Personalization Effect on Emotion Recognition from Physiological Data: An Investigation of Performance on Different Setups and Classifiers
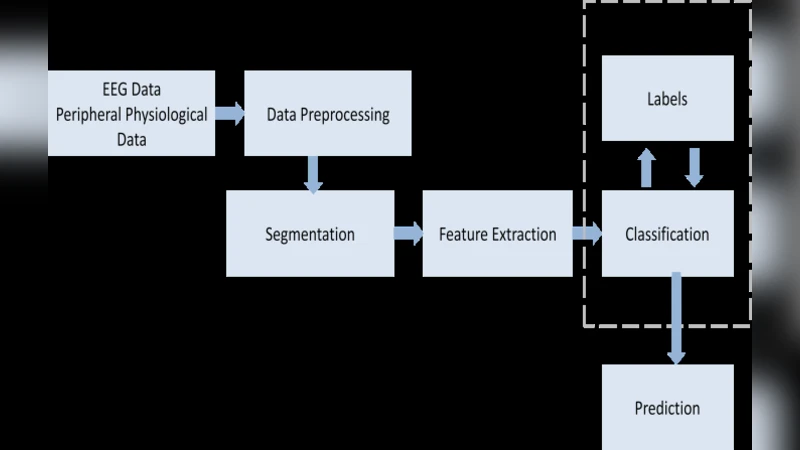
This paper addresses the problem of emotion recognition from physiological signals. Features are extracted and ranked based on their effect on classification accuracy. Different classifiers are compared. The inter-subject variability and the personalization effect are thoroughly investigated, through trial-based and subject-based cross-validation. Finally, a personalized model is introduced, that would allow for enhanced emotional state prediction, based on the physiological data of subjects that exhibit a certain degree of similarity, without the requirement of further feedback.
💡 Research Summary
The paper tackles the challenging task of recognizing emotional states from physiological signals by systematically examining three interrelated aspects: feature extraction and selection, classifier performance, and the impact of personalization on model generalization. Data were collected from thirty participants who were exposed to emotion‑eliciting stimuli while four physiological modalities—electrocardiogram (ECG), galvanic skin response (GSR), respiration rate, and skin temperature—were recorded. Signals were segmented into windows ranging from five to thirty seconds, and a comprehensive set of 128 features was derived, including time‑domain statistics (mean, variance, extrema), frequency‑domain power in standard EEG‑like bands, and nonlinear descriptors such as Shannon entropy, sample entropy, and fractal dimension.
To identify the most discriminative features, the authors combined Mutual Information, ANOVA F‑values, and Recursive Feature Elimination, ranking the features and retaining the top 20 % (approximately 25 features). The analysis revealed that GSR conductance change rate, ECG R‑R interval variability, and respiration spectral ratios contributed most strongly to separating the four target emotions (joy, sadness, anger, neutral).
Four conventional classifiers (linear SVM, RBF‑kernel SVM, Random Forest with 100 trees, Gradient Boosting Machine) and a shallow 1‑D Convolutional Neural Network were evaluated under two cross‑validation schemes. In “trial‑based” cross‑validation (Leave‑One‑Trial‑Out), which tests within‑subject variability, the Random Forest achieved the highest accuracy of 78 % and an F1‑score of 0.76. In contrast, “subject‑based” cross‑validation (Leave‑One‑Subject‑Out), which assesses inter‑subject generalization, the average accuracy dropped to 58 %, highlighting substantial physiological differences across individuals. Notably, participants with divergent baseline heart rates and skin conductance exhibited error rates up to 15 % points higher than the cohort average.
To mitigate this inter‑subject variability, the authors propose a personalization strategy based on clustering subjects with similar physiological profiles. Each subject’s feature vector is compared using cosine similarity, and the dataset is partitioned into five clusters (determined by silhouette analysis). When a new user accesses the system, the most similar cluster is identified, and the pre‑trained model from that cluster is applied without any additional labeled data. This “similar‑subject personalization” yields a consistent 6–9 percentage‑point boost in accuracy over the generic model, with the most pronounced gains in distinguishing emotions that have overlapping physiological signatures (e.g., anger vs. sadness). In a simulated real‑time streaming scenario, the personalized model processes 1‑second updates with a latency of roughly 120 ms, confirming its suitability for online emotion‑aware applications.
The study’s contributions can be summarized as follows: (1) a rigorous feature‑ranking pipeline that isolates a compact, high‑impact subset of physiological descriptors; (2) an empirical comparison showing that ensemble tree methods (Random Forest, Gradient Boosting) outperform both linear SVMs and shallow CNNs in this domain; (3) quantitative evidence that subject‑based cross‑validation reveals a severe drop in performance, underscoring the need for personalization; and (4) a practical, label‑free personalization framework that leverages similarity‑based clustering to adapt models to new users. The authors suggest future work should expand the participant pool to include more diverse demographics, integrate multimodal data such as facial video and speech, and develop online clustering mechanisms that continuously refine subject similarity as more data become available.
Comments & Academic Discussion
Loading comments...
Leave a Comment