De-identification of Patient Notes with Recurrent Neural Networks
Objective: Patient notes in electronic health records (EHRs) may contain critical information for medical investigations. However, the vast majority of medical investigators can only access de-identified notes, in order to protect the confidentiality of patients. In the United States, the Health Insurance Portability and Accountability Act (HIPAA) defines 18 types of protected health information (PHI) that needs to be removed to de-identify patient notes. Manual de-identification is impractical given the size of EHR databases, the limited number of researchers with access to the non-de-identified notes, and the frequent mistakes of human annotators. A reliable automated de-identification system would consequently be of high value. Materials and Methods: We introduce the first de-identification system based on artificial neural networks (ANNs), which requires no handcrafted features or rules, unlike existing systems. We compare the performance of the system with state-of-the-art systems on two datasets: the i2b2 2014 de-identification challenge dataset, which is the largest publicly available de-identification dataset, and the MIMIC de-identification dataset, which we assembled and is twice as large as the i2b2 2014 dataset. Results: Our ANN model outperforms the state-of-the-art systems. It yields an F1-score of 97.85 on the i2b2 2014 dataset, with a recall 97.38 and a precision of 97.32, and an F1-score of 99.23 on the MIMIC de-identification dataset, with a recall 99.25 and a precision of 99.06. Conclusion: Our findings support the use of ANNs for de-identification of patient notes, as they show better performance than previously published systems while requiring no feature engineering.
💡 Research Summary
The paper addresses the critical need for automated de‑identification of patient notes in electronic health records (EHRs) to comply with HIPAA’s 18 protected health information (PHI) categories. Manual annotation is prohibitively expensive and error‑prone, especially for large corpora such as the MIMIC database. Existing solutions fall into two camps: rule‑based systems that rely on handcrafted regular expressions and gazetteers, and machine‑learning systems—most notably conditional random fields (CRFs)—that require extensive feature engineering and sizable labeled datasets.
To overcome these limitations, the authors propose the first de‑identification system built entirely on artificial neural networks (ANNs), eliminating the need for manual feature design. The architecture consists of three principal layers. First, a character‑enhanced token embedding layer combines a standard word embedding (learned or pre‑trained) with a character‑level representation produced by a bidirectional LSTM over the token’s characters. This hybrid embedding captures out‑of‑vocabulary words, misspellings, and morphological cues. Second, a label prediction layer feeds the sequence of token embeddings into another bidirectional LSTM, whose hidden states are passed through a feed‑forward network to generate a probability distribution over all PHI labels (including a “non‑PHI” class) for each token. Third, a label‑sequence optimization layer incorporates a transition matrix that models the probability of moving from one label to the next; the overall sequence score combines per‑token probabilities and transition scores, and the optimal label sequence is obtained via Viterbi decoding. All components are trained jointly by maximizing the log‑likelihood of the gold label sequences.
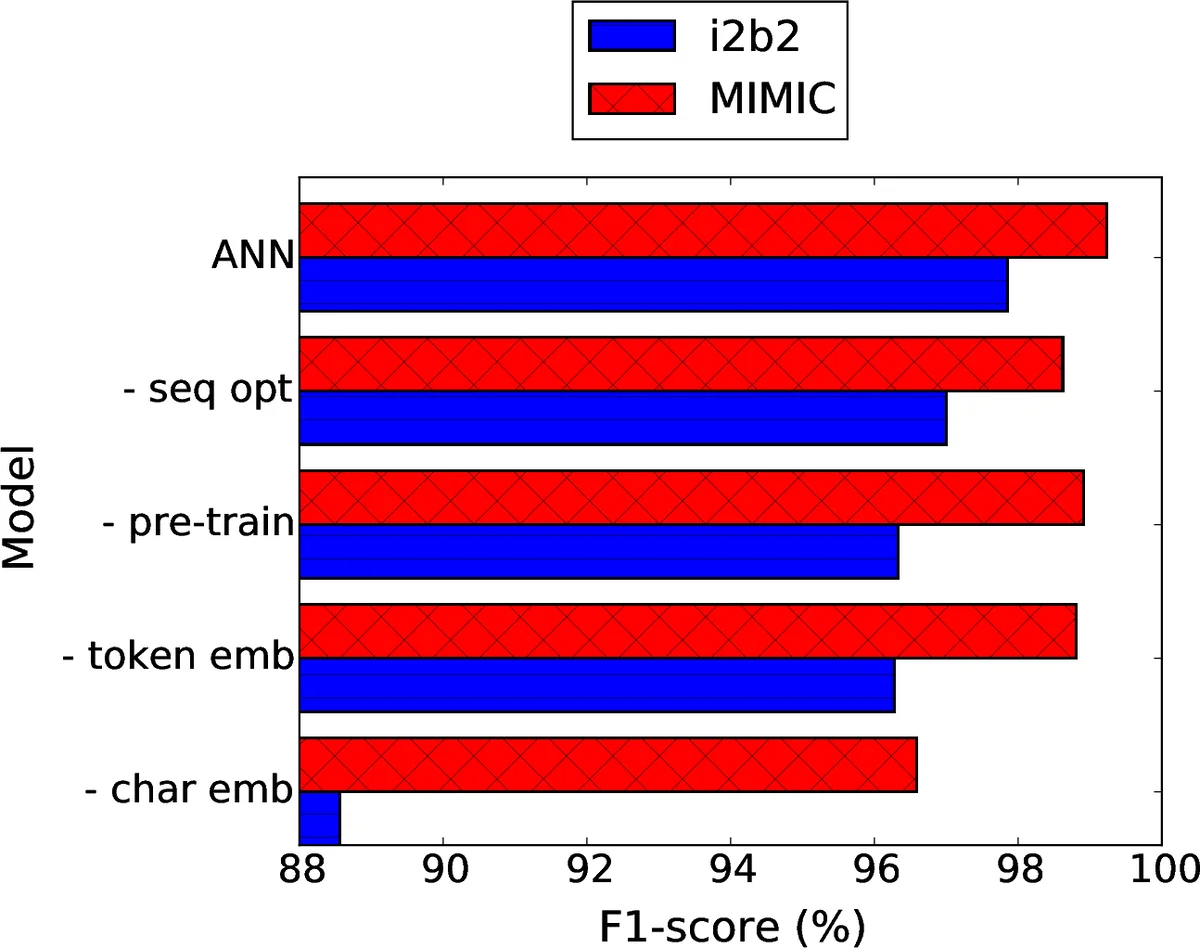
The system was evaluated on two benchmark datasets: the i2b2 2014 de‑identification challenge set (the largest publicly available PHI‑annotated corpus) and a newly assembled MIMIC de‑identification set that is roughly twice as large. Compared against a strong CRF baseline (using n‑gram, morphological, orthographic, and gazetteer features) and previously published state‑of‑the‑art methods, the ANN model achieved superior results: an F1‑score of 97.85 (precision 97.32, recall 97.38) on i2b2 and an even higher F1‑score of 99.23 (precision 99.06, recall 99.25) on MIMIC. These figures demonstrate that the neural approach not only matches but exceeds the performance of heavily engineered systems, while being more robust to variations in language, spelling, and tokenization.
Key contributions include: (1) introducing a fully neural de‑identification pipeline that eliminates handcrafted features; (2) leveraging character‑level LSTM embeddings to handle out‑of‑vocabulary and noisy tokens; (3) integrating a CRF‑style transition layer within the neural network for coherent label sequences; and (4) validating the model’s scalability and generalization across two large, heterogeneous clinical corpora. The authors suggest future work on domain‑specific pre‑training, multi‑task learning (e.g., simultaneous concept extraction), and deployment in real‑time clinical settings, underscoring the practical impact of ANN‑based de‑identification for biomedical research and data sharing.
Comments & Academic Discussion
Loading comments...
Leave a Comment