Gradient-based methods directly benefit from the proposed Bayesian optimization approach since we can warm start the gradient optimization at promising initialization points. Evaluating the microalgae metabolism model in our application so quickly leads to data set sized standard BO cannot manage. T o address this problem we introduce Bayesian Optimization with Dimension Scheduling Algorithm (DSA). The DSA distributes the training data across many GPs, with each GP containing training data of a subset of the dimensions. At each iteration, a new subset of the dimensions is sampled from a probability distribution, which reflects the importance of the corresponding parameters, and the utility or acquisition function is optimized with respect to this subset. DSA benefits from a faster computational performance because each iteration considers a relativ ely small number of prior experiments. 2. Backgr ound and Related W ork Bayesian Optimization (BO) (Kushner, 1964) is a global black-box technique for optimizing f : R d 7→ R , which may be non-con ve x, on a d -dimensional hyper-rectangle with finite bounds B (Jones et al., 1998): x ∗ ∈ arg min x ∈ B ⊂ R d f ( x ) . (1) In the BO context, we consider a regression problem y = f ( x ) + ε , where x ∈ R d , y ∈ R , and f is the unknown fitness/utility function. The Gaussian likelihood p ( y | f ( X )) = N ( f ( x ) , σ 2 ε ) accounts for the independently and identically distributed measurement noise ε ∼ N ( 0 , σ 2 ε ) . The objective is to infer the latent utility function f from a training data set X = { x i } N i = 1 , Y = { y i } N i = 1 . A GP is defined as a collection of random v ariables, any finite number of which is Gaussian distributed. A GP is fully specified by a mean function m and a covariance function k (kernel) with hyper- parameters ψ , which allows us to encode high-lev el structural assumptions, e.g., differentiability . W ithout loss of generality , we assume that the prior mean function is 0. A GP is trained by finding hyper-parameters θ = { ψ , σ ε } that maximize the log-marginal likelihood log p ( Y | X , θ ) = − 1 2 Y T ( K + σ 2 ε I ) − 1 Y + log | K + σ 2 ε I | + const , (2) Bayesian Optimization with Dimension Scheduling 3 Algorithm 1: Bayesian Optimization Evaluate the objecti ve function f N times; Obtain initial training set ( X , Y ) ; for n = N to maxIter do Update the GP with ( X , Y ) ; x n + 1 = arg min x a ( x | GP n ) ; Evaluate objecti ve function y n + 1 = f ( x n + 1 ) + ε ; Augment training set X , Y with ( x n + 1 , y n + 1 ) ; end where K = k ( X , X ) ∈ R N × N is the kernel matrix. For a gi ven set of hyper-parameters θ , a training set X , Y and a test input x ∗ ∈ R d , the GP posterior predictiv e distribution of the corresponding function value f ∗ = f ( x ∗ ) is Gaussian with mean and variance gi ven by E [ f ∗ ] = k T ∗ ( K + σ 2 ε I ) − 1 Y , var [ f ∗ ] = k ∗∗ − k T ∗ ( K + σ 2 ε I ) − 1 k ∗ , (3) respectiv ely , where we defined k ∗ = k ( X , x ∗ ) and k ∗∗ = k ( x ∗ , x ∗ ) . BO places a GP prior on the fitness function f , which serves as a probabilistic surrogate model for f . Instead of optimizing f , which will require many function ev aluations, BO optimizes an acquisition function a ( · ) that depends on the GP posterior predictive distribution to decide the next e valuation point (e xperiment). 1 This experiment yields an additional training point ( x i , y i ) , which updates the GP model. After updating the GP with the new observation, BO repeats the cycle until con ver gence or an upper bound on the total number of experiments. BO is summarized in Alg. 1. There hav e been approaches toward scaling BO to more dimensions. The Additive BO method (ABO) developed by Kandasamy et al. (2015) is the variant of BO most similar to our DSA contribution: ABO assumes that the objective function f ( x ) can be decomposed into M functions f ( x ) = ∑ m k = 1 f k ( x k ) operating on mutually disjoint dimensions, such that x i ∩ x j = / 0 for i 6 = j . Because of the decomposability assumption, each of the M functions can be optimized separately . ABO expedites typical BO approaches by explicitly breaking dependence between the functional domains. Although Kandasamy et al. (2015) show that their method is applicable to non-additiv e functions we find that, in the case of our parameter estimation problem, we do not get good results using ABO. W e no w introduce DSA, which shares the ABO goal of cutting BO computation time but abandons the assumption of functional decomposability and, in particular , the idea of mutually disjoint parameter domains. 3. Bay esian Optimization with Dimension Scheduling Algorithm The DSA algorithm is an extension of the traditional BO algorithm and is summarized in Alg. 2. The initialization corresponds to the standard procedure of BO and observed function values for a few sampled x i . Based on the set of initial samples X and corresponding observations Y we set x b = arg min X Y and y b = min ( Y ) , where x b stores our best kno wn argument for the best objecti ve value, and y b stores the best objective value. DSA samples coordinates from a probability v ector P , which represents the relativ e importance of the x coordinates. For this purpose, we apply PCA to the av ailable data X and set P proportional to the eigenv alue magnitude ev ery 50 iterations. DSA now samples a random dimension set Z from P . W e no w optimize the Z -coordinates of x while clamping the other coordinates to their current best values. For optimization we use a GP model GP Z , which possesses as input dimensions only the Z -coordinates of X . If GP Z does not yet exists, 1 In this paper, we choose Expected Improvement (Mockus et al., 1978) as acquisition function and use DIRECT (Jones et al., 1993) to optimize it. 4 Ulmasov , Bar oukh, Chachuat, Deisenr oth, Misener Algorithm 2: Bayesian Optimization with Dimension Scheduling Algorithm Evaluate objecti ve function N times, update the all GPs with sampled data ( X , Y ) ; Set x b = arg min f ( X ) and y b = min Y ; for n = N to maxIter do Update probability vector P from the observations; Randomly sample dimension set Z from P ; Find x Z n + 1 = arg min a ( x Z n + 1 , GP Z ) ; Replace coordinates Z of x n + 1 with x Z n + 1 ; Evaluate objecti ve function y n + 1 = f ( x n + 1 ) + ε n + 1 ; Augment training set of GP Z with ( x Z n + 1 , y Z n + 1 ) ; if y n + 1 < y b then ( x b , y b ) = ( x n + 1 , y n + 1 ) ; end T able 1: Best achie ved objective in four experiments using standard BO and DSA. Each e xperi- ment ran for 500 iterations Model d BO:1 DSA:1 BO:2 DSA:2 BO:3 DSA:3 BO:4 DSA:4 M19 10 54.62 32.32 52.66 47.20 39.35 30.46 41.70 24.48 M19o 10 77.78 66.09 114.78 354.81 78.64 133.36 80.05 69.54 M26 11 87.91 35.86 32.56 26.66 55.40 31.53 57.58 26.54 M29G 11 46.32 59.77 72.19 35.16 86.61 77.40 73.62 64.59 M29C 11 36.50 24.78 44.71 37.25 41.47 34.59 52.74 33.77 M31 11 39.24 24.37 60.39 29.75 49.52 26.26 44.18 48.53 M32C 12 43.33 33.14 52.19 31.15 50.62 28.02 51.13 26.18 M32G 12 42.51 27.12 47.55 26.78 52.38 31.45 53.21 23.54 M33 12 44.07 24.82 38.78 24.64 40.26 29.09 46.68 20.61 M35 12 50.16 25.86 57.72 22.86 57.06 28.97 57.00 40.11 we spawn a new GP model where we use the relev ant coordinates of the initial samples ( X , Y ) as training data. For the optimization, we optimize the acquisition function a ( · ) related to GP Z and obtain x ( Z ) n + 1 . W e perform an experiment with the ne w x b and if the observed function v alue y n + 1 is a better objecti ve v alue than y b , we update y b to y n + 1 and x b to x n + 1 . Note that we effecti vely only update the dimensions Z in x b with parameter values from x Z n + 1 . The new data point ( x n + 1 , y n + 1 ) is only added to GP Z , i.e., the data set size of the other GP models does not increase. 4. Results W e focus on parameter estimation for dynamic models of microalgae metabolism (Baroukh et al., 2014). The objective is to minimize a weighted sum of the squared errors between the model simulations and the real w orld e xperimental data. The weights were chosen as the mean of each kind of experimental data so that each kind of measurement (biomass, lipids, etc.) contributes equally to the error . Each model simulation is e v aluated in a fraction of the second, b ut the wide bounds and the number of dimensions creates a large search area which prohibits random sampling methods. T o compare traditional BO (Hoffman and Shahriari, 2014) and DSA, we tested 10 dif ferent model variants with 4 experiment runs per model. Each model variant, e.g., M19, represents a dif ferent biological hypothesis on microalgae metabolism (Baroukh et al., 2014) and may have a varying number of parameters or possible parameter bounds. Each experiment ran for 500 iterations, with the same set of randomly sampled data, a Squared Exponential kernel and the Expected Bayesian Optimization with Dimension Scheduling 5 Figure 1: A v erage running time best objectiv e v alue of dif ferent subset sizes with respect to fi ve av eraged runs of M19. Lower is better . Figure 2: Best running objecti ve value of DSA and classical BO for three models. Lo wer is better . Improv ement acquisition function. In all cases we chose dimension size 2 for the DSA. Figure 4 motiv ates using DSA dimension 2 by diagramming the average results from 5 experiments on model M19; note that the av erage running time and objectiv e values are consistently better for dimension 2. T able 1 summarizes model details and results of the 4 runs, the column d stands for dimensions in the model. In most cases, DSA terminated with a lower objective value than traditional BO. The exceptions arise due to the limitations of DSA discussed in the next section. Figure 4 presents a sample of running best objecti ve values. The traditional BO method tends to improve less frequently as the optimization process progresses. The DSA method, due to the constant changes of the dimensions, tends to improve the objectiv e value more frequently and achiev es an overall lower objecti ve value. Due to random sampling, the DSA performance is not uniformly superior to traditional BO, as seen in T able 1. The DSA algorithm completes each experiment on av erage at a fifth of the computation time of BO: Figure 4 sho ws the av erage computation time of the experiment runs with all the models. The performance increase stems from the reduced number of GP dimensions and reduced number of observations per GP . The computational complexity of the predictions are still O n 2 . Howe ver , by distributing the training data across multiple GPs the training sets per GP are relativ ely small (proportional to ho w often a set of dimensions Z has been sampled from the probability vector P ). The number of GPs used by the algorithm is upper bounded by the total number of permutations 6 Ulmasov , Bar oukh, Chachuat, Deisenr oth, Misener Figure 3: A v erage computation time of DSA and traditional BO for all models. DSA results in a substantial computational speed-up. of the set Z , which itself is based on the subset sizes and problem dimensionality . 5. Discussion Since the DSA algorithm never uses all of the data, we cannot assert global algorithm con ver gence. The issue of con vergence could be addressed, for example, by a hybrid BO solution with the function space sampled for near optimal solutions with DSA. The DSA addresses only a subset of problems with BO under specific conditions. The algorithm faces similar challenges when we increase number of total dimensions d , and the GP’ s accuracy would suffer since the observ ation points would be spread thinly ov er many GPs. In the current implementation, the maximum number of GPs is equiv alent to number of permutations of the subset Z used by the DSA. Howe ver , not all GPs are equally rele vant, and some pruning may be beneficial to limit the number of models. The mar ginal likelihood could be used for this purpose. One of the advantages of the Bayesian Optimization with Dimension Scheduler over traditional Bayesian Optimization is a f airly straightforward code parallelization for increased performance on multicore systems. The traditional Bayesian Optimization method can be parallelized to a certain extent. There has been some work on Gaussian process models in distributed systems (Deisenroth and Ng, 2015; Gal et al., 2014), which allo ws GPs to scale to larger data sets. The solvers can be parallelized by dividing the search space between different processes. These ap- proaches provide compartmental parallelization, b ut the whole process is still sequential. The DSA parallelization process is much simpler , and can use current GP and solver modules. The solution lies in GP distribution across man y processes, and a manager process to communicate be- tween each child-process and the objecti ve function. The manager would assign different iteration points to each process. Each process would contain a GP and a solver , and based on the iteration number the process receiv es (if we hav e exploration parameter scheduler), the process maximizes the acquisition function for the GP in the process and returns the solution. The manager would ev aluate the solution and return it to the process to update the GP and start a ne w iteration. 6. Conclusion W e have identified a performance issue with traditional BO when the experimentation time is significantly less than the optimization time and hav e therefore dev eloped DSA. In the case of the microalgae model, DSA out-performed traditional BO in terms of computation time and the best objectiv e value. The DSA could be applied to other models where a relativ ely quick com- putational model with high dimensionality requires ef ficient parameter estimation. Further DSA Bayesian Optimization with Dimension Scheduling 7 improv ement, such as parallel implementation and GPs reduction based on the marginal likeli- hood w ould lead to even greater performance benefits; the parallelization potential suggests that the algorithm may be able to manage more dimensions than traditional BO. Acknowledgments This work was supported by the Engineering and Physical Sciences Research Council [EP/M028240/1] and a Royal Academy of Engineering Research Fello wship to R.M. References C. Baroukh, R. Mu ˜ noz-T amayo, J.-P . Steyer , O. Bernard, 2014. DRUM: A ne w frame work for metabolic modeling under non-balanced growth. application to the carbon metabolism of unicellular microalgae. PLoS ONE 9 (8), e104499. R. Calandra, A. Seyfarth, J. Peters, M. P . Deisenroth, 2015. Bayesian optimization for learning gaits under uncertainty . Ann Math and AI, 1–19. G. E. Dahl, T . N. Sainath, G. E. Hinton, 2013. Improving deep neural netw orks for L VCSR using rectified linear units and dropout. In: Proc. ICASSP . pp. 8609–8613. M. P . Deisenroth, J. W . Ng, 2015. Distributed Gaussian processes. In: Proc. ICML. C. A. Floudas, P . M. Pardalos (Eds.), 2000. Optimization in computational chemistry and molecular biology: Local and global approaches. Kluwer. Y . Gal, M. van der W ilk, C. E. Rasmussen, 2014. Distributed v ariational inference in sparse Gaussian process re gression and latent variable models. In: Proc. NIPS. M. W . Hoffman, B. Shahriari, 2014. Modular mechanisms for Bayesian optimization. In: NIPS W orkshop on Bayesian Optimization. F . Hutter , H. H. Hoos, K. Leyton-Brown, 2011. Sequential model-based optimization for general algorithm configuration. In: Proc. LION. Springer , pp. 507–523. D. R. Jones, C. D. Perttunen, B. E. Stuckman, 1993. Lipschitzian optimization without the Lipschitz constant. J Optim Theory Appl 79 (1), 157–181. D. R. Jones, M. Schonlau, W . J. W elch, 1998. Efficient global optimization of expensive black-box functions. J Glob Optim 13 (4), 455–492. K. Kandasamy , J. Schneider , B. Poczos, 2015. High dimensional Bayesian optimization and bandits via additiv e models. arXiv preprint arXi v:1503.01673. H. J. Kushner, 1964. A ne w method of locating the maximum point of an arbitrary multipeak curve in the presence of noise. J of Basic Engineering 86, 97. T . Lacour, A. Sciandra, A. T alec, P . Mayzaud, O. Bernard, 2012. Diel variations of carbohydrates and neutral lipids in nitrogen-sufficient and nitrogen-starv ed cyclostat cultures of isochrysis sp.1. J Phycol 48 (4), 966–975. C. Michalik, B. Chachuat, W . Marquardt, 2009. Incremental global parameter estimation in dynamical systems. Ind Eng Chem Res 48 (11), 5489–5497. J. Mockus, V . T iesis, A. Zilinskas, 1978. The application of Bayesian methods for seeking the extremum. T o wards Global Optimization 2, 117–129. M. P . Rodrigues-Fernandez, J. R. Banga, 2006. A hybrid approach for efficient and robust parameter estimation in biochemical pathways. Biosystems 83, 248–265. M. P . Rodrigues-Fernandez, J. Egea, J. R. Banga, 2006. Novel metaheuristic for parameter estimation in nonlinear dynamic biological systems. BMC Bioinformatics 7 (1), 483. J. Snoek, H. Larochelle, R. P . Adams, 2012. Practical Bayesian optimization of machine learning algorithms. In: Proc. NIPS.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment